Abstract
Over the years, p53 has been shown to sit at the centre of an increasingly complex web of incoming stress signals and outgoing effector pathways. The number and diversity of stress signals that lead to p53 activation illustrates the breadth of p53's remit – responding to a wide variety of potentially oncogenic insults to prevent tumour development. Interestingly, different stress signals can use different and independent pathways to activate p53, and there is some evidence that different stress signals can mediate different responses. How each of the responses to p53 contributes to inhibition of malignant progression is beginning to be clarified, with the hope that identification of responses that are key to tumour suppression will allow a more focused and effective search for new therapeutic targets. In this review, we will highlight some recently identified roles for p53 in tumour suppression, and discuss some of the numerous mechanisms through which p53 can be regulated and activated.
Similar content being viewed by others
Introduction
Since its discovery over a quarter of a century ago, work on the tumour suppressor protein p53 has produced nearly 40 000 publications. This vast amount of work on one protein is arguably justified by the importance of p53 in cancer biology, as illustrated by the frequency with which p53 function is lost during tumour development. Approximately 50% of all malignancies carry a p53 mutation, and of those tumours that do not have mutated p53, a large proportion have inactivated p53 function by another mechanism (Hainaut and Hollstein, 2000; Bykov and Wiman, 2003). Taken together, p53 function is lost in almost all tumours. The importance of p53 as a tumour suppressor is illustrated by the observation that many individuals affected by Li Fraumeni syndrome – and so displaying an abnormally high and early incidence of tumour development – suffer a germline mutation in p53 (Malkin et al., 1990; Srivastava et al., 1990). Furthermore, mouse studies have corroborated the importance of p53 in tumour development; p53 null mice can develop normally but nearly all develop cancer before 6 months of age (Donehower et al., 1992; Jacks et al., 1994; Purdie et al., 1994). Taken together, a role for p53 as a key player in the inhibition of tumour development is beyond doubt. However, there is growing evidence that p53 also contributes to other pathologies beyond cancer – including a possible contribution in determining longevity – making it unlikely that the high degree of interest in p53 will wane.
The downstream response to p53 activation
The p53 protein is a transcription factor that regulates the expression of a large number of target genes (Vogelstein et al., 2000). Through this mechanism p53 can induce a number of different responses, ranging from the induction of cell cycle arrest, cell death and senescence to protective antioxidant activities, DNA repair and the control of mitochondrial respiration.
One of the first transcriptional targets of p53 to be identified was the cyclin dependent kinase (CDK) inhibitor p21Waf1/Cip1 (El-Deiry et al., 1994). CDKs play an important role in regulating progression through the cell cycle, and the inhibition of CDK activity by p21Waf1/Cip1 results in a cell cycle arrest (reviewed in Sherr and Roberts, 1999). Thus, p53 can effectively cause cells to stop proliferating, which – among other things – allows for the repair of any damaged DNA, preventing mutations from being passed on to daughter cells. Interestingly, a direct role for p53 in DNA repair has also been demonstrated (reviewed in Gatz and Wiesmuller (2006)). Recently, other activities of p53 that can inhibit the acquisition of potentially oncogenic mutations have been described. Several studies have shown an antioxidant role for p53 resulting from the expression of target genes that reduce intracellular reactive oxygen species (ROS) levels and so prevent genotoxic damage (Budanov et al., 2004; Yoon et al., 2004; Sablina et al., 2005; Bensaad et al., 2006). This function of p53 appears to be important even in the absence of acute, high levels of stress – and probably reflects a response to lower levels of constitutive stress that accompany everyday life of complex multicellular organisms (Sablina et al., 2005). These antioxidant functions of p53 are linked with enhanced survival of cells expressing p53, and so seem to be at odds with another important activity of p53, which is to induce cell death through apoptosis or autophagy. Why would p53 induce repair of a cell that it then targets for destruction? While some of the answer clearly lies in cell-type dependent differences, an interesting model is emerging where p53 behaves differently depending on the severity of the stress (Bensaad and Vousden, 2005). In response to low, constitutive stress, p53 activates expression of genes that drive a temporary cell cycle arrest, allowing for the removal of intracellular ROS and repair of any damage that may have occurred. But when the stress-induced damage is too severe for the cell to recover, p53 initiates programmed cell death – so eliminating cells that may have acquired irreparable and potentially oncogenic alterations. The transcriptional targets of p53 include a number of pro-apoptotic proteins such as the BH3 only proteins Puma and Noxa (Oda et al., 2000; Nakano and Vousden, 2001; Yu et al., 2001). These proteins function by inducing the loss of inner mitochondrial membrane potential, leading to the release of cytochrome C and other apoptogenic factors and the activation of the caspase cascade that results in apoptotic cell death (Jeffers et al., 2003; Villunger et al., 2003; Yu et al., 2003 and reviewed in Danial and Korsmeyer, 2004). Activation of autophagy by the p53-induced protein DRAM (damage-regulated autophagy modulator) has also been described to be an important contributor of the apoptotic response (Crighton et al., 2006).
