Abstract
We reconstituted a three-dimensional gastric carcinoma model similar to invasive gastric carcinoma tissue. This model consists of a human gastric carcinoma cell line, GCTM-1, a human fibroblast cell line, TIG-1-20, and transforming growth factor-β (TGF-β)-containing type I collagen gel. Using this model, we were able to observe the growth of the two cell types, especially carcinoma cell invasive growth, in real time for more than 30 days. TGF-β and TIG-1-20 were essential for GCTM-1 invasive growth and proliferation, respectively. TGF-β induced the enhanced expression of matrix metalloproteinase 9 (MMP9) and urokinase-type plasminogen activator (uPA) in GCTM-1 at both the protein and enzymatic activity levels. The TGF-β-induced invasion of GCTM-1 was inhibited by MMP9- or uPA-antisense (AS) oligonucleotide transfection to GCTM-1. When exogenous interferon-γ (IFN-γ) was added to this model, TGF-β-dependent GCTM-1 invasion was significantly inhibited, concomitant with the decreased expression of MMP9 and uPA. The intracellular signal transduction of Smad was examined to analyse the mechanism of the inhibitory effect of IFN-γ. TGF-β accelerated the phosphorylation of Smad2/3 and nuclear translocation of the Smad2/3–Smad4 complex in GCTM-1, but these TGF-β-induced effects were significantly inhibited by IFN-γ-induced Smad7 expression. When GCTM-1 was cotransfected with AS oligonucleotide of Smad2 and Smad3, the TGF-β-induced invasion of GCTM-1 disappeared. In addition, the inhibitory effect of IFN-γ on TGF-β-dependent GCTM-1 invasion vanished by the AS oligonucleotide of Smad7 transfection. These results indicate that IFN-γ inhibits TGF-β-dependent GCTM-1 invasion through cross-talk in the Smad pathway. IFN-γ may be a new therapeutic tool for TGF-β-expressed invasive carcinomas.
Similar content being viewed by others
Introduction
Carcinoma tissue consists essentially of carcinoma cells, stromal cells, and the extracellular matrix. Stromal cells are an integral part of carcinoma tissue and account for some of its most destructive features (Ellis et al., 1994; Kunz-Schughart et al., 2001; Matrisian et al., 2001). Fibroblasts, vascular endothelial cells, macrophages, and other cells organize the tumor microenvironment. Of the various stromal cells, fibroblasts play an essential role in producing the growth factors and the extracellular matrix, which may promote the proliferation and invasion of carcinoma cells (Yashiro et al., 1996; Kunz-Schughart et al., 2001). The extracellular matrix is another important component forming the tumor microenvironment, providing the structure, generating biological signals, storing factors that generate biological signals, and exerting mechanical influence on both the carcinoma and stromal cells (Matrisian et al., 2001). These facts motivated the establishment of an in vitro organotypic culture system for an in-depth understanding of complicated carcinoma tissue and the forward development of valuable therapeutic strategies.
Various growth factors that effect carcinoma invasion are produced by both the carcinoma and stromal cells in carcinoma tissue (Ura et al., 1991; Yashiro et al., 1996; Kunz-Schughart et al., 2001). It is generally thought that the transforming growth factor-β (TGF-β) plays an important role in the aggression of gastric carcinoma, especially in promoting the highly invasive nature of carcinoma cells (Nakamura et al., 1998; Ellenrieder et al., 2001; Pasche, 2001). In fact, we and other investigators have shown that TGF-β is an indicator of a poor prognosis in cases of advanced gastric carcinoma (Nakamura et al., 1998; Saito et al., 1999). However, data concerning the role of TGF-β in carcinoma cell invasion are mainly derived from in vitro monolayer culture systems (Festuccia et al., 2000; Ellenrieder et al., 2001). These data indicate that TGF-β promotes the invasion of carcinoma cells through the proliferation of fibroblasts (Ura et al., 1991), enhanced production of the extracellular matrix from fibroblasts (Ignotz and Massague, 1986; Mahara et al., 1994), and the increased production of invasion-related factors such as urokinase-type plasminogen activator (uPA) and matrix metalloproteinase-9 (MMP9) from carcinoma cells (Festuccia et al., 2000; Ellenrieder et al., 2001).
Interferon-γ (IFN-γ) is a cytokine with pleiotropic effects on various types of cells (David, 1995; Shyu et al., 2000). Although the mechanisms of pleiotropic effects of IFN-γ are not fully understood, one of interest is the interaction with TGF-β signals (Ulloa et al., 1999; Gurujeyalakshmi et al., 1995; Tredget et al., 2000). It has been shown that IFN-γ inhibits TGF-β-induced phosphorylation of Smad2/3, required for the nuclear translocation of the Smad2/3–Smad4 complex, through the induction of antagonistic Smad7 (Ulloa et al., 1999). It has been shown that IFN-γ can suppress both fibroblast proliferation and collagen synthesis induced by TGF-β (Gurujeyalakshmi et al., 2000; Tredget et al., 2000). On the basis of these new findings, we hypothesized that IFN-γ inhibits TGF-β-dependent invasion of carcinoma cells through integration of IFN-γ and TGF-β signals.
We reconstituted an in vitro preclinical invasive gastric carcinoma model in which we were able to observe morphological changes of both carcinoma cells and fibroblasts in real time for more than 30 days. Using this model, we were able to show that exogenous IFN-γ suppresses TGF-β-dependent carcinoma cell invasion through a possible integration of TFG-β and IFN-γ signals, which could be considered as cross-talk in the Smad pathway. A possible IFN-γ therapy for TGF-β-related invasive gastric carcinoma is also suggested by our results.
