Abstract
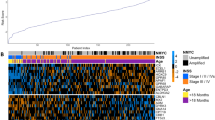
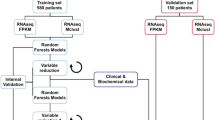
Neuroblastoma is a common childhood tumor comprising cases with rapid disease progression as well as spontaneous regression. Although numerous prognostic factors have been identified, risk evaluation in individual patients remains difficult. To define a reliable prognostic predictor and gene signatures characteristic of biological subgroups, we performed mRNA expression profiling of 68 neuroblastomas of all stages. Expression data were analysed using support vector machines (SVM-rbf), prediction analysis of microarrays (PAM), k-nearest neighbors (k-NN) algorithms and multiple decision trees. SVM-rbf performed best of all methods, and predicted recurrence of neuroblastoma with an accuracy of 85% (sensitivity 77%, specificity 94%). PAM identified a classifier of 39 genes reliably predicting outcome with an accuracy of 80%. In comparison, conventional risk stratification based on stage, age and MYCN-status only reached a predictive accuracy of 64%. Kaplan–Meier analysis using the PAM classifier indicated a 5-year survival of 20 versus 78% for patients with unfavorably versus favorably predicted neuroblastomas, respectively (P=0.0001). Significance analysis of microarrays (SAM) identified additional genes differentially expressed among subgroups. MYCN-amplification and high expression of NTRK1/TrkA demonstrated a strong association with specific gene expression patterns. Our data suggest that microarray-derived data in addition to traditional clinical factors will be useful for risk assessment and defining biological properties of neuroblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Abbreviations
- AWD:
-
alive with disease
- CR:
-
complete remission
- EFS:
-
event-free survival
- FDR:
-
false discovery rate
- k-NN:
-
k-nearest neighbors
- NED:
-
no evidence of disease
- PAM:
-
prediction analysis of microarrays
- SAM:
-
significance analysis of microarrays
- SVM-rbf:
-
support vector machines with a radial basis function kernel
- VGPR:
-
very good partial remission
References
Alaminos M, Mora J, Cheung NK, Smith A, Qin J, Chen L and Gerald WL . (2003). Cancer Res., 63, 4538–4546.
Berwanger B, Hartmann O, Bergmann E, Bernard S, Nielsen D, Krause M, Kartal A, Flynn D, Wiedemeyer R, Schwab M, Schafer H, Christiansen H and Eilers M . (2002). Cancer Cell, 2, 377–386.
Bown N, Cotterill S, Lastowska M, O'Neill S, Pearson AD, Plantaz D, Meddeb M, Danglot G, Brinkschmidt C, Christiansen H, Laureys G, Speleman F, Nicholson J, Bernheim A, Betts DR, Vandesompele J and Van Roy N . (1999). N. Engl. J. Med., 340, 1954–1961.
Brodeur GM . (2003). Nat. Rev. Cancer, 3, 203–216.
Brodeur GM, Pritchard J, Berthold F, Carlsen NL, Castel V, Castelberry RP, de Bernardi B, Evans AE, Favrot M, Hedborg H, Kaneko M, Kemshead J, Lampert F, Lee REJ, Look ATh, Pearson AD, Philip T, Roald B, Sawada T, Seeger RC, Tsuchida Y and Voute PA . (1993). J. Clin. Oncol., 11, 1466–1477.
Brodeur GM, Seeger RC, Schwab M, Varmus HE and Bishop JM . (1984). Science, 224, 1121–1124.
Caron H . (1995). Med. Pediatr. Oncol., 24, 215–221.
Chang CC and Lin CJ . (2001). Neural. Comput., 13, 2119–2147.
Draghici S, Khatri P, Bhavsar P, Shah A, Krawetz SA and Tainsky MA . (2003). Nucleic Acids Res., 31, 3775–3781.
Elbaz N, Bedecs K, Masson M, Sutren M, Strosberg AD and Nahmias C . (2000). Mol. Endocrinol., 14, 795–804.
Godfried MB, Veenstra M, v Sluis P, Boon K, v Asperen R, Hermus MC, v Schaik BD, Voute TP, Schwab M, Versteeg R and Caron HN . (2002). Oncogene, 21, 2097–2101.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD and Lander ES . (1999). Science, 286, 531–537.
Hiyama E, Hiyama K, Nishiyama M, Reynolds CP, Shay JW and Yokoyama T . (2003). J. Pediatr. Surg., 38, 1730–1734.
Hiyama E, Hiyama K, Yamaoka H, Sueda T, Reynolds CP and Yokoyama T . (2004). Pediatr. Surg. Int, 20, 33–38.
Katoh M . (2003). Int. J. Oncol., 23, 1219–1224.
Maris JM and Matthay KK . (1999). J. Clin. Oncol., 17, 2264–2279.
Matthay KK, Villablanca JG, Seeger RC, Stram DO, Harris RE, Ramsay NK, Swift P, Shimada H, Black CT, Brodeur GM, Gerbing RB and Reynolds CP . (1999). N. Engl. J. Med., 341, 1165–1173.
Ntzani EE and Ioannidis JP . (2003). Lancet, 362, 1439–1444.
Nutt CL, Mani DR, Betensky RA, Tamayo P, Cairncross JG, Ladd C, Pohl U, Hartmann C, McLaughlin ME, Batchelor TT, Black PM, von Deimling A, Pomeroy SL, Golub TR and Louis DN . (2003). Cancer Res., 63, 1602–1607.
