Abstract
As mitosis progresses, the chromosomal passenger proteins (CPPs) Survivin, Aurora B, INCENP and Borealin dynamically colocalize to mitotic structures. Chromosomal passenger proteins are already expressed during G2, whereas the nuclear envelope is only disassembled at the end of prophase. However, the mechanisms that modulate their nucleocytoplasmic localization before nuclear envelope breakdown (NEB) are poorly characterized. Using epitope-tagged proteins, we show that Aurora B, like Survivin, undergoes CRM1-mediated nucleocytoplasmic shuttling, although both proteins lack identifiable ‘classical’ nuclear transport signals. On the other hand, INCENP resides more stably in the nucleus and contains multiple nuclear localization signals. Finally, Borealin was detected in the nucleolus and the cytoplasm, but its cytoplasmic localization is not directly regulated by CRM1. Coexpression experiments indicate that the nuclear localization of Aurora B, but not of Survivin, is modulated by INCENP and that Survivin prevents the nucleolar accumulation of Borealin. Our data reveal that, in contrast to their closely related localization during mitosis, the nucleocytoplasmic localization of the CPPs before NEB is largely unrelated. Furthermore, the specific effect of INCENP and Survivin on the localization of Aurora B and Borealin, respectively, suggests that different complexes of CPPs may exist not only during mitosis, as recently reported, but also before NEB.
Similar content being viewed by others
Introduction
The chromosomal passenger proteins (CPPs) Survivin, Aurora B, INCENP and Borealin interact with each other during mitosis forming one or more protein complexes that play important roles in early and late mitotic events. Aurora B is a member of a conserved family of serine/threonine protein kinases originally identified as a mammalian homologue of Drosophila and yeast proteins required for chromosome segregation (Bischoff et al., 1998; Terada et al., 1998). Aurora B phosphorylates a variety of substrate proteins and regulates many aspects of cell division, from chromosome condensation to cytokinesis (reviewed by Andrews et al., 2003). INCENP is a large protein that has been proposed to act as a scaffold for the binding of the other passenger proteins (Bolton et al., 2002). Aurora B phosphorylates INCENP, which, in turn, activates the kinase activity of Aurora B in a positive feedback loop (Bishop and Schumacher, 2002; Honda et al., 2003). Survivin is a member of the inhibitor of apoptosis protein family that plays a dual role as regulator of both apoptosis and cell division (reviewed by Altieri, 2003a). Like INCENP, Survivin has been reported to constitute both a substrate and an activator for Aurora B (Chen et al., 2003; Wheatley et al., 2004). Borealin, which was independently isolated as DasraB, is the most recently identified and the least characterized subunit of the chromosomal passenger complex (Gassman et al., 2004; Sampath et al., 2004). Borealin is phosphorylated by Aurora B in vitro but, unlike Survivin and INCENP, does not appear to increase the activity of the kinase (Gassman et al., 2004). Several lines of evidence implicate at least two of the CPPs, Survivin and Aurora B, in the development and progression of human tumors, and these proteins hold potential as targets for novel anticancer therapies (reviewed by Altieri, 2003b; Giet et al., 2005).
The hallmark of CPPs is their dynamic pattern of subcellular localization during mitosis: they associate with the centromeric region of chromosomes from prophase to metaphase, relocate to the spindle midzone and equatorial cortex during anaphase and finally accumulate in the midbodies during cytokinesis (Adams et al., 2001a). Although their levels are maximal during mitosis, the CPPs are already expressed before the disassembly of the nuclear envelope, or nuclear envelope breakdown (NEB), takes place at the transition between prophase and prometaphase (reviewed by Burke and Ellenberg, 2002). Before NEB, the transport of proteins between the nucleus and cytoplasm occurs exclusively through the nuclear pore complex, and is usually an active process, mediated by soluble import/export receptors that bind to nuclear localization signals (NLSs) and nuclear export signals (NESs) in the cargo protein, respectively (Mattaj and Englmeier, 1998). Dynamic nucleocytoplasmic transport constitutes an important mechanism that regulates the subcellular localization and function of several cancer-associated proteins (Fabbro and Henderson, 2003).
In contrast to the detailed knowledge on the mitotic localization of CPPs, the mechanisms that contribute to regulate their subcellular distribution in the presence of an intact nuclear envelope are poorly characterized. In this respect, we have previously shown that Survivin behaves as a nuclear shuttling protein before NEB, and is actively excluded from the nucleus by a mechanism mediated by the nuclear export receptor CRM1 (Rodriguez et al., 2002). In the present study, we show that Aurora B also accesses, directly or indirectly, the CRM1-dependent pathway of nuclear export. In contrast, CRM1 does not appear to play a direct role in the nucleocytoplasmic distribution of INCENP or Borealin. Furthermore, we have functionally tested several potential nuclear transport signals in Aurora B and INCENP, and identified three independent NLSs in INCENP. Interestingly, our results suggest that, despite their closely related localization during mitosis, the subcellular localization of the CPPs before NEB is partially unrelated. In this respect, coexpression experiments showed that INCENP induces the nuclear accumulation of Aurora B, but not of Survivin. On the other hand, Survivin, through its carboxy-terminal domain, prevents the targeting of Borealin to the nucleolus. Such specific effect of INCENP and Survivin on the localization of Aurora B and Borealin, respectively, suggests that different complexes of these CPPs may exist before NEB, in line with recent evidence indicating that chromosomal passenger complexes of different composition may exist later during mitosis (Gassman et al., 2004). Our findings, in summary, begin to elucidate the molecular mechanisms that modulate the subcellular distribution of the CPPs in the presence of an intact nuclear envelope, during periods of the cell cycle at which our knowledge of the biology of these proteins is more limited.
Results
Subcellular localization of epitope-tagged chromosomal passenger proteins before nuclear envelope breakdown
We have previously shown that Survivin undergoes continuous shuttling between nucleus and cytoplasm in the presence of an intact nuclear envelope (Rodriguez et al., 2002). The information regarding the subcellular localization of the other CPPs before NEB is very limited, which contrasts with the large number of studies addressing the subcellular trafficking of these proteins later during mitosis. To investigate the mechanisms that contribute to regulate their nucleocytoplasmic distribution, we generated mammalian expression plasmids encoding full-length human Aurora B, INCENP and Borealin proteins with small VSV-epitope tags fused at their amino-terminal end, and their localization in transiently transfected MCF-7 cells was analysed by immunofluorescence using antitag antibodies (Figure 1a). As a marker of nuclear envelope integrity, green fluorescent protein (GFP)-lamin A was co-transfected with the CPPs. Consistent with our previous findings, Flag-Survivin was predominantly detected in the cytoplasm in cells where the nuclear envelope remained intact. VSV-Aurora B was detected in the nucleus and cytoplasm. Variable levels of the protein in each compartment were observed in different cells, ranging from predominantly nuclear (N>C) to mostly cytoplasmic (C>N). Similar results were obtained using Aurora B-HA, which bears a carboxy-terminal HA tag (not shown). On the other hand, VSV-INCENP showed a clear nuclear localization in most transfected cells. Finally, VSV-Borealin was detected in the nucleus, with a prominent nucleolar accumulation, and in the cytoplasm. Importantly, the three VSV-tagged CPPs readily localized to the centromeres during metaphase and the midbody during telophase/cytokinesis as expected (Figure 1a), thus confirming that the small tags do not grossly interfere with the subcellular targeting of these proteins. We determined the localization of each of the CPPs in at least 200 transfected cells. As illustrated in Figure 1b, Survivin and INCENP were consistently localized in the cytoplasm and nucleus, respectively, whereas Aurora B and Borealin showed a more heterogeneous distribution in the population of transfected cells.