Results
Effects of growth factors on GCTM-1 invasion
When GCTM-1 cells were cocultured with TIG-1-20, the GCTM-1 colonies grew slowly but were rounded, suggesting the expansive growth of GCTM-1 (Figure 1a). To establish a three-dimensional model similar to aggressive gastric carcinoma tissue, various growth factors, such as TGF-α, TGF-β, and the basic fibroblast growth factor (b-FGF), were included in the culture at several concentrations. In this study, we considered that the greatest diameter of a tumor colony reflected the invasiveness of GCTM-1. Our preliminary studies identified two significant concentrations (0.2 and 2.0 ng/ml) of growth factors. TGF-α did not affect GCTM-1 invasion, whereas both TGF-β and b-FGF enhanced it (Figure 1a and b). GCTM-1 invasion was significantly strengthened even at a low dose of TGF-β. However, TGF-β (2.0 ng/ml) did not affect the cell numbers of GCTM-1 in three-dimensional culture (control, 1.0±0.11 × 105 cells/sample; TGF-β, 1.1±0.14 × 105 cells/sample) on day 10. The invasiveness of GCTM-1 strengthened by these growth factors in a three-dimensional culture showed a similar pattern to the results of the Matrigel invasion assay (data not shown). We therefore focused on the effect of TGF-β on the invasiveness of GCTM-1.
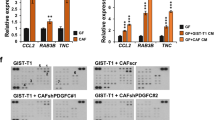
Effects of various growth factors on tumor invasion. (a) Morphology of GCTM-1-colonies in control and each growth factor-added cultures after 10 days incubation. Morphological features of GCTM-1 seen with a phase-contrast microscopy are shown. The bars in the photographs represent the length of the greatest diameter of a colony. The shape of the colonies in the control and TGF-α-added culture is round. However, colonies of GCTM-1 became irregular in shape in TGF-β- and b-FGF-added culture. (b) Effects of TGF-α, TGF-β, or b-FGF (0.2 or 2.0 ng/ml) on tumor invasion. The invasion index is shown. Each bar is identified by the growth factor and dose. Lines connecting bars identify compared indices for the t-test and are labeled by the P-value of the comparison
When TGF-β was added to the three-dimensional model, GCTM-1 colonies showed invasive characteristics within 5 days after the initial culture and continued to grow invasively until day 20. In paraffin-embedded sections on day 14, GCTM-1 showed expansive tubular growth and a signet ring-like appearance was not observed without TGF-β (Figure 2a). Under the existence of TGF-β, GCTM-1 showed invasive growth without tubular formation and some cells had a signet ring-like appearance (Figure 2b) similar to primary carcinoma tissue (Figure 2c).
Optical images of the model. The model was fixed at day 14 and 3 μm sections were made. Sections were stained with hematoxylin. (a) Control culture without TGF-β. (b) TGF-β-added culture. Some GCTM-1 cells have a signet ring appearance (arrow). (c) Primary gastric carcinoma tissue of GCTM-1. Primary carcinoma tissue was similar to our model. (b). Signet ring-like tumor cells were observed in this primary tissue (arrow)
Effects of TGF-β on the expression of MMP9 and uPA
The molecular mechanism of TGF-β-induced GCTM-1 invasion was analysed by examining whether TGF-β induces the production of invasion-related molecules, such as MMP9 and uPA. Immunohistochemical analysis revealed that TGF-β promoted the expression of uPA and MMP9 in GCTM-1 (Figure 3). This result suggests that TGF-β promotes GCTM-1 invasion through the upregulation of MMP9 and uPA.
Inhibitory effect of IFN-γ on TGF-β-induced GCTM-1 invasion
Since the integration of TGF-β and IFN-γ signals has been suggested by several investigators, we examined the effects of IFN-γ on TGF-β-induced GCTM-1 invasion in our model. The representative GCTM-1 colonies formed in different culture conditions on day 10 are shown in Figure 4a. In a complete culture medium without TGF-β and IFN-γ (control), GCTM-1 cells formed round colonies consisting of 3–20 cells that grew expansively. When TGF-β was added to the culture, the GCTM-1 colonies became irregular in shape, suggesting invasive growth. However, the numbers of cells forming a colony did not differ significantly different from the control numbers as described already. These results indicate again that TGF-β increases the invasive ability of GCTM-1 cells, but not their proliferation. When IFN-γ (100 IU/ml) alone was added, no significant effect on the morphology and the size of the GCTM-1 colony was found, but IFN-γ inhibited TGF-β-induced GCTM-1 invasion. The effect of IFN-γ and TGF-β on the invasive growth of GCTM-1 was quantified as an invasion index. TGF-β significantly increased the invasion index (P<0.001). Although INF-γ alone had no significant effect on the invasion index of the control culture, it prevented the TGF-β-dependent increase (P<0.001) (Figure 4b).
Inhibitory effects of IFN-γ on tumor invasion. (a) Morphology of GCTM-1 colonies after 10 days incubation. Phase-contrast images of GCTM-1 are shown. The colonies in both the control and IFN-γ-added cultures are round. However, colonies of GCTM-1 are irregular in shape, and are degenerating around the collagen in TGF-β-added culture. IFN-γ also inhibited TGF-β-induced GCTM-1 invasion. The bars in photographs represent the length of the greatest diameter of a colony in each culture. (b) Evaluation of tumor invasion. The invasion index shows a significant difference between the control culture and TGF-β-treated culture (P<0.001), and IFN-γ completely abolished the increase of the invasion index (P<0.001) by TGF-β
Inhibitory effects of IFN-γ on TGF-β-induced MMP9 and uPA expression
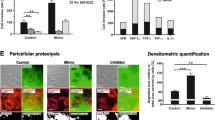
Since IFN-γ completely inhibited TGF-β-induced GCTM-1 invasion, we speculated that IFN-γ would inhibit TGF-β-induced MMP9 and uPA expression. The expression of MMP9 (Figure 5a) was determined by gelatin zymography and Western blot analysis. As regards uPA, the expression was analysed with casein-uPA zymography and uPA activity assay (Figure 5b). Three-dimensional culture gels were cultured in a serum-free culture medium with TGF-β (2.0 ng/ml) and/or IFN-γ (100 IU/ml) for 48 h. Then the proteolytic activities and production of these enzymes in the culture supernatants were measured. TGF-β significantly increased MMP9 and uPA at both the protein and enzymatic activity levels, and IFN-γ inhibited this TGF-β-induced production and the activation of these proteolytic enzymes.
