Abstract
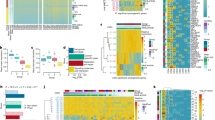
We used a classical rodent model of transformation to understand the transcriptional processes, and hence the molecular and cellular events a given cell undergoes when progressing from a normal to a transformed phenotype. Src activation is evident in 80% of human colon cancer, yet the myriad of cellular processes effected at the level of gene expression has yet to be fully documented. We identified a Src ‘transformation fingerprint’ within the gene expression profiles of Src-transformed rat 3Y1 fibroblasts demonstrating a progression in transformation characteristics. To evaluate the role of this gene set in human cancer development and progression, we extracted the orthologous genes present on the Affymetrix Hu95A GeneChip™ (12k named genes) and compared expression profiles between the Src-induced rodent cell line model of transformation and staged colon tumors where Src is known to be activated. A similar gene expression pattern between the cell line model and staged colon tumors for components of the cell cycle, cytoskeletal associated proteins, transcription factors and lysosomal proteins suggests the need for co-regulation of several cellular processes in the progression of cancer. Genes not previously implicated in tumorigenesis were detected, as well as a set of 14 novel, highly conserved genes with here-to-fore unknown function. These studies define a set of transformation associated genes whose up-regulation has implications for understanding Src mediated transformation and strengthens the role of Src in the development and progression of human colon cancer. Supportive Supplemental Data can be viewed at http://pga.tigr.org/PGApubs.shtml.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Akamatsu H, Ichihara-Tanaka K, Ozono K, Kamiike W, Matsuda H, Sekiguchi K . 1996 Cancer Res. 56: 4541–4546
Alter O, Brown PO, Botstein D . 2000 Proc. Natl. Acad. Sci. USA 97: 10101–10106
Ben-Dor A, Shamir R, Yakhini Z . 1999 J. Comput. Biol. 6: 281–297
Bjorge JD, Bellagamba C, Cheng HC, Tanaka A, Wang JH, Fujita DJ . 1995 J. Biol. Chem. 270: 2422–2428
Brown MT, Cooper JA . 1996 Biochim. Biophys. Acta 1287: 121–149
Byrne JA, Tomasetto C, Garnier JM, Rouyer M, Mattei MG, Bellocq JP, Rio MC, Basset P . 1995 Cancer Res. 55: 2896–2903
Chew CS, Parente JAJ, Chen X, Chaponnier C, Cameron RS . 2000 J. Cell Sci. 113: 2035–2045
DeRisi JL, Iyer VR, Brown PO . 1997 Science 278: 680–686
Eisen MB, Spellman PT, Brown PO, Botstein D . 1998 Proc. Natl. Acad. Sci. USA 95: 14863–14868
Freier S, Weiss P, Eran M, Flyvbjerg A, Dahan R, Nephesh I, Safra T, Shiloni E, Raz I . 1999 Gut 44: 704–708
Guo QM, Malek RL, Kim S, Chiao C, He M, Ruffy M, Sanka K, Lee NH, Dang CV, Liu ET . 2000 Cancer Res. 60: 5922–5928
Iba H, Takeya T, Cross FR, Hanafusa T, Hanafusa H . 1984 Proc. Natl. Acad. Sci. USA 81: 4424–4428
Inoue H, Pan J, Hakura A . 1998 J. Virol. 72: 2532–2537
Iravani S, Mao W, Fu L, Karl R, Yeatman T, Jove R, Coppola D . 1998 Lab. Invest. 78: 365–371
Irby RB, Mao W, Coppola D, Kang J, Loubeau JM, Trudeau W, Karl R, Fujita DJ, Jove R, Yeatman TJ . 1999 Nat. Genet. 21: 189–190
Johnson PJ, Coussens PM, Danko AV, Shalloway D . 1985 Mol. Cell. Biol. 5: 1073–1083
Jove R, Hanafusa H . 1987 Annu. Rev. Cell Biol. 3: 31–56
Jove R, Hanafusa T, Hamaguchi M, Hanafusa H . 1989 Oncogene Res. 5: 49–60
Kato JY, Takeya T, Grandori C, Iba H, Levy JB, Hanafusa H . 1986 Mol. Cell. Biol. 6: 4155–4160
Kraker AJ, Hartl BG, Amar AM, Barvian MR, Showalter HD, Moore CW . 2000 Biochem. Pharmacol. 60: 885–898
Kumble S, Omary MB, Cartwright CA, Triadafilopoulos G . 1997 Gastroenterology 112: 348–356
Lee NH, Malek RL . 1998 J. Biol. Chem. 273: 22317–22325
Lee Y, Sultana R, Pertea G, Cho J, Karamycheva S, Tsai J, Parvizi B, Cheug F, Antoescu V, White J, Holt I, Liang F, Quackenbush J . 2002 Genome Res. 12: 493–502
Levy JB, Iba H, Hanafusa H . 1986 Proc. Natl. Acad. Sci. USA 83: 4228–4232
Lutz MP, Esser IB, Flossmann-Kast BB, Vogelmann R, Luhrs H, Friess H, Buchler MW, Adler G . 1998 Biochem. Biophys. Res. Commun. 243: 503–508
Manenti S, Malecaze F, Chap H, Darbon JM . 1998 Cancer Res. 58: 1429–1434
Martin GS . 2001 Nat. Rev. Mol. Cell Biol. 2: 467–475