Subcellular localization of epitope-tagged chromosomal passenger proteins (CPPs) before nuclear envelope breakdown. (a) Examples of the subcellular localization of epitope-tagged CPPs in transfected MCF-7 cells. Flag-Survivin is predominantly located in the cytoplasm, as previously reported. Three different patterns of VSV-Aurora B nucleocytoplasmic localization can be observed: predominantly nuclear (N>C), nuclear and cytoplasmic (N=C) and predominanlty cytoplasmic (C>N). VSV-INCEP efficiently accumulates in the nucleus. Finally, VSV-Borealin localizes into the nucleus, where it accumulates prominently in nucleoli and in the cytoplasm. Co-transfected green fluorescent protein-Lamin A was used as a marker to confirm the integrity of the nuclear envelope. VSV-tagged Aurora B, INCENP and Borealin localize to the centromeres during metaphase and to the midbody during telophase/cytokinesis. Cells were counterstained with Hoechst 33285 to visualize the DNA. (b) Quantification of the subcellular distribution of CPPs in transfected MCF-7 cells. The localization of each protein was determined using fluorescence microscopy in a single-cell basis, and classified as predominantly cytoplasmic (C>N), cytoplasmic and nuclear (C=N) or predominantly nuclear (N>C). In the case of Borealin, the localization was scored as cytoplasmic (C), cytoplasmic and nucleolar (C=Nol) or predominantly nucleolar (Nol). Bars represent the mean of two independent experiments with less than 10% variation. At least 200 cells were counted per sample.
These results indicate that despite their close related targeting during mitosis, the CPPs show different patterns of subcellular localization before NEB, suggesting that the distribution of these proteins in the presence of an intact nuclear envelope may be, at least partially, regulated in an independent manner.
Aurora B undergoes CRM1-mediated nuclear export
The nucleocytoplasmic distribution of Aurora B fused to a MYC tag, similar in size to the VSV and HA epitopes used here, has been previously shown to be predominantly nuclear in U2OS cells (Honda et al., 2003), and both nuclear and cytoplasmic in COS-7 cells (Kawajiri et al., 2003; Yasui et al., 2004). These previous observations, and the heterogeneous distribution we found in MCF-7 cells, suggested that the nucleocytoplasmic distribution of Aurora B before NEB could be a dynamic process. Given that the nucleocytoplasmic shuttling of Survivin is mediated by the CRM1 nuclear export receptor (Rodriguez et al., 2002), we tested the possibility that a similar mechanism may operate in the case of Aurora B, by evaluating the effect of the specific CRM1 inhibitor leptomycin B (LMB) on the nucleocytoplasmic distribution of epitope-tagged Aurora B. Incubation of transfected MCF-7 cells with LMB led to a relocation of VSV-Aurora B (Figure 2a) and Aurora B-HA (not shown) from the cytoplasm to the nucleus. These results indicate that, as in the case of Survivin, the nucleocytoplasmic localization of Aurora B before NEB is the result of a dynamic equilibrium between nuclear import and CRM1-dependent export. The steady-state localization and the LMB-induced nuclear relocation of a kinase dead (KD) Aurora B (the catalytically inactive mutant K106R) was comparable to that of the wild-type protein (not shown), indicating that nucleocytoplasmic transport of Aurora B does not require its kinase activity.
Nucleocytoplasmic distribution of Aurora B is regulated by CRM1-dependent nuclear export. (a) Images show an example of the re-localization of VSV-Aurora B to the nucleus of MCF-7 cells after treatment with the specific CRM1 inhibitor leptomycin B (LMB; 6 ng/ml for 6 h). Graphs show a quantification of the effect of LMB on the localization of VSV-Aurora. Bars represent the proportion of transfected MCF-7 cells showing each pattern of VSV-Aurora B nucleocytoplasmic distribution and represent the results (mean±s.d.) of four independent experiments. (b) Time-course analysis of LMB-induced nuclear relocation of Flag-Survivin and VSV-Aurora B in MCF-7 cells. Graph shows the proportion of cells showing predominantly cytoplasmic (C>N) localization of Flag-Survivin or VSV-Aurora B after treatment with 6 ng/ml LMB for the indicated time.
Next, we used a time-course analysis of cells treated with LMB to compare the nuclear transport kinetics of Aurora B and Survivin. As illustrated in Figure 2b, Flag-Survivin in untreated cells was more cytoplasmic than VSV-Aurora B and, upon LMB treatment, the nuclear accumulation of Survivin was noticeably faster than that of Aurora B. These results indicate that Survivin is more dynamic than Aurora B in terms of nuclear shuttling before NEB.
CRM1 is not a direct determinant of the nucleocytoplasmic localization of INCENP and Borealin before nuclear envelope breakdown
As CRM1-mediated export contributes, directly or indirectly, to determine the localization of Aurora B and Survivin before NEB, we tested the possibility that the same pathway may also regulate the nucleocytoplasmic distribution of INCENP and Borealin.
In the case of INCENP, its predominantly nuclear localization hampered the use of LMB to assess the role of CRM1-dependent nuclear export. As an alternative experimental approach, we coexpressed VSV-INCENP with an yellow fluorescent protein (YFP)-tagged version of the export receptor CRM1. We have used this approach previously to demonstrate the role of CRM1 in the export of the predominantly nuclear protein BARD1 (Rodriguez et al., 2004). As a positive control, we used Flag-Survivin+NLS, a modified version of Survivin fused to the strong NLS of SV-40 large T antigen (Kalderon et al., 1984). As shown in Figure 3a (left panels), the SV-40 NLS-mediated import overcomes endogenous CRM1-dependent export of Survivin and enforces its steady-state localization in the nucleus when expressed alone. However, coexpression with YFP-CRM1 efficiently relocates Flag-Survivin+NLS to the cytoplasm, revealing its nuclear shuttling ability. In contrast to Survivin+NLS, VSV-INCENP remained in the nucleus when coexpressed with YFP-CRM1 (Figure 3a, right panels).