Inhibitory effects of IFN-γ on TGF-β-induced MMP9 and uPA expression. (a) Gelatin zymography (upper panel) and Western blot analysis (lower panel) of MMP9 and (b) casein-uPA zymography (upper panel) and uPA activity assay (lower panel). (a) The 92 kDa bands represent enzymatic activity (upper panel) and protein level (lower panel) of MMP9. The densities of bands of Western blot analysis were quantified by NIH imaging densitometry. The expression of MMP9 was 1.0 (control), 1.7 (TGF-β), 0.8 (IFN-γ), and 0.9 (TGF-β/IFN-γ). (b) Enzymatic activity (upper panel) of uPA (54 kDa) and uPA activity (lower panel) are shown. TGF-β significantly increased uPA activity, and IFN-γ inhibited this TGF-β-upregulated uPA activity
Cross-talk between TGF-β and IFN-γ signals
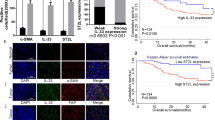
It is generally accepted that the main signal pathway of TGF-β is the Smad pathway. Therefore, we examined immunohistochemically the effects of IFN-γ on the TGF-β-related Smad pathway (Figure 6). The phosphorylation of Smad2/3 and nuclear translocation of the p-Smad2/3–Smad4 complex of GCTM-1 is not detectable in the control culture, but TGF-β accelerated both phosphorylation and nuclear translocation. IFN-γ inhibited the phosphorylation of Smad2/3 and nuclear translocation of Smad complex in the presence of TGF-β. The expression of Smad7, which inhibits Smad2/3 phosphorylation and Smad-complex nuclear translocation, was also examined. IFN-γ induced the cytoplasmic expression of Smad7 in GCTM-1 in both the absence and presence of TGF-β. These findings suggest that phosphorylation and p-Smad2/3–Smad4 complex nuclear translocation, induced by TGF-β, may be inhibited by the overexpression of Smad7 induced by IFN-γ.
Invasion assays with antisense oligonucleotide-transfected tumor cells
To verify the effects of MMP9 and uPA on the invasiveness of GCTM-1 and the inhibitory effect of IFN-γ on TGF-β-induced invasiveness through Smad7 expression, invasion assays were performed using MMP9-, uPA-, Smad2-, Smad3-, or Smad7-antisense (AS) oligonucleotide-transfected GCTM-1 (Figure 7a). TGF-β did not increase the invasive ability of MMP9-, or uPA-AS oligonucleotide-transfected GCTM-1. The cotransfection of Smad2- and Smad3-AS oligonucleotide significantly inhibited TGF-β-induced invasion of GCTM-1. On the other hand, IFN-γ did not inhibit the TGF-β-induced invasiveness of Smad7-AS-transfected GCTM-1. A Matrigel invasion assay with AS-transfected GCTM-1 showed similar results (Figure 7b).
Invasion assays with MMP9-, uPA-, Smad2-, Smad3-, or Smad7-AS-transfected GCTM-1. (a) Phase-contrast images of a three-dimensional culture. GCTM-1 were transfected scramble-, MMP9-, or uPA-AS oligonucleotide and cultured in three-dimensional condition in the presence of TGF-β (2 ng/ml). MMP9- or uPA-antisense transfection inhibited TGF-β-induced GCTM-1 invasion. Smad2- or Smad3-AS oligonucleotide transfection did not significantly inhibit TGF-β-induced GCTM-1 invasion, but their cotransfection inhibited it. Smad7-AS-transfected GCTM-1 were cultured in a three-dimensional condition with TGF-β (2 ng/ml) and IFN-γ (100 IU/ml). Smad7-AS transfection restored TGF-β-dependent GCTM-1 invasion, (b) Result of the Matrigel invasion assay. The Matrigel invasion assay was performed with AS-transfected GCTM-1. Results of the Matrigel invasion assay were corresponding with that in the three-dimensional assay. #P<0.001; ##P<0.003; *no significant difference compared with the control culture
Discussion
We developed a three-dimensional collagen gel matrix culture model of invasive gastric carcinoma tissue. We were able to observe morphological changes of both GCTM-land TIG-1-20 in real time for more than 30 days with a phase-contrast microscopy. Using this model, we indicated that IFN-γ inhibits TGF-β-induced GCTM-1 invasion through cross-talk in the Smad pathway.
Several types of three-dimensional collagen gel matrix cultures have been developed (Ura et al., 1991; Kikuchi et al., 2000; Rossi et al., 2000; Kunz-Schughart et al., 2001) to understand the complicated pathophysiology of carcinoma tissue. Most consist of carcinoma cells, fibroblasts, and collagen gel. It is usually impossible to observe cells in real time with phase-contrast microscopy in those models (Ura et al., 1991; Kikuchi et al., 2000; Meier et al., 2000; Rossi et al., 2000). We sought a suitable cell density of GCTM-1 and TIG-1-20 for long-term observation in collagen gel. When GCTM-1 and TIG-1-20 were adjusted to 1.0 × 106 and 2.0 × 106 cells/ml, respectively, both types of cell proliferated slowly, and their growths could be observed for more than 30 days. When the TIG-1-20 density was less than 1.0 × 106 cells/ml, GCTM-1 showed rapid apoptosis within 5 days after the initial culture, substantiating that TIG-1-20 produced growth factors, as reported previously (Ankrapp and Bevan, 1993; Hlatky et al., 1994; Hofland et al., 1995). Actually, TGF-β induced the expression of the hepatocyte growth factor and vascular endothelial growth factor mRNA in TIG-1-20 (data not shown).