Inkeeping with its function as a transcription factor, p53 is predominately a nuclear protein. However, a fraction of p53 can be found in the cytoplasm. Recent studies have demonstrated an unexpected transcriptionally independent function for this pool of p53 as a BH3-only protein, inducing apoptosis by directly binding to the pro-apoptotic proteins Bax and Bak, as well as the anti-apoptotic proteins like BclxL (Mihara et al., 2003; Chipuk et al., 2004; Leu et al., 2004). Interestingly PUMA, a transcriptional target of p53, can activate this cytoplasmic function of p53 (Chipuk et al., 2005), suggesting that both transcriptional and non-transcriptional activities of p53 are important for the induction of cell death (reviewed in Yee and Vousden, 2005).
Regulation of p53 function
The anti-proliferative activities of p53 play an important role in tumour suppression, but require strong negative regulation to allow for normal growth and development. Not surprisingly, this regulation of p53 takes many forms, including the regulation of expression, protein stability and protein activity. Many post-transcriptional modifications of p53 have been described including phosphorylation, acetylation, methylation, glycosylation, ribosylation and, most recently, O-GlcNAcylation (Bode and Dong, 2004; Yang et al., 2006c). However, in vivo models have suggested that these modifications play only very subtle, modulatory roles in regulating p53 function (Toledo and Wahl, 2006). The interaction of p53 with proteins that control p53 function, either directly or through the regulation of p53 stability, appears to be key in turning on and off the p53 response.
Regulation of p53 protein stability
p53 is degraded via the proteasome by a predominately ubiquitin-dependent mechanism, and a number of ubiquitin ligases that promote the ubiquitination and subsequent degradation of p53 have been identified (Table 1).
By far the best studied of these is Mdm2 (sometimes referred to as HDM2 in humans) – a RING finger protein that is essential to regulate p53 and allow survival during normal development (Jones et al., 1995; Montes de Oca Luna et al., 1995). Mdm2 functions as a ubiquitin ligase, in which the RING domain is necessary to promote the ubiquitination and degradation of its target proteins, including p53 (Fang et al., 2000). Mdm2 is itself a transcriptional target of p53 (Zauberman et al., 1993), establishing a negative feedback loop in which Mdm2 can keep the physiological levels of p53 low, and contribute to the recovery phase at the end of a p53 response.
While Mdm2 is sufficient to target p53 for degradation, there is good evidence that it does not function alone. Several proteins have recently been shown to cooperate with Mdm2 in the regulation of p53. The Yin Yang 1 (YY1) transcription factor, a protein that plays a key role in development, can increase the interaction between p53 and Mdm2, so enhancing Mdm2-dependent p53 poly-ubiquitination and degradation (Gronroos et al., 2004; Sui et al., 2004).
A similar function has also been attributed to gankyrin, with the gankyrin/Mdm2 interaction promoting p53 binding and degradation (Higashitsuji et al., 2005a, 2005b). The mechanism underlying this activity of gankyrin is suggested by the observation that gankyrin associates with the S6 proteasomal ATPase. This interaction of gankyrin, Mdm2 and p53 also promotes increased association of the Mdm2-p53 complex to the 26S proteasome, and so promotes more efficient degradation of p53 and Mdm2 (Dawson et al., 2002; Higashitsuji et al., 2005a). The frequent overexpression of gankyrin in hepatocellular carcinomas may therefore be an important mechanism for inactivating the p53 pathway in these tumours (Fu et al., 2002; Nagao et al., 2003; Higashitsuji et al., 2005b).
p300/CBP has also been shown to cooperate with Mdm2 to promote poly-ubiquitination of p53 (Grossman et al., 2003). p300 has a well established role in promoting acetylation of p53, thereby enhancing its transcriptional activity (Grossman, 2001). However, a recent report has suggested that under most conditions Mdm2 can only promote the mono-ubiquitination of p53, and that the binding of p300 to Mdm2 is necessary for efficient poly-ubiquitination to occur (Grossman et al., 2003). As poly-ubiquitination is required for recognition by the proteasome and degradation (reviewed in Weissman, 2001), proteins like p300 may be key in controlling p53 stability.