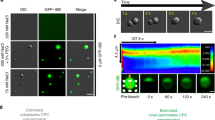
CRM1 is not a direct determinant of the nucleocytoplasmic localization of INCENP and Borealin before nuclear envelope breakdown. (a) Coexpression with the export receptor CRM1 does not induce cytoplasmic relocation of INCENP. As a positive control (left panels), a nuclear version of Survivin, tagged with the SV-40 NLS (Flag-Surv+NLS), efficiently relocalizes to the cytoplasm when coexpressed with YFP-CRM1. In contrast, the nuclear localization of VSV-INCENP (right panels) does not change when coexpressed with the export receptor. (b) Leptomycin B (LMB) treatment does not significantly reduce the partial cytoplasmic localization of VSV-Borealin. The localization of VSV-Borealin was determined by immunofluorescence in transfected MCF-7 cells treated with 6 ng/ml LMB for 6 h. In the same cells, nuclear relocation of endogenous NF-κB (p65 subunit) was evaluated as an internal control for inhibition of the CRM1-mediated nuclear export pathway. (c) Coexpression with CRM1 does not induce cytoplasmic relocation of Borealin. When expressed alone or coexpressed with YFP-CRM1, VSV-Borealin is similarly localized to the nucleoli or both the nucleoli and the cytoplasm in untreated cells (left panels). Upon incubation with 5 μg/ml actinomycin D for 3 h (right panels), the nucleolar localization of Borealin is disrupted, and the protein disperses through the nucleus, but does not relocate to the cytoplasm when coexpressed with CRM1.
On the other hand, given the partial cytoplasmic localization of Borealin, we first evaluated the effect of LMB on its nucleocytoplasmic distribution. As illustrated in Figure 3b, the levels of cytoplasmic Borealin did not significantly decrease in cells exposed to LMB for 6 h. As an internal control, nuclear accumulation of endogenous NF-κB, whose cytoplasmic localization requires active CRM1-dependent export from the nucleus (Johnson et al., 1999), confirmed the effective blockage of CRM1 in these cells. In line with the lack of LMB responsiveness, the fraction of Borealin localized in the nucleus did not undergo relocation to the cytoplasm when coexpressed with YFP-CRM1 (Figure 3c). As nuclear Borealin was largely accumulated in the nucleoli, it remained possible that trapping in this organelle could prevent its export. To test this possibility, cells were incubated in the presence of actinomycin D to disrupt nucleoli. Actinomycin D treatment resulted in a diffuse nuclear localization of VSV-Borealin. However, VSV-Borealin did not relocate to the cytoplasm in CRM1-transfected, actinomycin D-treated cells (Figure 3c).
These observations strongly suggest that CRM1-mediated export does not play a major role as a direct determinant of the nucleocytoplasmic localization of these proteins, although our results do not exclude the possibility that INCENP and Borealin may be actively exported from the nucleus under some circumstances.
Sustained treatment with leptomycin B disrupts the localization of Survivin and Aurora B in mitotic cells
It has been recently reported that, besides its role as a nuclear export receptor, CRM1 is a critical effector of Ran-GTP during mitosis (Arnaoutov et al., 2005). Thus, inhibition of CRM1 by LMB treatment was shown to disrupt mitotic progression and chromosome segregation. Given the role of CRM1 as a determinant of the nucleocytoplasmic distribution of Survivin and Aurora B before NEB, we tested the possibility that LMB could also affect the localization of these CPPs during mitosis. To this end, MCF-7 cells were incubated overnight in the presence of 6 ng/ml LMB and the localization of endogenous Survivin and Aurora B was evaluated by immunofluorescence. In those interphase cells where endogenous Survivin and Aurora B were expressed at detectable levels, their localization was predominantly nuclear both in the absence and presence of LMB (not shown). Consistent with the LMB-induced mitotic defects reported by Arnaoutov et al. in HeLa and U2OS cells, very few MCF-7 cells in anaphase could be detected after the treatment. Interestingly, the levels of endogenous Survivin (Figure 4a, upper set of panels) and Aurora B (Figure 4a, lower set of panels) in the centromeric region of chromosomes during prophase and (pro)metaphase were significantly reduced in many cells after LMB treatment. Next, the effect of shorter LMB incubation on the localization of Survivin and Aurora B was determined in synchronized mitotic cells that were released from a nocozadole arrest in the absence or presence of LMB. As a positive control, the localization of RanBP2, whose mitotic targeting is rapidly disrupted upon CRM1 inactivation (Arnaoutov et al., 2005), was determined. In contrast to that of RanBP2 (Figure 4b, upper set of panels), the localization of Survivin (Figure 4b, lower set of panels) and Aurora B (not shown) was not disrupted after 1 h in the presence of LMB. Similar results (not shown) were obtained using time-lapse microscopy analysis of HeLa cells stably transfected with GFP-Survivin. Thus, in our setting, sustained blockage of CRM1 is required to disrupt CPP targeting to mitotic structures, suggesting that the mechanism by which CRM1 activity affects the localization of these CPPs is different from the direct mechanism by which the export receptor mediates the targeting of RanBP2. We are presently carrying out experiments to further investigate the relationship between the CRM1 pathway and the localization and activity of CPPs during mitosis.
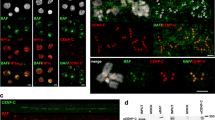
Sustained leptomycin B (LMB) treatment disrupts the localization of endogenous Survivin and Aurora B in mitotic cells. (a) Immunofluorescence microscopy analysis of the localization of endogenous Survivin (upper set of panels) and Aurora B (lower set of panels) in mitotic MCF-7 cells untreated or treated with 6 ng/ml LMB for 14 h. A clear colocalization of Survivin and Aurora B with the centromere marker CREST is consistently observed in untreated samples during prophase and metaphase. In LMB-treated samples, the colocalization of the chromosomal passenger proteins (CPPs) with CREST is severely disrupted in many prophase and metaphase cells. The larger panels in the ‘+14 h LMB’ samples show microscopy fields containing cells with correctly localized CPPs (green arrowheads) and cells with disrupted localization of Survivin or Aurora B (red arrowheads). A higher magnification of the cells inside the square is shown on the right. (b) Mitotic localization of Survivin is not altered by a short-term LMB treatment that efficiently disrupts RanBP2 targeting. Cytospins were made from nocodazole-arrested MCF-7 cells that were collected by shake-off, washed and incubated for 1 h in the absence or presence of LMB (6 ng/ml). Localization of RanBP2 to kinetochores is disrupted under these conditions (upper set of panels). A higher magnification of the cells inside the square is shown on the right. In contrast to RanBP2, Survivin remains correctly localized after short-term LMB treatment (lower set of panels).
