Abstract
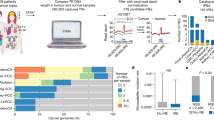
Messenger RNA contains untranslated 3′ and 5′ regions (3′ and 5′ UTRs) with sequence elements that are essential for the regulation of gene expression. A systematic search of GenBank revealed a large number of mononucleotide repeats within these UTRs. We selected 35 such mononucleotide repeats ranging in length from 15 bp to 32 bp and analysed their size in a series of 60 normal individuals. The conservation of repeats correlated inversely to their length, with longer repeats generally being more polymorphic than shorter repeats, irrespective of 3′ or 5′ location. Several long repeats were identified however to be monomorphic and we postulate that their conservation may be due to selective pressures relating to a possible functional role. We analysed 19 conserved UTR repeats in 117 colorectal cancers (CRC), 43 of which had defective mismatch repair characterized by widespread microsatellite instability (MSI-H). The UTR repeats were very often deleted in MSI-H tumors, with the length of deletion being proportional to the size of the repeat. Because of the high frequency of deletion observed in the conserved UTR repeats of MSI-H tumors, these could serve as a useful model for the study of possible changes in gene expression resulting from such mutations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aaltonen LA, Peltomäki P, Leach FS, Sistonen P, Pylkkänen L, Mecklin JP, Järvinen H, Powell SM, Jen J, Hamilton SR, Petersen GM, Kinzler KW, Vogelstein B, de la Chapelle A . 1993 Science 260: 812–816
Aharoni A, Baran N, Manor H . 1993 Nucleic Acids Res. 21: 5221–5228
Buchman AR, Kornberg RD . 1990 Mol. Cell. Biol. 10: 887–897
Conne B, Stutz A, Vassalli JD . 2000 Nature Med. 6: 637–641
Duret L, Dorkeld F, Gautier C . 1993 Nucleic Acids Res. 21: 2315–2322
Duval A, Rolland S, Compoint A, Tubacher E, Iacopetta B, Thomas G, Hamelin R . 2001 Hum. Mol. Gen. 10: 513–518
Duval A, Gayet J, Zhou XP, Iacopetta B, Thomas G, Hamelin R . 1999 Cancer Res. 59: 4213–4215
Eshleman JR, Markowitz SD . 1995 Curr. Opin. Oncol. 7: 83–89
Fu L, Minden MD, Benchimol S . 1996 EMBO J. 15: 4392–4401
Goller M, Funke B, Gehe-Becker C, Kroger B, Lottspeich F, Horak I . 1994 Nucleic Acids Res. 22: 1885–1889
Hoang JM, Cottu PH, Thuille B, Salmon RJ, Thomas G, Hamelin R . 1997 Cancer Res. 57: 300–303
Ionov Y, Peinado M, Malkhosyan S, Shibata D, Perucho M . 1993 Nature 363: 558–561
Johnson AC, Jinno Y, Merlino GT . 1988 Mol. Cell Biol. 8: 4174–4184
Kashi Y, King D, Soller M . 1997 Trends in Genet. 13: 74–78
Levinson G, Gutman GA . 1987 Mol. Biol. Evol. 4: 202–221
Lue NF, Buchman AR, Kornberg RD . 1989 Proc. Natl. Acad. Sci. USA 86: 486–490
Markowitz S, Wang J, Myeroff L, Parsons R, Sun L, Lutterbaugh J, Fan RS, Zborowska E, Kinzler KW, Vogelstein B, Brattain M, Willson JKV . 1995 Science 268: 1336–1338
Nowak R . 1994 Science 263: 608–610
Percesepe A, Pedroni M, Sala E, Menigatti M, Borghi F, Losi L, Viel A, Genuardi M, Benatti P, Roncucci L, Peltomaki P, Ponz de Leon M . 2000 Genes Chrom. Cancer 27: 424–429
Pesole G, Liuni S, D'Souza M . 2000 Bioinformatics 16: 439–450
Pesole G, Liuni S, Grillo G, Saccone C . 1997 Gene 205: 95–102
Rampino N, Yamamoto H, Ionov Y, Li Y, Sawai H, Reed JC, Perucho M . 1997 Science 275: 967–969
Sonenberg N . 1994 Curr. Opin. Genet. Dev. 4: 310–315
Souza RF, Appel R, Yin J, Wang S, Smolinski KN, Abraham JM, Zou TT, Shi YQ, Lei J, Cottrell J, Cymes K, Biden K, Simms L, Leggett B, Lynch PM, Frazier M, Powell SM, Harpaz N, Sugimura H, Young J, Meltzer SJ . 1996 Nature Genet 14: 255–257
Sturzeneker R, Bevilacqua RAU, Haddad LA, Simpson AJG, Pena SDJ . 2000 Hum. Mol. Gen. 9: 347–352
Thibodeau SN, Bren G, Schaid D . 1993 Science 260: 816–819
Thompson MA, Flegg R, Westin EH, Ramsey RG . 1997 Oncogene 14: 1715–1723
Winter E, Varshavsky A . 1989 EMBO J. 8: 1867–1877
Yee HA, Wong AK, Van de Sande JH, Rattner JB . 1991 Nucleic Acids Res. 19: 949–953
Zhang L, Yu J, Willson JKV, Markowitz SD, Kinzler KW, Vogelstein B . 2001 Cancer Res. 61: 3801–3805
Zhou XP, Hoang JM, Cottu P, Thomas G, Hamelin R . 1997 Oncogene 15: 1713–1718
Acknowledgements
This work was funded by grant 5502 from the Association pour la Recherche sur le Cancer, the Raine Medical Research Foundation, and the Cancer Foundation of Western Australia.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Suraweera, N., Iacopetta, B., Duval, A. et al. Conservation of mononucleotide repeats within 3′ and 5′ untranslated regions and their instability in MSI-H colorectal cancer. Oncogene 20, 7472–7477 (2001). https://doi.org/10.1038/sj.onc.1204952
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204952
Keywords
This article is cited by
-
Microsatellite instability assessment is instrumental for Predictive, Preventive and Personalised Medicine: status quo and outlook
EPMA Journal (2023)
-
Frequent mutations in the 3′-untranslated region of IFNGR1 lack functional impairment in microsatellite-unstable colorectal tumours
European Journal of Human Genetics (2008)