GCTM-1 cells were established from ascites of a patient with poorly differentiated adenocarcinoma. Although carcinoma cells show highly invasive features in the original carcinoma tissue, in which carcinoma cells were positive for TGF-β, GCTM-1 grew in an expansive manner in our modified conventional model in which no exogenous TGF-β exists. To convert the expanding growth pattern of GCTM-1 in to an invasive growth pattern, several kinds of growth factors were added to our original model. TGF-β induced the most aggressive invasion pattern. In addition, signet ring-like GCTM-1 cells, similar to the original carcinoma tissue cells, appeared in the TGF-β-added model (Figure 2b). TGF-β inhibits cellular growth in normal epithelial cells (Shipley et al., 1986; Coffey et al., 1988; Moses et al., 1990; Alexandrow and Moses, 1995), and in early tumor stages the transformed epithelial cells are usually sensitive to TGF-β-mediated growth inhibition (Markowitz and Roberts, 1996; Ellenrieder et al., 2001). In later stages of tumorigenesis, however, epithelial tumor cells frequently escape from TGF-β-induced growth control; and once escape has occurred, TGF-β can act as a promoter of tumor progression (Torre-Amione et al., 1990; Welch et al., 1990). Our three-dimensional model showed that TGF-β had no effect on tumor cell proliferation. On the contrary, TGF-β increased GCTM-1 invasion with upregulation of the expressions of matrix proteinase enzymes (Figures 3 and 5). The results of invasion assays using MMP9- or uPA-AS-transfected GCTM-1 suggested that the TGF-β-dependent GCTM-1 invasion is mainly due to the enhanced expression of proteolytic enzymes such as MMP9 and uPA (Figure 7). These findings are similar to those of recent reports in which TGF-β upregulates the tumor cell expression of uPA, uPA receptor, and MMP1, 2, 7, and 9 in a monolayer culture system (Keski-Oja et al., 1988; Overall et al., 1989; Liotta et al., 1991; Arnoletti et al., 1995; Lund et al., 1995; Teti et al., 1997; Ellenrieder et al., 2001).
The TGF-β receptors are transmembrane serine/threonine kinases that propagate signals downstream (Chen et al., 1999; Itoh et al., 2000; Mori et al., 2000). Smads associate with activated TGF-β receptors and play a crucial role in TGF-β signal transduction. Smad2 and Smad3 are direct substrates of the TGF-β receptor kinase, and they interact with the common partner Smad4 (Giehl et al., 2000; Itoh et al., 2000; Inagaki et al., 2001). Smad4-containing heterometric Smad complexes then translocate from the cytoplasm into the nucleus where they function as transcriptional regulators (Fink et al., 2001; Montgomery et al., 2001). Transcriptional activation by Smad-complex requires coactivators p300 and cAMP response element-binding protein (CBP) (Attisano and Lee-Hoeflich, 2001; Ghosh et al., 2001). In contrast to receptor-activated Smads, Smad7 binds stably to TGF-β receptors and interferes with ligand-induced phosphorylation of Smad2 and Smad3 (Hayashi et al., 1997; Nakao et al., 1997). It has been shown that IFN-γ inhibits the TGF-β signaling pathway through the induction of Smad7 or competition between Smads and STAT1 for limited amounts of the shared cellular coactivators p300/CBP (Ghosh et al., 2001; Ma et al., 2001). In our experiments, IFN-γ inhibited the effects of TGF-β-dependent GCTM-1 invasion (Figure 4). In addition, our results show the possibility that IFN-γ inhibits the phosphorylation of Smad2/3 and the nuclear translocation of Smad-complex accelerated by TGF-β through the upregulation of Smad7 expression (Figure 6). Since phosphorylation and nuclear translocation of Smads are inhibited by IFN-γ, inhibition of the TGF-β signaling pathway by IFN-γ may be due to the induction of Smad7 rather than p300/CBP competition. However, we cannot completely rule out the possibility of interactions with some other signaling pathways of TGF-β, such as phosphatidylinositol 3-kinase (Bakin et al., 2000), p38 mitogen-activated protein kinase (Leivonen et al., 2002; Yu et al., 2002), extracellular signal-regulated kinase (Funaba et al., 2002; Sowa et al., 2002), and c-Jun N-terminal kinase pathway (Verrecchia et al., 2003). Furthermore, the expression of Smad7, which is stimulated by IFN-γ, induces Smad7–Smurf1 (Ebisawa et al., 2001) or Smad7–Smurf2 (Kavsak et al., 2000) complex formation and increases TGF-β receptor turnover. Our result of invasion assays using Smad2-, Smad3-, and Smad7-AS-transfected GCTM-1 (Figure 7) supported the fact that the main pathway of strengthened invasiveness in GCTM-1 is the Smad pathway.
Although the precise mechanisms of IFN-γ inhibition of TGF-β-dependent GCTM-1 invasion are still unclear, IFN-γ may be therapeutically useful for patients with TGF-β-related invasive gastric carcinoma. In addition, our three-dimensional invasive gastric carcinoma model should prove valuable in further study of cell–cell and cell–matrix interactions in complicated carcinoma tissue.
Materials and methods
Cells and reagents
TIG-1-20 (JCRB0501), a human embryonic pulmonary fibroblast cell line, was obtained from Human Science Research and Resource Bank (Osaka, Japan). GCTM-1, a human poorly differentiated gastric carcinoma cell line, was obtained from malignant ascites of a patient with relapsed scirrhous gastric cancer and were established in our laboratory (Morisaki et al., 2000). The cells were grown in tissue culture flasks in DMEM (Gibco BRL, Grand Island, NY, USA) for TIG-1-20 and RPMI-1640 (Gibco BRL) for GCTM-1. Both culture media were supplemented with 10% fetal calf serum (FCS; Gibco BRL), 100 U/ml of penicillin, and 100 μg/ml of streptomycin. Recombinant human TGF-β1 (Genzyme-Techne, Mpls, MN, USA), TGF-α (Pepro Tech, London, England), and b-FGF (Pepro Tech) were purchased. Recombinant IFN-γ was provided by Shionogi Pharmaceutical Company (Osaka, Japan). Aqueous type I collagen of cow tendon was purchased from Koken (Tokyo, Japan).