Functional analysis of putative ‘classical’ nuclear transport signals in Aurora B and INCENP
To further characterize the mechanisms that control the nucleocytoplasmic distribution of the CPPs, we next aimed to identify amino-acid sequences in these proteins that may function as nuclear transport signals. In our previous work, we mapped the region responsible for the nuclear shuttling of Survivin to its carboxy-terminal coiled-coil domain, although we could not identify active ‘classical’ nuclear transport signals in this protein (Rodriguez et al., 2002). On the other hand, two evolutionarily conserved functional NLSs have been reported in Borealin sequence (Gassman et al., 2004). Thus, we focused our analysis on Aurora B and INCENP. By visual inspection of Aurora B primary sequence, we identified two segments of amino acids that resemble the leucine-rich NESs recognized by CRM1 (Table 1). The presence of an NES in the second of these segments was also predicted by using the NetNES software (La Cour et al., 2004). On the other hand, analysis of Aurora B sequence using the PSORT II program (Nakai and Horton, 1999) identified a stretch of basic residues as a putative ‘pat7’ NLS. We tested the ability of each of these putative transport sequences to act as autonomous NLSs or NESs. As shown in Table 1, none of the NES-like sequences in Aurora B was active in a Rev-GFP-based in vivo nuclear export assay (Henderson and Eleftheriou, 2000), and the putative NLS was not able to mediate the nuclear accumulation of YFP. Thus, none of the candidate nuclear transport signals identified in Aurora B are functional, raising the possibility that Aurora B may contain non-classical transport signals or access the pathways of nucleocytoplasmic transport in an indirect way, through binding to other molecule(s) that may bridge its interaction with transport receptors.
In the case of INCENP, two amino-acid segments resembling bipartite NLSs have been noted previously in the murine protein (Saffery et al., 1999). These sequences are conserved in the human homologue (Adams et al., 2001b), but their functionality has not been experimentally tested. INCENP segments encompassing these amino-acid sequences and several other candidate NLSs identified by PSORT II analysis were fused to YFP to evaluate their ability to mediate nuclear import in vivo. We identified three short amino-acid sequences that showed nuclear import activity (Figure 5a). As illustrated in Figure 5b, each of these amino-acid segments was able to promote the nuclear accumulation of YFP, which per se is evenly distributed between nucleus and cytoplasm. We termed these sequences NLS 1, NLS 2 (both located in the amino-terminal end) and NLS 3 (in the middle region of the protein). In addition, we noted that INCENP NLS 1 and NLS 2 led to efficient accumulation of the fused YFP in the nucleoli. Next, using YFP-tagged proteins, we observed that an N-terminal fragment of INCENP containing NLS 1, 2 and 3 localized exclusively in the nucleus (Figure 5c), whereas a C-terminal fragment distributed throughout the cell. Importantly, small interstitial deletions that eliminate the NLSs readily disrupted the nuclear and nucleolar localization of the amino-terminal fragment of INCENP. Altogether, these results indicate that human INCENP contains at least three independent functional NLSs, two of which may also function as nucleolar targeting motifs.
INCENP contains multiple functional nuclear localization signals (NLSs). (a) Amino-acid sequence of the INCENP fragments containing putative NLSs that were tested for import activity. Underlined residues indicate the sequences in human INCENP homologous to the previously proposed putative NLS in the mouse protein (Saffery et al., 1999). Each of these INCENP fragments was fused to YFP and the localization of the fusion proteins was compared to that of YFP alone (negative control) and YFP-NLS (positive control) using fluorescence microscopy. This analysis identified three small INCENP segments that induce nuclear accumulation of YFP and thus, behave as autonomous functional NLSs. (b) Representative examples of transfected MCF-7 cells showing an even distribution of YFP through the nucleus and the cytoplasm, and the nuclear accumulation of YFP induced by the fusion of INCENP NLS 1, NLS 2 and NLS 3. Note that both INCENP NLS 1 and NLS 2 efficiently targeted YFP to the nucleoli. (c) Schematic representation and subcellular localization of INCENP fragments containing or lacking NLS 1, 2 and 3. YFP-INCENP(1–454) is exclusively nuclear, whereas YFP-INCENP(455–919) is more diffusely distributed throughout the cell. Small interstitial deletions eliminating the NLSs efficiently abrogate the nuclear accumulation of YFP-INCENP(1–454).
INCENP and Survivin modulate the subcellular localization of Aurora B and Borealin, respectively, before nuclear envelope breakdown
The closely related and interdependent trafficking of the CPPs during mitosis (Speliotes et al., 2000; Wheatley et al., 2001; Bolton et al., 2002; Lens et al., 2003; Gassman et al., 2004; Sampath et al., 2004) raised the possibility that these proteins might also modulate the localization of each other before NEB. We addressed this issue using co-transfection experiments. We reasoned that concomitantly raising the levels of two CPPs would reveal any influence on each other's localization, although it must be noted that endogenous levels of the four CPPs are present in all cases. Figure 6A illustrates the localization of Aurora B, Survivin, INCENP and Borealin when coexpressed with each of the other CPPs. As a control for specificity, we used YFP-NLS, a fusion protein of YFP and the SV-40 NLS, whose coexpression should not alter the localization of the CPPs.
INCENP and Survivin modulate the subcellular localization of Aurora B and Borealin, repectively, before nuclear envelope breakdown. (A) Images show representative examples of the localization of epitope-tagged Aurora B, Survivin, INCENP and Borealin when coexpressed with each of the other chromosomal passanger proteins (CPPs) or YFP-nuclear localization signals (NLS) as a control for specificity. The tag used is indicated inside the panels. Coexpression with INCENP induced nuclear relocation of Aurora B (panel c1), but not of Survivin (panel c2). The largely cytoplasmic localization of Survivin and the nuclear localization of INCENP were not substantially altered by coexpression with any of the other CPPs. Coexpression with Survivin prevented accumulation of Borealin in the nucleoli, and increased its cytoplasmic localization (panel b4). Coexpression with the negative control YFP-NLS (panels e1–e4) did not alter the localization of any of the CPPs. (B) Quantification of the effect of INCENP coexpression on the localization of Aurora B. Graphs show the proportion of cells expressing HA-Aurora B predominantly in the cytoplasm (C>N), at similar levels in the cytoplasm and nucleus (C=N) or predominantly in the nucleus (N>C), when expressed alone or coexpressed with VSV-INCENP. The percentage of cells expressing predominantly nuclear Aurora B drastically increases in the presence of coexpressed INCENP. Bars represent the mean±s.d. of three independent experiments. (C) Quantification of the effect of Survivin coexpression on the localization of Borealin. Graphs show the proportion of cells expressing VSV-Borealin predominantly in the cytoplasm (C), in both the cytoplasm and nucleolus (C+Nol) or predominantly in the nucleolus (Nol), when expressed alone or coexpressed with YFP-Survivin. Bars represent the mean of two independent experiments with less than 10% variation. More than 200 cells per sample were counted in each experiment.
The nucleocytoplasmic distribution of Aurora B coexpressed with Survivin or Borealin was similar to its localization when expressed alone or with the YFP-NLS control. In contrast, coexpression with INCENP led to a prominent relocation of Aurora B to the nucleus. As shown in Figure 6B, localization of Aurora B was predominantly nuclear in more than 60% of INCENP-co-transfected cells. We confirmed that the effect of INCENP on the localization of Aurora B was specific using control experiments (data not shown) in which coexpression with INCENP did not induce nuclear accumulation of the shuttling protein Snurportin I (Paraskeva et al., 1999), or the cytoplasmic protein TRAF2 (Birbach et al., 2002).