The recent identification of HAUSP (herpesvirus associated ubiquitin-specific protease) and other de-ubiquitinating (DUB) enzymes has added a further level of complexity to the regulation of p53 stability. Although HAUSP can de-ubiquitinate p53 directly (Li et al., 2002), its principal function in the p53 pathway seems to be in the de-ubiquitination and stabilization of Mdm2 (Cummins and Vogelstein, 2004; Li et al., 2004), resulting in enhanced degradation of p53. This modification is influenced by yet another protein, Daxx (death domain-associated protein), which forms a complex with Mdm2 and HAUSP, preventing the auto-ubiquitination of Mdm2 and thus promoting p53 degradation (Tang et al., 2006).
Of the other p53 ubiquitin ligases listed in Table 1, Cop1, Pirh2, HectH9/MULE/ARF-BP1, E6AP and CHIP have all been shown to promote the degradation of p53 under varying conditions. Like Mdm2, Cop1 and Pirh2 also reside in negative feedback loops by being themselves transcriptional targets of p53 (Leng et al., 2003; Dornan et al., 2004). While each of these ligases have been shown to regulate p53 stability in cells, mouse knockout studies will prove useful in determining their relative contribution to regulating p53 levels in vivo.
Finally, it has been shown that p53 can be degraded through an ubiquitin-independent mechanism as well. This pathway involves the 20S proteasome and is regulated by the NAD(P)H quinone oxidoreductase 1 (NQO1) (Asher et al., 2001). The major pool of NQO1 can be found at the 20S proteasome, where it is thought to act as a gatekeeper. NQO1 can bind directly to p53, preventing it from being degraded in a 20S proteasomal manner. Genotoxic insults such as ionizing radiation lead to increased association between p53 and NQO1, and so to p53 stabilization (Asher and Shaul, 2005).
SUMOylation and NEDDylation
While ubiquitination is the most studied protein modification of p53, the ubiquitin-like proteins SUMO-1 and Nedd8 can also be conjugated to p53 and modify its function (recently reviewed in Watson and Irwin (2006)). As in ubiquitination, specific E3 ligases mediate the modification of p53 by SUMO-1 and Nedd8. Interestingly, Mdm2 also functions as an E3 for Nedd8 – promoting the conjugation of Nedd8 to three C-terminal lysines in p53 that are also targeted by ubiquitination (Xirodimas et al., 2004). Neddylation of p53 can also be mediated by FBX011, a component of the SCF complex (Abida et al., 2006). A number of proteins have been shown to function in the regulation of p53 SUMOylation, including the family of PIAS ligases (PIAS1, PIASxβ and PIASy), (Kahyo et al., 2001; Nelson et al., 2001; Megidish et al., 2002; Schmidt and Muller, 2002). While Neddylation appears to result in the inhibition of p53 function, there have been conflicting reports of the effect of SUMOylation (reviewed in Melchior and Hengst (2002)). However, recent studies have demonstrated that SUMO-modification can activate p53 and so contribute to the induction of p53-mediated senescence and apoptosis (Bischof et al., 2006; Li et al., 2006). Whereas the modification by SUMO-1 and Nedd8 does not appear to modulate p53 stability directly, SUMOylation can mediate an indirect effect by regulating Mdm2's stability and function (Lee et al., 2006).
MdmX
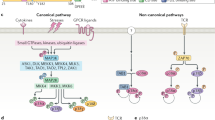
Another key regulator of p53 is MdmX (Mdm4), a structural relative of Mdm2. Like Mdm2, MdmX contains an N-terminal p53-binding domain and a RING domain at its C-terminus (Shvarts et al., 1996). However, MdmX does not show E3 activity (Jackson and Berberich, 2000; Stad et al., 2001) and functions predominantly by directly inhibiting p53's transcriptional activity (Shvarts et al., 1996; Francoz et al., 2006). Although this function of MdmX is independent of Mdm2 (Francoz et al., 2006; Xiong et al., 2006), there are some interesting levels of cross talk between these two proteins. MdmX and Mdm2 hetero-oligomerize through their RING domains (Sharp et al., 1999; Tanimura et al., 1999), which results in the stabilization of Mdm2 (Stad et al., 2001; Gu et al., 2002) or the Mdm2-mediated degradation of MdmX (de Graaf et al., 2003; Kawai et al., 2003; Pan and Chen, 2003), depending on the relative expression levels of the two proteins. Recent studies have also suggested that a heteromeric complex between Mdm2 and MdmX can retain ubiquitin ligase activity, suggesting that despite the inactive RING, MdmX may play a direct role in E3 activity (Poyurovsky et al., 2006; Uldrijan et al., 2006) (Figure 1).