Three-dimensional culture model for long-term observation
TIG-1-20 and GCTM-1 were incorporated at various cell densities in type I collagen gel and cultured in RPMI-1640 medium with 10% FCS (complete culture medium). The cellular growth was continuously observed with a phase-contrast microscopy. For long-term observation, a suitable cell density in the collagen gel was 2.0 × 106 cells/ml for TIG-1-20 and 1.0 × 106 cells/ml for GCTM-1. Cells suspended in RPMI-1640 medium with 20% FCS were mixed with the same volume of chilled type I collagen. Each mixture was dispensed into 48 wells at 300 μl/well and allowed to gel for approximately 30 min at 37°C before the addition of a complete culture medium. After the complete culture medium was added, the three-dimensional models were incubated at 37°C in a humidified atmosphere of 5% CO2 in air. The complete culture medium was renewed every 2 days. A schematic and the macroscopic features of the three-dimensional model are shown in Figure 8a. In this model, morphological observations of both TIG-1-20 and GCTM-1 were possible in real time for more than 30 days.
Schematic and macroscopic features of the three-dimensional gastric carcinoma model. (a) Gastric carcinoma model for long-term observation by phase-contrast microscopy. This model consists of GCTM-1, TIG-1-20, and type I collagen. The final density of GCTM-1 is 1.0 × 106 cells/ml, and of TIG-1-20 is 2.0 × 106 cells/ml in collagen gel. Culture supernatants without serum were harvested for Western blot analysis and zymography. (b) Model for immunohistochemistry. A high density of cells (5.0 × 106 cells/ml of GCTM-1 and 1.0 × 107 cells/ml of TIG-1-20) in collagen gel is required to obtain this model. The high density of TIG-1-20 caused collagen gel contraction. The model block was fixed with buffered formalin, and immunohistochemical analysis was performed
Assessment of tumor invasion in three dimensions
At 10 days after the initial incubation, three sections of each culture well were photographed (× 100) at random with a digital camera (NIKON, COOL PIX, Tokyo, Japan) connected to a phase-contrast microscopy (NIKON, ECLIPSE TE 300). The pictures were printed and the greatest diameter of each GCTM-1 colony was measured. Invasion ability refers to the mean value of the greatest diameters (Bell et al., 1999; Bauman et al., 1999) of at least 20 colonies. In this study, the invasion ability of GCTM-1 was expressed as a tumor invasion index as follows: invasion index=(invasion ability in a complete culture medium supplemented with growth factor/invasion ability in a complete culture medium alone) × 100.
Matrigel invasion assay
In vitro Matrigel invasion assays were performed in cell culture chambers (Becton Dickinson, Franklin Lakes, NJ, USA) containing a polyvinylpyrrolidone-free polycarbonate filter (pore size 8.0 μm) according to the method of Albini et al. (1987) with some modifications. Briefly, the upper surface of the filter was coated with basement membrane Matrigel (Collaborative Biomedical Products, Bedford, MA, USA) at a concentration of 250 μg/cm2 and air-dried overnight at room temperature. The tumor cells were suspended at a concentration of 1 × 106 cells/ml in RPMI-1640 and treated with TGF-β (2 ng/ml) and/or IFN-γ (100 IU/ml) for 12 h. After incubation, the cells were washed twice with PBS to remove the agents, cell suspension (200 μl) was added to the upper compartment, and 20% Matrigel-contained RPMI-1640 medium (400 μl) was added to the lower compartment. The cells were then incubated at 37°C in a humidified atmosphere of 5% CO2 in air for 12 h. The cells that traverse the Matrigel adhere to the opposite surface of the filter. After incubation, the filter was fixed with 100% methanol and stained with Giemsa solution, after which cells on the upper surface were completely removed by wiping with a cotton swab. Tumor cells that had migrated from the upper to the lower side of the filter were counted with a light microscope at a magnification of × 400. Tumor cell invasiveness was defined as the mean cell number of 10 microscopic fields.
Assessment of tumor proliferation in three dimensions
The tumor proliferation in three-dimensional culture was analysed by Coulter counter cell analysis and fluorescence-activated cell sorter (FACS) analysis. TIG-1-20 cells were stained by fluorescence-PKH-67 dye (Sigma, St Louis, MO, USA) prior to three-dimensional culture and according to the manufacturer's protocol. The three-dimensional culture was performed with GCTM-1 and PKH-67-labeled TIG-1-20. After a 10-day incubation, the cultures were treated with 1% collagenase (Wako, Osaka, Japan) at 37°C for 15 min. The cells were then washed with RPMI-1640 and centrifuged. The cell pellet was resuspended with 500 μl of RPMI-1640, and the total cell number was counted with a Coulter counter (Beckman Coulter, Fullerton, CA, USA). The fractions of GCTM-1 and TIG-1-20 were determined by FACS-Calibur (Becton Dickinson). The number of GCTM-1 cells and TIG-1-20 was calculated separately from the total cell number and the fraction of GCTM-1.