The predominantly cytoplasmic localization of Survivin, on the other hand, did not significantly change when coexpressed with any of the other CPPs, although a faint nucleolar signal was noted in some cells coexpressing Borealin. It is remarkable that, in contrast to Aurora B, Survivin remained in the cytoplasm when coexpressed with INCENP.
The localization of INCENP, like that of Survivin, was not significantly altered by coexpression of any of the other CPPs, and this protein remained nuclear in virtually all transfected cells.
Finally, a striking change in the localization of Borealin was observed in cells coexpressing Survivin. In these cells, the nucleolar localization of Borealin was completely abrogated. Survivin and Borealin colocalized in the cytoplasm in approximately 90% of the co-transfected cells (Figure 6C). In contrast to Survivin, neither Aurora B nor INCENP disrupted the nucleolar accumulation of Borealin. We must point out, however, that the coexpression efficiency of VSV-INCENP and YFP-Borealin was consistently low in the several experiments we attempted and only a limited number of co-transfected cells could be examined.
In summary, the results of these experiments uncover a novel role for INCENP and Survivin as specific modulators of the subcellular distribution of Aurora B and Borealin, respectively, before NEB.
Modulation of the subcellular localization of Borealin by Survivin
As Borealin is the less well characterized of the CPPs, we explored in more detail how Survivin modulates its localization before NEB. First, we carried out a deletion analysis (Figure 7a) to determine what region of Survivin mediates its effect on Borealin localization. As shown in Figure 7b, the results of this analysis indicated that the carboxy-terminal domain of Survivin is both necessary and sufficient to prevent the nucleolar accumulation of Borealin. This observation is consistent with our finding that this region of Survivin is sufficient to bind Borealin in co-immunoprecipitation experiments (data not shown), and raised two interesting possibilities. First, it is known that alternative splicing of Survivin mRNA gives origin to several Survivin protein isoforms (Mahotka et al., 1999) that contain different carboxy-terminal amino-acid sequences (depicted in Figure 7a) and thus might differ in their ability to modulate Borealin localization. Indeed, Survivin-2B, which contains an in-frame novel sequence of 23 amino acids, prevented nucleolar localization of Borealin as efficiently as Survivin, whereas Survivin-ΔEx3, which bears a carboxy-terminal domain completely unrelated to that of Survivin, was unable to prevent the nucleolar localization of Borealin (Figure 7b). Second, we have previously reported that the carboxy-terminal region of Survivin mediates its nucleocytoplasmic transport (Rodriguez et al., 2002). Thus, binding of Borealin could modulate the access of Survivin to the nuclear transport machinery and vice versa. As illustrated in Figure 7c, LMB treatment of co-transfected cells led to the efficient nuclear accumulation of YFP-Survivin, demonstrating that the nucleocytoplasmic shuttling ability of Survivin is not impaired in cells coexpressing VSV-Borealin. In some cells, we observed simultaneous nuclear relocation of Survivin and Borealin, but, in most cases, only Survivin relocated to the nucleus after LMB treatment (Figure 7c). Next, we coexpressed VSV-Borealin with YFP-Survivin+NLS, a version of Survivin targeted to the nucleus by fusion of the SV40 NLS. As shown in Figure 7d, coexpression of YFP-Survivin+NLS led to a diffuse nuclear localization of Borealin. In contrast to the LMB-induced nuclear accumulation of Survivin, SV40 NLS-mediated nuclear targeting of Survivin invariably led to the concomitant nuclear relocation of coexpressed Borealin, suggesting that the interaction of Borealin and Survivin is more efficient when the nuclear import of Survivin is mediated by a heterologous sequence outside its carboxy-terminal domain.
Modulation of the subcellular localization of Borealin by Survivin before nuclear envelope breakdown. (a) Schematic representation of the YFP-tagged deletion mutants and splice variants of Survivin used in the experiments. The BIR domain of Survivin is depicted in gray. The regions of Survivin-2B and Survivin-ΔEx3 containing amino-acid sequences different from Survivin are depicted in black. (b) The carboxy-terminal region of Survivin is necessary and sufficient to prevent nucleolar localization of Borealin. Images show representative examples of the effect of coexpressing each of the fragments and isoforms of Survivin on the subcellular localization of VSV-Borealin. Like the full-length protein, the carboxy-terminal region of Survivin (74–142), and Survivin-2B abrogate nucleolar accumulation of Borealin. In contrast, Borealin remains in the nucleoli when coexpressed with the amino-terminal region of Survivin (1–74) or with Survivin-ΔEx3. (c) Survivin remains a nuclear shuttling protein when coexpressed with Borealin. MCF-7 cells coexpressing YFP-Survivin and VSV-Borealin were treated with leptomycin B (LMB) (6 ng/ml for 3 h). In untreated samples, YFP-Survivin and VSV-Borealin colocalize in the cytoplasm. After LMB treatment, Survivin efficiently accumulates in the nucleus. In a proportion of co-transfected cells (the mean±s.d. of three independent experiments is indicated inside the panels), simultaneous nuclear relocation of Borealin was observed. (d) Nuclear targeting of Survivin mediated by a heterologous nuclear localization signal (NLS) leads to nuclear localization of coexpressed Borealin. Images show that VSV-Borealin is diffusely localized within the nucleus (not accumulated in the nucleoli) when coexpressed with YFP-Survivin+NLS, a nuclear version of Survivin bearing an SV40 NLS fused to its carboxy-terminal end.
Discussion
The CPPs Survivin, Aurora B, INCENP and Borealin sequentially colocalize to the chromosome centromeres, the spindle midzone, the cleavage furrow and finally to the midbody as mitosis progresses (Adams et al., 2001a, 2001b; Gassman et al., 2004; Sampath et al., 2004). The CPPs are already expressed at the G2/M transition and, at least in the case of Survivin, expression at earlier cell cycle stages has been reported in different types of both normal and tumor cells (Krysan et al., 2004; Song et al., 2005). Before NEB, which takes place at the transition between prophase and prometaphase, the nuclear envelope poses a physical barrier between the nucleus and the cytoplasm. During most of the cell cycle, dynamic nucleocytoplasmic transport across the nuclear envelope determines the localization of many cancer-related proteins, including Survivin (Rodriguez et al., 2002). Here, we have used epitope-tagged versions of the other CPPs to investigate the molecular mechanisms that determine the nucleocytoplasmic distribution of Aurora B, INCENP and Borealin in the presence of an intact nuclear envelope.