Ohira M, Morohashi A, Inuzuka H, Shishikura T, Kawamoto T, Kageyama H, Nakamura Y, Isogai E, Takayasu H, Sakiyama S, Suzuki Y, Sugano S, Goto T, Sato S and Nakagawara A . (2003). Oncogene, 22, 5525–5536.
Ohira M, Oba S, Nakamura Y, Isogai E, Kaneko S, Nakagawa A, Hirata T, Kubo H, Goto T, Yamada S, Yoshida Y, Fuchioka M, Ishii S and Nakagawara A . (2005). Cancer Cell, 7, 337–350.
Pomeroy SL, Tamayo P, Gaasenbeek M, Sturla LM, Angelo M, McLaughlin ME, Kim JY, Goumnerova LC, Black PM, Lau C, Allen JC, Zagzag D, Olson JM, Curran T, Wetmore C, Biegel JA, Poggio T, Mukherjee S, Rifkin R, Califano A, Stolovitzky G, Louis DN, Mesirov JP, Lander ES and Golub TR . (2002). Nature, 415, 436–442.
Quinlan JR . (1993). Programs for Machine Learning. C4.5 Morgan Kaufmann: San Francicso, CA.
Rosenwald A and Staudt LM . (2002). Semin. Oncol., 29, 258–263.
Schilling FH, Spix C, Berthold F, Erttmann R, Fehse N, Hero B, Klein G, Sander J, Schwarz K, Treuner J, Zorn U and Michaelis J . (2002). N. Engl. J. Med., 346, 1047–1053.
Schoch C, Kohlmann A, Schnittger S, Brors B, Dugas M, Mergenthaler S, Kern W, Hiddemann W, Eils R and Haferlach T . (2002). Proc. Natl. Acad. Sci. USA, 99, 10008–10013.
Schulte JH, Schramm A, Pressel T, Klein-Hitpass L, Kremens B, Eils J, Havers W and Eggert A . (2003). Klin. Padiatr., 215, 298–302.
Schwab M, Westermann F, Hero B and Berthold F . (2003). Lancet Oncol., 4, 472–480.
Scott D, Elsden J, Pearson A and Lunec J . (2003). Cancer Lett., 197, 81–86.
Simon R, Radmacher MD, Dobbin K and McShane LM . (2003). J. Natl. Cancer Inst., 95, 14–18.
Su WT, Alaminos M, Mora J, Cheung NK, La Quaglia MP and Gerald WL . (2004). Cancer Genet. Cytogenet., 154, 131–137.
Teitz T, Wei T, Valentine MB, Vanin EF, Grenet J, Valentine VA, Behm FG, Look AT, Lahti JM and Kidd VJ . (2000). Nat. Med., 6, 529–535.
Thompson PM, Gotoh T, Kok M, White PS and Brodeur GM . (2003). Oncogene, 22, 1002–1011.
Tibshirani R, Hastie T, Narasimhan B and Chu G . (2002). Proc. Natl. Acad. Sci. USA, 99, 6567–6572.
Tusher VG, Tibshirani R and Chu G . (2001). Proc. Natl. Acad. Sci. USA, 98, 5116–5121.
van de Vijver MJ, He YD, van't Veer LJ, Dai H, Hart AA, Voskuil DW, Schreiber GJ, Peterse JL, Roberts C, Marton MJ, Parrish M, Atsma D, Witteveen A, Glas A, Delahaye L, van der Velde T, Bartelink H, Rodenhuis S, Rutgers ET, Friend SH and Bernards R . (2002). N. Engl. J. Med., 347, 1999–2009.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A and Speleman F . (2002). Genome Biol., 3, RESEARCH0034.1–RESEARCH0034.11.
Wei JS, Greer BT, Westermann F, Steinberg SM, Son CG, Chen QR, Whiteford CC, Bilke S, Krasnoselsky AL, Cenacchi N, Catchpoole D, Berthold F, Schwab M and Khan J . (2004). Cancer Res., 64, 6883–6891.
Witten A and Frank E . (2002). Data Mining: Practical Machine Learning Tools and Techniques with Java Implemenations. Morgan Kaufmann: San Francicso, CA.
Yamamoto K, Hanada R, Kikuchi A, Ichikawa M, Aihara T, Oguma E, Moritani T, Shimanuki Y, Tanimura M and Hayashi Y . (1998). J. Clin. Oncol., 16, 1265–1269.
Acknowledgements
We thank Frank Berthold and Thorsten Simon from the German Neuroblastoma Study Trial Office at the University Children's Hospital of Cologne for providing the clinical patient data as well as a part of the primary tumor material from the neuroblastoma tumor bank of the German competence network ‘Pediatric Oncology and Hematology’. This work was funded by a Grant from the Kind-Philipp Stiftung (to AE) and the German National Genome Network (BMBF/NGFN2) to AE and AS.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Schramm, A., Schulte, J., Klein-Hitpass, L. et al. Prediction of clinical outcome and biological characterization of neuroblastoma by expression profiling. Oncogene 24, 7902–7912 (2005). https://doi.org/10.1038/sj.onc.1208936
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208936
Keywords
This article is cited by
-
Development and validation of a 21-gene prognostic signature in neuroblastoma
Scientific Reports (2023)
-
Artificial neural network classifier predicts neuroblastoma patients’ outcome
BMC Bioinformatics (2016)
-
Hybrid Mixture Model for Subpopulation Identification
Statistics in Biosciences (2016)
-
HOXD-AS1 is a novel lncRNA encoded in HOXD cluster and a marker of neuroblastoma progression revealed via integrative analysis of noncoding transcriptome
BMC Genomics (2014)
-
Pathway activity inference for multiclass disease classification through a mathematical programming optimisation framework
BMC Bioinformatics (2014)