Three-dimensional culture model for immunohistochemical analysis
The three-dimensional culture material needs to be of higher consistency for immunohistochemical analysis than that for long-term observation. Preliminary studies showed that a suitable cell density is 1.0 × 107 cells/ml for TIG-1-20 and 5.0 × 106 cells/ml for GCTM-1 (Figure 8b). Blocks of cultures for immunohistochemical analysis were fixed with buffered formalin (10% methanol, 4% formaldehyde) and embedded into paraffin. Paraffin sections were deparaffinized, rehydrated, incubated in an antigen retrieval solution (pH 6 citrate buffer), and heated in a high-pressure steam sterilizer at 120°C for 20 min. The slides were washed in phosphate-buffered saline, and endogenous peroxidases were quenched in 3% H2O2–methanol for 30 min, followed by blocking with 10% goat serum for 30 min at room temperature. The primary antibodies against uPA, MMP9, p-Smad2/3, Smad4, and Smad7 were obtained from Santa Cruz Biotechnologies (Santa Cruz, CA, USA) and applied at 2 μg/ml. Primary antibodies were applied and incubations were performed overnight at 4°C. After washings, secondary antibodies were applied according to the specificity of the primary antibody: uPA, MMP9, and Smad7 for Histofine Simple Stain MAX PO (goat) (Nichirei, Tokyo, Japan), and p-Smad2/3 and Smad4 for EnVision Detection Reagent Peroxidase Rabbit/Mouse (Dako Cytomation, Glostrup, Denmark). After the secondary antibodies were applied, the slides were incubated for 30 min at room temperature. The reaction sites were visualized with diaminobenzidine used as the chromogen, and the nuclei were counterstained with hematoxylin.
Western blot analysis and zymography
The proteinase secretion in the supernatant from a three-dimensional culture was determined with Western blot analysis and SDS–PAGE zymography with gelatin and casein as the substrate forMMP-9 and uPA, respectively (Sehgal et al., 1996; Ellenrieder et al., 2001). Briefly, the mixtures of TIG-1-20, GCTM-1, and type I collagen with or without the growth factor were dispensed into 48 wells at 300 μl/well, and 1000 μl of fresh serum-free RPMI was added to each well. The supernatants were collected after a 48-h incubation, centrifuged at 1500 g for removal of cellular particles, and then concentrated to 50 μl with a membrane dialysis concentrator (Millipore, Bedford, MA, USA) that excludes molecules less than 30 kDa.
Western blot analysis
Concentrated supernatants (10 μl) were resolved by SDS–PAGE, and the gels were transferred onto nitrocellulose membranes (Hybond ECL: Amersham, Arlington Heights, IL, USA). The membranes were subsequently blocked with 5% powdered milk solution for 2 h at room temperature, followed by incubation with first antibodies against MMP9 (Santa Cruz Biotechnologies) at a 1 : 1000 dilution for 1 h at room temperature. The membranes were washed and then incubated with FITC-labeled anti-goat antibody (Santa Cruz Biotechnologies) at a 1 : 100 dilution for 1 h at room temperature. After the membranes were washed, detections were performed with Molecular Imager FX (BIO RAD, Hercules, CA, USA).
Gelatin and casein-uPA zymography
Concentrated supernatants (20 μl) were mixed with SDS sample buffer without prior denaturation, and were run on a 10% SDS–PAGE containing 1 mg/ml of gelatin for gelatin MMP9 zymography or 2 mg/ml of casein and 15 μg/ml of plasminogen for casein uPA zymography. After electrophoresis, the gels were washed in 2.5% Triton X-100 for 1 h to remove the SDS and then incubated for 48 h at 37°C in a renaturing buffer containing 50 mM Tris (pH 7.5), 10 mM CaCl2, 150 mM NaCl, and 0.05% NaN3 to allow digestion of the gelatin (50 mM Tris (pH 7.5), 10 mM CaCl2, to allow digestion of the casein). The gels were subsequently stained in a solution of 0.25% Coomassie brilliant blue G-250 for 30 min and destained for 1 h with acetic acid and methanol. Proteolytic activity appeared as clear bands (zones of gelatin or casein degradation) against the blue background of stained gelatin or casein.
uPA activity assay
uPA secretion in the supernatant from a three-dimensional culture was quantified with a uPA activity assay kit (Chemicom International, Temecula, CA, USA), according to the manufacturer's protocol. Briefly, the mixtures of TIG-1-20, GCTM-1, and type I collagen with or without the growth factor were dispensed into 48 wells at 300 μl/well, and 1000 μl of fresh serum-free RPMI was added to each well. Supernatants of the three-dimensional culture were collected using the same method as that described in the Western blot analysis section. In this assay, Supernatants were applied without concentration. Supernatants were applied on a 96-well plate, and then the assay buffer and substrate were added. These mixtures were incubated at 37°C for 5 h. After incubation, absorbance on a microplate reader at 405 nm was measured.
Antisense oligonucleotide treatments
The oligonucleotides used in the AS treatments were purchased from Sigma Genosys Japan (Hokkaido, Japan). The oligonucleotides were scramble oligonucleotide:
CTACGGGTAGGTGGGGAATGGGTC AS oligonucleotide for MMP 9:
CAGGGGCTGCCAGAGGCTCAT AS oligonucleotide for uPA:
CGCCTCCGGTTGTCTGGGTTCC AS oligonucleotide for Smad 2:
GAATGGCAAGATGGACGACAT AS oligonucleotide for Smad 3:
GCAGGATGGACGACAT AS oligonucleotide for Smad 7:
GATCGTTTGGTCCTGAACAT (Morrissey et al., 1999; Chen et al., 2000; Dong et al., 2000; Sandusky et al., 2002; Razandi et al., 2003). Cells were transfected using Lipofectamine Plus Reagent (Invitrogen, Carlsbad, CA, USA) following a procedure protocol. Briefly, the oligonucleotides (1 μg)–Lipofectamine complexes were added to cells with a serum-free medium and the cells were incubated for 4 h. After 4 h incubation, the same volume of 15% FCS-RPMI-1640 was added and incubated for 24 h. After suitable incubation (24 h incubation for MMP9-, uPA-, Smad7-AS oligonucleotide transfected cells and 5 days incubation for Smad2- and Smad3-AS oligonucleotide transfected cells), invasion assays were performed.
Statistical analysis
The differences between treated and untreated control samples were analysed with the Student's t-test, and P<0.05 was considered significant.