Our results reveal that the localization of the CPPs before NEB is not as closely related as it is during mitosis and that their nucleocytoplasmic transport is, at least in part, independently regulated. Our findings are summarized in the model depicted in Figure 8. We show that Aurora B, like Survivin, accesses the CRM1-dependent nuclear export pathway and undergoes LMB-sensitive shuttling between the nucleus and the cytoplasm, whereas CRM1-mediated export does not seem to play a major role as a direct determinant of the nucleocytoplasmic distribution of INCENP and Borealin. Several groups, including ours, have shown that the nucleocytoplasmic localization of Survivin in tumor cells determined by immunohistochemistry (IHC) is related to prognosis in patients with different types of cancer (Altura et al., 2003; Vischioni et al., 2004; Tonini et al., 2005). It is presently unknown if a similar relationship with clinical parameters exists in the case of Aurora B localization, as the number of studies evaluating the expression Aurora B in human tumors using IHC is very limited. The functional consequences of the nucleocytoplasmic shuttling of Survivin and Aurora B before NEB remain to be established. However, it has been recently reported that, besides its function as a nuclear export receptor, CRM1 has a role determining the localization of proteins to chromosome kinetochores during mitosis (Arnaoutov et al., 2005). In this respect, our initial experiments demonstrate that extended LMB treatment disrupts the localization of endogenous Survivin and Aurora B to the centromeric region of prophase and (pro)metaphase cells. We show that, unlike that of RanBP2 (Arnaoutov et al., 2005), the localization of the CPPs is only disrupted after sustained blockage of CRM1 activity, suggesting that the presence of LMB during the preceding interphase may contribute to the observed effect. Therefore, further experimental evidence is needed to clarify the relationship between the CRM1 pathway and the CPPs during mitosis. For example, it remains to be established if the altered localization of Survivin and Aurora B is a direct or indirect consequence of blocking the CRM1 pathway. In the light of these results, however, it is tempting to speculate that the CRM1-dependent nuclear shuttling of Aurora B and Survivin before NEB may reflect a functional interaction of the CPPs with the CRM1 pathway that is relevant for events that take place after NEB.
Nucleocytoplasmic localization of the chromosomal passenger proteins (CPPs) before nuclear envelope breakdown. Model summarizing the mechanisms that regulate the nucleocytoplasmic distribution of the CPPs in the presence of an intact nuclear envelope. Aurora B undergoes CRM1-mediated nuclear export and is, like Survivin, able to shuttle between nucleus and cytoplasm. INCENP contains multiple functional nuclear localization signals (NLSs) and is efficiently imported/retained into the nucleus. Borealin also contains functional NLSs (Gassman et al., 2004) and, after translocation into the nucleus, it accumulates in the nucleolus. Although not depicted in the model, the presence of Aurora B and INCENP (but not of Survivin) in the nucleolus has also been reported (Andersen et al., 2005). INCENP modulates the nucleocytoplasmic distribution of Aurora B (probably through increased nuclear import and/or reduced nuclear export/enhanced retention of the kinase in the nucleus), but not of Survivin. Finally, Survivin prevents the targeting of Borealin to the nucleolus.
Despite their dynamic nucleocytoplasmic transport, our functional assays failed to identify active NLSs or NESs of the ‘classical’ type in Aurora B (this report) or Survivin (Rodriguez et al., 2002). Although it remains possible that other ‘non-classical’ signals in these proteins mediate their direct interaction with nuclear transport receptors, our findings suggest that the nucleocytoplasmic traffic of Survivin or Aurora B could be indirectly modulated through interaction with other NES- or NLS-containing proteins. In this regard, both INCENP (this report) and Borealin (Gassman et al., 2004) contain functional ‘classical’ NLSs, and therefore they were obvious candidates to modulate the nucleocytoplasmic distribution of Survivin and Aurora B.
Indeed, coexpression with INCENP dramatically increased the nuclear accumulation of Aurora B. It could be argued that coexpression of any protein with an NLS-bearing binding partner would lead to the nuclear localization of both proteins. Remarkably, however, coexpression with INCENP did not induce nuclear relocation of Survivin, although the ability of these two proteins to physically interact is well established (Bolton et al., 2002; Wheatley et al., 2001). Thus, these results indicate that INCENP plays a specific role as modulator of Aurora B nucleocytoplasmic distribution by increasing its nuclear import and/or its retention in the nucleus.
In contrast to INCENP, Borealin did not significantly alter the nucleocytoplasmic distribution of neither Aurora B nor Survivin. Rather, our data indicate that it is Survivin that modulates the subcellular localization of Borealin before NEB. As previously described using a GFP-tagged version (Gassman et al., 2004), a prominent characteristic of the subcellular distribution of VSV-Borealin before NEB was its accumulation in nucleoli. The nucleolar localization of GFP-Borealin was considered a non-physiological feature in the initial report, because the authors did not detect endogenous Borealin in this organelle using immunocytochemistry (ICC) (Gassman et al., 2004). More recently, however, a highly sensitive proteomics approach using mass spectrometry has revealed the presence of endogenous Borealin in the nucleolus (Andersen et al., 2005, see the Nucleolar Proteome Database available at http://www.lamondlab.com/NOPdb/), suggesting that Borealin is a bona fide nucleolar protein. In fact, not only Borealin but also Aurora B and INCENP were found to copurify with human nucleoli. These findings raise the intriguing possibility that the nucleolus may function as a reservoir for a fraction of Borealin, Aurora B and INCENP before NEB, and it would be interesting to dissect the mechanisms that modulate the trafficking of the CPPs in and out of this nuclear organelle. In this context, we noted that two of the INCENP NLSs strongly targeted the fused YFP to the nucleoli, suggesting that they may contribute to the nucleolar localization of INCENP.
More importantly, our data reveal a previously unknown role for Survivin as a potent negative modulator of the nucleolar accumulation of Borealin. We show that the coexpression of both proteins results in Survivin efficiently blocking the nucleolar localization of Borealin. Thus, the low levels of endogenous Borealin in nucleoli, undetectable by ICC, could be explained by its balanced expression with endogenous Survivin. This novel role of Survivin is mediated by its carboxy-terminal end and is retained in the splice variant Survivin-2B, but lost in Survivin-ΔEx3. Survivin isoforms have been previously reported to differ in their antiapoptotic activity (Mahotka et al., 1999). Our results identify another functional divergence between these proteins that is most likely non-related to apoptosis regulation, as the isoform Survivin-2B, which has reportedly lost the antiapoptotic ability (Mahotka et al., 1999), is still able to prevent the nucleolar localization of Borealin.
Finally, we show that nuclear shuttling of Survivin is not impaired when coexpressed with Borealin, although the carboxy-terminal region of Survivin mediates both its nuclear transport and its effect on Borealin. Importantly, only a minor fraction of Borealin relocates to the nucleus along with Survivin in the presence of LMB. In contrast, enforced nuclear accumulation of Survivin by fusion of a heterologous NLS invariably results in the concomitant nuclear relocation of Borealin. These results are consistent with a model in which the binding of proteins that mediate its nuclear transport would largely impair the interaction of Survivin with Borealin. Therefore, we suggest that, although Survivin modulates the nucleolar localization of Borealin before NEB, the nucleocytoplasmic transport of both proteins is mostly unrelated.