Miller LD, Park KS, Guo QM, Alkharouf NW, Malek RL, Lee NH, Liu ET, Cheng S . 2001 Mol. Cell. Biol. 21: 6626–6639
Muschel RJ, Williams JE, Lowy DR, Liotta LA . 1985 Am. J. Pathol. 121: 1–8
Niinaka Y, Paku S, Haga A, Watanabe J, Raz A . 1998 Cancer Res. 58: 2667–2674
Odajima J, Matsumura I, Sonoyama J, Daino H, Kawasaki A, Tanaka H, Inohara N, Kitamura T, Downward J, Nakajima K, Hirano T, Kanakura Y . 2000 J. Biol. Chem. 275: 24096–24105
Okamoto-Inoue M, Taniguchi S, Sadano H, Kawano T, Kimura G, Gabbiani G, Baba T . 1990 J. Cell Sci. 96: 631–637
Parker RC, Varmus HF, Bishop JM . 1984 Cell 37: 131–139
Quackenbush J, Liang F, Holt I, Petrea G, Upton J . 2000 Nucleic Acids Res. 28: 141–145
Raychaudhuri S, Stuart JM, Altman RB . 2000 Pac. Symp. Biocomput. 5: 455–466
Reimer CL, Borras AM, Kurdistani SK, Garreau JR, Chung M, Aaronson SA, Lee SW . 1999 J. Biol. Chem. 274: 11022–11029
Sabo SL, Ikin AF, Buxbaum JD, Greengard P . 2001 J. Cell Sci. 153: 1403–1414
Saitoh T, Sundsmo M, Roch JM, Kimura N, Cole G, Schubert D, Oltersdorf T, Schenk DB . 1989 Cell 58: 615–622
Schreiber V, Moog-Lutz C, Regnier CH, Chenard MP, Boeuf H, Vonesch JL, Tomasetto C, Rio MC . 1998 Mol. Med. 4: 675–687
Schubert D, Jin LW, Saitoh T, Cole G . 1989 Neuron 3: 689–694
Sefton BM, Hunter T, Beemon K, Eckhart W . 1980 Cell 20: 807–816
Shalloway D, Coussens PM, Yaciuk P . 1984 Proc. Natl. Acad. Sci. USA 81: 7071–7105
Skotzko M, Wu L, Anderson WF, Gordon EM, Hall FL . 1995 Cancer Res. 55: 5493–5498
Studler JM, Glowinski J, Levi-Strauss M . 1993 Eur. J. Neurosci 5: 614–623
Takekura N, Yasui W, Yoshida K, Tsujino T, Nakayama H, Kameda T, Yokozaki H, Nishimura Y, Ito H, Tahara E . 1990 Int. J. Cancer 45: 847–851
Takeya T, Hanafusa H . 1982 J. Virol. 44: 12–18
Talamonti MS, Roh MS, Curley SA, Gallick GE . 1993 J. Clin. Invest. 91: 53–60
Taylor SJ, Shalloway D . 1996 Bioessays 18: 9–11
Thomas SM, Brugge JS . 1997 Annu. Rev. Cell Dev. Biol. 13: 513–609
Wickus G, Gruenstein E, Robbins PW, Rich A . 1975 Proc. Natl. Acad. Sci. USA 72: 746–749
Yamashita A, Hakura A, Inoue H . 1999 Oncogene 18: 4777–4787
Yano M, Naito Z, Yokoyama M, Shiraki Y, Ishiwata T, Inokuchi M, Asano G . 1999 Cancer Lett. 22: 45–51
Yeung KY, Haynor DR, Ruzzo WL . 2001 Bioinformatics 17: 309–318
Zuber J, Tchernitsa OI, Hinzmann B, Schmitz AC, Grips M, Hellriegel M, Sers C, Rosenthal A, Schafer R . 2000 Nat. Genet. 24: 144–152
Acknowledgements
We would like to thank Terry Utterback and the TIGR Sequencing Core Facility for their excellent sequencing support and Michael Heaney for database support. We would also like to acknowledge the exceptional efforts of the TIGR Gene Indices Group as well as the TMEV software developers. We would like to thank Dr Alan Kraker for Pfizer Global Research for generously providing PG180970. This study was supported in part by a National Heart Lung and Blood Institute Grant (HL59781, NH Lee), the National Cancer Institute (CA85052-01-A1 and CA85429-01, TJ Yeatman; CA77049-02 and 6120-119-L0-A, J Quackenbush), the American Cancer Society (ACSRPG99-099-01-MGO, TJ Yeatman).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Malek, R., Irby, R., Guo, Q. et al. Identification of Src transformation fingerprint in human colon cancer. Oncogene 21, 7256–7265 (2002). https://doi.org/10.1038/sj.onc.1205900
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205900
Keywords
This article is cited by
-
Tumor protein D52 is upregulated in oral squamous carcinoma cells under hypoxia in a hypoxia-inducible-factor-independent manner and is involved in cell death resistance
Cell & Bioscience (2021)
-
Characterization of the Src-regulated kinome identifies SGK1 as a key mediator of Src-induced transformation
Nature Communications (2019)
-
Role of Non Receptor Tyrosine Kinases in Hematological Malignances and its Targeting by Natural Products
Molecular Cancer (2018)
-
Adhesion-mediated cytoskeletal remodeling is controlled by the direct scaffolding of Src from FAK complexes to lipid rafts by SSeCKS/AKAP12
Oncogene (2013)
-
Tumor protein D54 is a negative regulator of extracellular matrix-dependent migration and attachment in oral squamous cell carcinoma-derived cell lines
Cellular Oncology (2013)