Abbreviations
- TGF:
-
transforming growth factor
- MMP:
-
matrix metalloproteinase
- uPA:
-
urokinase-type plasminogen activator
- IFN:
-
interferon
- b-FGF:
-
basic fibroblast growth factor
- AS:
-
antisense
- CBP:
-
cAMP response element-binding protein
- JAK:
-
janus kinase
- STAT:
-
signal transducer and activator of transcription
- FCS:
-
fetal calf serum
- FACS:
-
fluorescence-activated cell sorter
- SDS:
-
sodium dodecyl sulfate
- PAGE:
-
polyacrylamide gel electrophoresis
References
Albini A, Iwamoto Y, Kleinman HK, Martin GR, Aaronson SA, Kozlowski JM and McEwan RN . (1987). Cancer Res., 47, 3239–3245.
Alexandrow MG and Moses HL . (1995). Cancer Res., 55, 1452–1457.
Ankrapp DP and Bevan DR . (1993). Cancer Res., 53, 3399–3404.
Arnoletti JP, Albo D, Granick MS, Solomon MP, Castiglioni A, Rothman VL and Tuszynski GP . (1995). Cancer, 76, 998–1005.
Attisano L and Lee-Hoeflich ST . (2001). Genome Biol., 2, 3010.1–3010.8..
Bakin AV, Tomlinson AK, Bhowmick NA, Moses HL and Arteaga CL . (2000). J. Biol. Chem., 275, 36803–36810.
Bauman GS, Fisher BJ, Mcdonald W, Amberger VR, Moore E and Del Maestro RF . (1999). Int. J. Dev. Neurosci., 17, 643–651.
Bell HS, Wharton SB, Leaver HA and Whittle IR . (1999). J. Neurosurg., 91, 989–996.
Chen SJ, Yuan W, Lo S, Trojanowska M and Varga J . (2000). J. Cell. Physiol., 183, 381–392.
Chen SJ, Yuan W, Mori Y, Levenson A, Trojanowska M and Varga J . (1999). J. Invest. Dermatol., 112, 49–57.
Coffey Jr RJ, Sipes NJ, Bascom CC, Graves-Deal R, Pennington CY, Weissman BE and Moses HL . (1988). Cancer Res., 48, 1596–1602.
David M . (1995). Pharmacol. Ther., 65, 149–161.
Dong C, Li Z, Alvarez Jr R, Feng XH and Goldschmidt-Clermont PJ . (2000). Mol. Cell, 5, 27–34.
Ebisawa T, Fukuchi M, Murakami G, Chiba T, Tanaka K, Imamura T and Miyazono K . (2001). J. Biol. Chem., 276, 12477–12480.
Ellenrieder V, Hendler SF, Ruhland C, Boeck W, Adler G and Gress TM . (2001). Int. J. Cancer, 93, 204–211.
Ellis MJ, Singer C, Hornby A, Rasmussen A and Cullen KJ . (1994). Breast Cancer Res. Treat., 31, 249–261.
Festuccia C, Angelucci A, Gravina GL, Villanova I, Teti A, Albini A, Bologna N and Abini A . (2000). Int. J. Cancer, 85, 407–415.
Fink SP, Swinler SE, Lutterbaugh JD, Massague J, Thiagalingam S, Kinzler KW, Vpgelstein B, Willson JKV and Markowitz S . (2001). Cancer Res., 61, 256–260.
Funaba M, Zimmerman CM and Mathews LS . (2002). J. Biol. Chem., 277, 41361–41368.
Ghosh AK, Yuan W, Mori Y, Chen SJ and Varga J . (2001). J. Biol. Chem., 276, 11041–11048.
Giehl K, Seidel B, Gierschik P, Adler G and Menke A . (2000). Oncogene, 19, 4531–4541.
Gurujeyalakshmi G and Giri SN . (1995). Exp. Lung Res., 21, 791–808.
Hayashi H, Abdolla, S, Qiu Y, Cai J, Xu YY, Grinnell BW, Richardson MA, Topper JN, Gimbrone Jr MA, Wrana JL and Falb D . (1997). Cell, 89, 1165–1173.
Hlatky L, Tsionou C, Hahnfeldt P and Coleman CN . (1994). Cancer Res., 54, 6083–6086.
Hofland LJ, van der Burg B, van Eijck CH, Sprij DM, van Koetsveld PM and Lamberts SW . (1995). Int. J. Cancer, 60, 93–99.
Ignotz RA and Massague J . (1986). J. Biol. Chem., 261, 4337–4345.
Inagaki Y, Mamura M, Kanamaru Y, Greenwel P, Nemoto T, Takehara K, ten Dijke P and Nakao A . (2001). J. Cell Physiol., 187, 117–123.
Itoh S, Itoh F, Goumans MJ and ten Dijke P . (2000). Eur. J. Biochem., 267, 6954–6967.
Kavsak P, Rasmussen RK, Causing CG, Bonni S, Zhu H, Thomsen GH and Wrana JL . (2000). Mol. Cell, 6, 1365–1375.
Keski-Oja J, Blasi F, Leof EB and Moses HL . (1988). J. Cell Biol., 106, 451–459.
Kikuchi S, Sato T, Arai Y, Tsutsumi O, Hiki Y and Kakita A . (2000). Hepatogastroenterology, 47, 1256–1259.
Kunz-Schughart LA, Heyder P, Schroeder J and Knuechel R . (2001). Exp. Cell Res., 266, 74–86.
Leivonen SK, Chantry A, Hakkinen L, Han J and Kahari VM . (2002). J. Biol. Chem., 277, 46338–46346.
Liotta LA, Steeg PS and Stetler-Stevenson WG . (1991). Cell, 64, 327–336.
Lund LR, Ellis V, Ronne E, Pyke C and Dano K . (1995). Biochem. J., 310, 345–352.