In conclusion, we show that INCENP and Survivin are able to specifically modulate the subcellular localization of Aurora B and Borealin, respectively, before NEB. The existence of at least two different chromosomal passenger complexes in mitotic cells has been recently proposed (Gassman et al., 2004). One of these complexes contains only Aurora B and INCENP, whereas the second complex also includes Survivin and Borealin. Consistent with this view, our findings suggest that different complexes of CPPs may also exist before NEB and undergo different dynamics of transport between the nucleus and the cytoplasm.
Materials and methods
Cell culture, transfection and drug treatments
Human breast carcinoma cells MCF-7 were grown in Dulbecco's modified Eagle's medium (DMEM) (BioWhittaker, Walkersville, MD, USA), supplemented with 10% fetal calf serum, 100 U/ml penicillin and 100 μg/ml streptomycin (Gibco BRL, Gaithersburg, MD, USA). Cells were seeded onto sterile glass coverslips in 12-well trays and transfected with 0.5–2 μg of plasmid DNA using the FuGene6 transfection reagent (Roche Molecular Biochemicals). Leptomycin B (LMB, a generous gift from Dr Minoru Yoshida, University of Tokyo, Japan) and actinomycin D (Sigma, St. Louis, MO, USA) were added to the culture medium to a final concentration of 6 ng/ml and 5 μg/ml, respectively. Mitotic-arrested cells were collected by shake-off after overnight incubation with 50 ng/ml nocodazole (Sigma) and released from the arrest in nocodazole-free DMEM with or without LMB (6 ng/ml). After 1 h incubation, cytospins were prepared using a Cytospin 2 (Shandon, Waltham, MA, USA).
Plasmids and cloning procedures
The plasmids encoding HA-TRAF2, HA-Snurportin I and GFP-Lamin A were generously provided by Dr Colin Duckett (University of Michigan, USA), Dr Sylvain Meloche (Institute de Recherches Cliniques de Montreal, Canada) and Dr David Gilbert (State University of New York, USA). The plasmids encoding YFP-CRM1, Flag-Survivin, YFP-Survivin, YFP-Survivin-2B and YFP-Survivin-ΔEx3 have been previously described (Rodriguez and Henderson, 2000; Rodriguez et al., 2002). Survivin deletion mutants YFP-Survivin(1–74) and YFP-Survivin(74–142) were generated by PCR using Survivin complementary DNA (cDNA) as template. Aurora B, Borealin and INCENP cDNAs were obtained by reverse transcriptase–PCR and cloned in-frame to a 5′ sequence encoding the VSV epitope into pCR3 (Aurora B and Borealin) or pCDNA3 (INCENP) vectors to generate VSV-Aurora B, VSV-Borealin and VSV-INCENP, as described in detail elsewhere (Vader et al., 2006). Carboxy-terminally tagged Aurora B-HA, YFP-Borealin, YFP-INCENP(1–454) and YFP-INCENP(455–919) were generated by PCR using Aurora B, Borealin and INCENP cDNA as templates. The PCR products were cloned into pCDNA3 (Aurora B) and pEYFP-C1 (Borealin and INCENP). On the other hand, site-directed mutagenesis was used to introduce the K106R point mutation in Aurora B sequence, generating the catalytically inactive Aurora B KD, and to delete INCENP NLSs in YFP-INCENP(1–454). Survivin+NLS was constructed using PCR to add a DNA sequence encoding the SV-40 large T-antigen nuclear localization signal (SV-40 NLS) to the 3′ end of Survivin. The PCR product was cloned as a HindIII/BamHI fragment into the pCDNA3 and pEYFP-C1 vectors to generate Flag-Survivin+NLS and YFP-Survivin+NLS, respectively. Finally, cDNA oligonucleotides encoding the SV-40 NLS were annealed, phosphorylated and cloned into HindIII/BamHI-digested pEYFP-C1 to make the control construct YFP-NLS. The high-fidelity DNA polymerase Pfu (Stratagene, La Jolla, CA) was used in all PCR reactions and the sequence of the inserts was verified by DNA sequencing. The sequence of the oligonucleotides used in plasmid construction is available upon request.
Functional analysis of putative transport signals in Aurora B and INCENP
DNA fragments encoding two putative NESs identified in Aurora B protein sequence were generated by PCR using Aurora B cDNA as template. PCR products were cloned into the pRev(1.4)-GFP plasmid and the export activity of these sequences was tested in a nuclear export assay, as previously described (Henderson and Eleftheriou, 2000). On the other hand, Aurora B and INCENP sequences encoding putative NLSs were cloned into HindIII/BamHI-digested pEYFP-C1. The import activity of these sequences was tested by comparing the nucleocytoplasmic distribution of the fusion proteins to that of the YFP protein alone (negative control) and YFP-NLS (positive control).
Immunofluorescence and microscopy analysis
Cells were fixed with 3.7% formaldehyde in phosphate-buffered saline (PBS) for 30 min and permeabilized with 0.2% Triton X-100 in PBS for 10 min. After 1 h incubation in blocking solution (3% bovine serum albumin (BSA) in PBS), immunocytochemical detection of endogenous proteins was carried out using a rabbit anti-NF-κB polyclonal antibody (diluted 1:300; Santa Cruz, Biotechnology, Santa Cruz, CA, USA), mouse anti-Survivin monoclonal antibody 6E4 (diluted 1:200; Cell Signaling Technology, Danvers, MA, USA), mouse anti-AIM-1 (Aurora B) monoclonal antibody (diluted 1:300; BD Biosciences, Franklin Lakes, NJ, USA), a human anti-CREST antibody provided by J Kuijpers (University Medical Center Utrecht, The Netherlands) and a goat anti-RanBP2 antibody provided by F Melchior (Georg-August Universität Gottingen, Germany). Detection of epitope-tagged proteins was performed with anti-VSV monoclonal antibody (diluted 1:20 000; Sigma), anti-HA rabbit polyclonal antibody Y-11 (diluted 1:300; Santa Cruz) or anti-Flag M2 monoclonal antibody (diluted 1:200; Stratagene). Primary antibodies were detected using anti-mouse, anti-rabbit, anti-human or anti-goat secondary antibodies conjugated to fluorescein isothiocyanate (Santa Cruz and Sigma), Alexa Fluor 594 or Alexa Fluor 488 (Molecular Probes, Carlsbad, CA, USA). The chromosome dye Hoechst 33285 (Sigma) was used to counterstain the nuclei. Finally, the coverslips were mounted onto microscope slides using Vectashield (Vector, Burlingame, CA, USA).
Fluorescence microscopy analysis was carried out using an inverted Leica DMIRB/E fluorescence microscope. Images were collected using the Q500MC Quantimet software V01.01 (Leica Cambridge Ltd, Cambridge, UK). To quantitatively determine the subcellular distribution of each protein, its localization was assessed in a single-cell basis in at least 200 transfected cells per sample.
References
Adams RR, Carmena M, Earnshaw WC . (2001a). Trends Cell Biol 11: 49–54.
Adams RR, Eckley DM, Vagnarelli P, Wheatley SP, Gerloff DL, Mackay AM et al. (2001b). Chromosoma 110: 65–74.
Altieri DC . (2003a). Oncogene 22: 8581–8589.
Altieri DC . (2003b). Nat Rev Cancer 3: 46–54.
Altura RA, Olshefski RS, Jiang Y, Boue DR . (2003). Br J Cancer 89: 1743–1749.
Andersen JS, Lam YW, Leung AKL, Ong S-E, Lyon CE, Lamond AI et al. (2005). Nature 433: 77–83.
Andrews PD, Knatko E, Moore WJ, Swedlow JR . (2003). Curr Opin Cell Biol 15: 672–683.
Arnaoutov A, Azuma Y, Ribbeck K, Joseph J, Boyarchuk Y, Karpova T et al. (2005). Nat Cell Biol 7: 626–632.
Birbach A, Gold P, Binder BR, Hofer E, de Martin R, Schmid JA . (2002). J Biol Chem 277: 10842–10851.
Bischoff JR, Anderson L, Zhu Y, Mossie K, Ng L, Souza B et al. (1998). EMBO J 17: 3052–3065.
Bishop JD, Schumacher JM . (2002). J Biol Chem 277: 27577–27580.
Bolton MA, Lan W, Powers SE, McCleland ML, Kuang J, Stukenberg PT . (2002). Mol Biol Cell 13: 3064–3077.
Burke B, Ellenberg J . (2002). Nat Rev Mol Cell Biol 3: 487–497.
Chen J, Jin S, Tahir SK, Zhang H, Liu X, Sarthy AV et al. (2003). J Biol Chem 278: 486–490.
Fabbro M, Henderson BR . (2003). Exp Cell Res 282: 59–69.
Gassman R, Carvalho A, Henzing AJ, Ruchaud S, Hudson DF, Honda R et al. (2004). J Cell Biol 166: 179–191.
Giet R, Petretti C, Pringent C . (2005). Trends Cell Biol 15: 241–250.
Henderson BR, Eleftheriou A . (2000). Exp Cell Res 256: 213–224.
Honda R, Körner R, Nigg EA . (2003). Mol Biol Cell 14: 3325–3341.
Johnson C, Van Antwerp D, Hope TJ . (1999). EMBO J 18: 6682–6693.
Kalderon D, Roberts BL, Richardson WD, Smith AE . (1984). Cell 39: 499–509.
Kawajiri A, Yasui Y, Goto H, Tatsuka M, Takahashi M, Nagata K et al. (2003). Mol Biol Cell 14: 1489–1500.
Krysan K, Merchant FH, Zhu L, Dohadwala M, Luo J, Lin Y et al. (2004). FASEB J 18: 206–208.
La Cour T, Kiemer L, Molgaard A, Gupta R, Skriver K, Brunak S . (2004). Protein Eng Des Sel 17: 527–536.
Lens SMA, Wolthuis RM, Klompmaker R, Kauw J, Agami R, Brummelkamp T et al. (2003). EMBO J 22: 2934–2947.
Mahotka C, Wenzel M, Springer E, Gabbert HE, Gerharz CD . (1999). Cancer Res 59: 6097–6102.
Mattaj IW, Englmeier L . (1998). Annu Rev Biochem 67: 265–306.
Nakai K, Horton P . (1999). Trends Biochem Sci 24: 34–35.
Paraskeva E, Izaurralde E, Bischoff FR, Huber J, Kutay U, Hartmann E et al. (1999). J Cell Biol 145: 255–264.
Rodriguez JA, Henderson BR . (2000). J Biol Chem 275: 38589–38596.
Rodriguez JA, Schüchner S, Au WWY, Fabbro M, Henderson BR . (2004). Oncogene 23: 1809–1820.
Rodriguez JA, Span SW, Ferreira CGM, Kruyt FAE, Giaccone G . (2002). Exp Cell Res 275: 44–53.
Saffery R, Irvine DV, Kile BT, Hudson DF, Cutts SM, Choo KHA . (1999). Mamm Genome 10: 415–418.
Sampath SC, Ohi R, Leismann O, Salic A, Pozniakovski A, Funabiki H . (2004). Cell 118: 187–202.
Song J, So T, Cheng M, Tang X, Croft M . (2005). Immunity 22: 621–631.
Speliotes EK, Uren A, Vaux D, Horvitz HR . (2000). Mol Cell 6: 211–223.
Terada Y, Tatsuka M, Suzuki F, Yasuda Y, Fujita S, Otsu M . (1998). EMBO J 17: 667–676.
Tonini G, Vincenzi B, Santini D, Scarpa S, Vasaturo T, Malacrino C et al. (2005). Br J Cancer 92: 2225–2232.
Vader G, Kauw JJW, Medema RH, Lens SMA . (2006). EMBO Rep 7: 85–96.
Vischioni B, van der Valk P, Span SW, Kruyt FAE, Rodriguez JA, Giaccone G . (2004). Ann Oncol 15: 1654–1660.
Wheatley SP, Carvalho A, Vagnarelli P, Earnshaw WC . (2001). Curr Biol 11: 886–890.
Wheatley SP, Henzing AJ, Dodson H, Khaled W, Earnshaw WC . (2004). J Biol Chem 279: 5655–5660.
Yasui Y, Urano T, Kawajiri A, Nagata K, Tatsuka M, Saya H et al. (2004). J Biol Chem 279: 12997–13003.
Acknowledgements
We thank Drs C Duckett, S Meloche, D Gilbert and BR Henderson for providing plasmids, Drs J Kuijpers and F Melchior for providing antibodies and Dr M Yoshida for providing LMB. JA Rodriguez was supported by the Walter Bruckerhoff Stiftung.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rodriguez, J., Lens, S., Span, S. et al. Subcellular localization and nucleocytoplasmic transport of the chromosomal passenger proteins before nuclear envelope breakdown. Oncogene 25, 4867–4879 (2006). https://doi.org/10.1038/sj.onc.1209499
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209499
Keywords
This article is cited by
-
A cellular reporter to evaluate CRM1 nuclear export activity: functional analysis of the cancer-related mutant E571K
Cellular and Molecular Life Sciences (2016)
-
Potential effects of CRM1 inhibition in mantle cell lymphoma
Chinese Journal of Cancer Research (2012)
-
Expression of HuR, COX-2, and survivin in lung cancers; cytoplasmic HuR stabilizes cyclooxygenase-2 in squamous cell carcinomas
Modern Pathology (2011)
-
The Survivin–Crm1 interaction is essential for chromosomal passenger complex localization and function
EMBO reports (2006)