Ma Z, Qin H and Benveniste EN . (2001). J. Immunol., 167, 5150–5159.
Mahara K, Kato J, Terui T, Takimoto R, Horimoto M, Murakami T, Mogi Y, Watanabe N, Kohgo Y and Niitsu Y . (1994). Br. J. Cancer, 69, 777–783.
Markowitz SD and Roberts AB . (1996). Cytokine Growth Factor Rev., 7, 93–102.
Matrisian LM, Cunha GR and Mohla S . (2001). Cancer Res., 61, 3844–3846.
Meier F, Nesbit M, Hsu MY, Martin B, van Belle P, Elder DE, Schaumburg-Lever G, Garbe C, Walz TM, Donatien P, Crombleholme TM and Herlyn M . (2000). Am. J. Pathol., 156, 193–200.
Montgomery E, Goggins M, Zhou S, Argani P, Wilentz RE, Kaushal M, Booker S, Romans K, Bhargava P, Hruban RH and Kern SE . (2001). Am. J. Pathol., 158, 537–542.
Mori Y, Chen SJ and Varga J . (2000). Exp. Cell Res., 258, 374–383.
Morisaki T, Matsunaga H, Beppu K, Ihara E, Hirano K, Kanaide H, Mori M and Katano M . (2000). Anticancer Res., 20, 3363–3373.
Morrissey D, O'Connell J, Lynch D, O'Sullivan GC, Shanahan F and Collins JK . (1999). Clin. Exp., Metastasis, 17, 77–85.
Moses HL, Yang EY and Pietenpol JA . (1990). Cell, 63, 245–247.
Nakamura M, Katano M, Kuwahara A, Fujimoto K, Miyazaki K, Morisaki T and Mori M . (1998). Br. J. Cancer, 78, 1373–1378.
Nakao A, Afrakhte M, Moren A, Nakayama T, Christian JL, Heuchel R, Itoh S, Kawabata M, Heldin NE, Heldin CH and ten Dijke P . (1997). Nature, 389, 631–635.
Overall CM, Wrana JL and Sodek J . (1989). J. Biol. Chem., 264, 1860–1869.
Pasche B . (2001). J. Cell Physiol., 186, 153–168.
Razandi M, Pedram A, Park ST and Levin FR . (2003). J. Biol. Chem., 278, 2701–2712.
Rossi L, Reverberi D, Podesta G, Lastraioli S and Corvo R . (2000). Int. J. Cancer, 85, 667–673.
Saito H, Tsujitani S, Oka S, Kondo A, Ikeguchi M, Maeta M and Kaibara N . (1999). Cancer, 86, 1455–1462.
Sandusky G, Berg DT, Richardson MA, Myers L and Grinnell BW . (2002). J. Biol. Chem., 277, 49815–49819.
Sehgal I, Baley PA and Thompson TC . (1996). Cancer Res., 56, 3359–3365.
Shipley GD, Pittelkow MR, Wille Jr JJ, Scott RE and Moses HL . (1986). Cancer Res., 46, 2068–2071.
Shyu RY, Su HL, Yu JC and Jiang SY . (2000). J. Surg. Oncol., 75, 122–130.
Sowa H, Kaji H, Yamaguchi T, Sugimoto T and Chihara K . (2002). J. Biol. Chem., 277, 36024–36031.
Teti A, De Giorgi A, Spinella MT, Migliaccio S, Canipari R, Onetti Muda A and Faraggiana T . (1997). Int. J. Cancer, 72, 1013–1020.
Torre-Amione G, Beauchamp RD, Koeppen H, Park BH, Schreiber H, Moses HL and Rowley DA . (1990). Proc. Natl. Acad. Sci. USA, 87, 1486–1490.
Tredget EE, Wang R, Shen Q, Scott PG and Ghahary A . (2000). J. Interferon Cytokine Res., 20, 143–151.
Ulloa L, Doody J and Massague J . (1999). Nature, 397, 710–713.
Ura H, Obara T, Yokota K, Shibata Y, Okamura K and Namiki M . (1991). Cancer Res., 51, 3550–3554.
Verrecchia F, Tacheau C, Wangner EF and Mauviel A . (2003). J. Biol. Chem., 278, 1585–1593.
Welch DR, Fabra A and Nakajima M . (1990). Proc. Natl. Acad. Sci. USA, 87, 7678–7682.
Yashiro M, Chung YS, Kubo T, Hato F and Sowa M . (1996). Br. J. Cancer, 74, 1096–1103.
Yu L, Herert MC and Zhang YE . (2002). EMBO J., 21, 3749–3759.
Acknowledgements
We thank Kaori Nomiyama for providing expert technical assistance. This work was supported by grants for General Scientific Research (12557106) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kuga, H., Morisaki, T., Nakamura, K. et al. Interferon-γ suppresses transforming growth factor-β-induced invasion of gastric carcinoma cells through cross-talk of Smad pathway in a three-dimensional culture model. Oncogene 22, 7838–7847 (2003). https://doi.org/10.1038/sj.onc.1207046
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1207046
Keywords
This article is cited by
-
c-Kit signaling potentiates CAR T cell efficacy in solid tumors by CD28- and IL-2-independent co-stimulation
Nature Cancer (2023)
-
Regulation gene expression of miR200c and ZEB1 positively enhances effect of tumor vaccine B16F10/GPI-IL-21 on inhibition of melanoma growth and metastasis
Journal of Translational Medicine (2014)
-
Inclusive estimation of complex antigen presentation functions of monocyte-derived dendritic cells differentiated under normoxia and hypoxia conditions
Cancer Immunology, Immunotherapy (2012)
-
Nuclear factor kappaB-activated monocytes contribute to pancreatic cancer progression through the production of Shh
Cancer Immunology, Immunotherapy (2010)
-
IFN-γ down-regulates TGF-β1-induced IgA expression through Stat1 and p300 signaling
Molecules and Cells (2010)