Mdm2 and MdmX. Comparison of the structural domains of Mdm2 and MdmX, showing the regions of similarity. Although both proteins contain a RING domain, only Mdm2 is functional as an E3 ubiquitin ligase (Marine and Jochemsen, 2005).
p53 subcellular localization
As might be expected for a protein with both nuclear and cytoplasmic functions, the subcellular localization of p53 can also play an important role in regulating its activity. Mdm2 can promote the nuclear export of p53 in a ubiquitin-ligase-dependent manner (Boyd et al., 2000; Lohrum et al., 2001; Li et al., 2003). Mdm2 and p53 do not leave the nucleus together; rather it seems that mono-ubiquitination of p53 by Mdm2 leads to an unmasking of the nuclear export sequence within the C-terminus of p53 (Gu et al., 2001; Li et al., 2003). Two other ubiquitin ligases also promote the cytoplasmic localization of p53, Cullin 7 (CUL7) and WWP1 (Andrews et al., 2006; Laine and Ronai, 2006). Neither of these E3s target p53 for degradation, but result in the accumulation of transcriptionally inactive p53 in the cytoplasm. Similarly, regulation of p53 ubiquitination by the E2 ubiquitin-conjugating enzyme Ubc13 has been shown to drive nuclear export of p53 (Laine et al., 2006). In a further twist, ubiquitination of p53 by E4F1 results in neither degradation nor nuclear export, but enhances transcriptional activation resulting selectively in the cell cycle arrest response (Le Cam et al., 2006).
While cytoplasmic sequestration inhibits p53 – and is seen as a mechanisms to inactivate p53 in some breast cancers and neuroblastomas (Moll et al., 1992, 1995) – the relocalization of at least some p53 from the nucleus is important for the cytoplasmic functions of p53. The effect of Mdm2 on p53 function is therefore complex – while degradation of p53 clearly inactivates all functions, some level of Mdm2-mediated ubiquitination may be required for export and full apoptotic activity (Moll et al., 2006).
A delicate balance
In both humans and mice, there is exquisite sensitivity to modulation of the p53 pathway, with relatively modest changes in p53 function resulting in profound effects. This has been highlighted by studies investigating the consequences of modifying the naturally occurring ratio between Mdm2 and p53. Increasing potential p53 activity, by adding an extra copy of p53 or reducing (but not eliminating) expression of Mdm2, resulted in apparently normal mice that were resistant to tumour development (Garcia-Cao et al., 2002, 2006; Mendrysa et al., 2006). However, mice in which overexpression of p53 was accompanied by an imbalance in the normal ratios of different p53 isoforms, showed an alarming premature aging phenotype (Tyner et al., 2002; Maier et al., 2004) that may be associated with p53 functions of driving senescence and modulating the insulin pathway. The contribution of p53 to aging is still not clear, but it would seem that just enough p53 – neither too much nor too little – is required for optimal longevity (Mendrysa and Perry, 2006; Poyurovsky and Prives, 2006).
The consequences of small changes in p53 activity have also been highlighted by the study of polymorphisms in the p53 pathway in humans. A SNP was recently discovered at position 309 of the first intron of Mdm2 (Bond et al., 2004). This polymorphism alters the affinity of an SP1 transcription factor-binding site, and results in a slightly higher expression of Mdm2 in these individuals, dampening the activation of p53 in response to DNA damage. While this results in a relatively modest decrease in the activation of p53, there is a clear effect on tumour susceptibility in individuals carrying this SNP.
A number of polymorphisms of p53 have also been described, including a SNP at amino acid position 72 that changes the amino acid from a proline (P) to an arginine (R). The P72 variant is somewhat better at promoting a cell cycle arrest, while the R72 variant is better at inducing apoptosis (Pim and Banks, 2004) and as such, tumours carrying the R72 variant are generally more responsive to treatment and have a better survival rate, when compared to tumours carrying the P72 variant (reviewed in Pietsch et al. (2006)).
Stress induced stabilization and activation of p53
Efficient control of p53 is critical to allow normal cell growth. But clearly these brakes to p53 function must be released in order for p53 to act as an effective tumour suppressor, and a number of mechanisms that lead to the stabilization and activation of p53 have been identified. A growing number of stress signals that can lead to p53 activation are being identified, including DNA damage, oncogene activation and more. Interestingly, different stress signals seem to utilize different pathways to allow for the activation of p53.
DNA damage
DNA damage efficiently induces a p53 response, in part through the activation of ATM/ATR kinases that result in the phosphorylation of p53, Mdm2 and MdmX. While phosphorylation of p53 and Mdm2 can inhibit the interaction between these two proteins, allowing the stabilization of p53 (Shieh et al., 1997; Siliciano et al., 1997; Banin et al., 1998; Khosravi et al., 1999; de Toledo et al., 2000; Sakaguchi et al., 2000; Maya et al., 2001), phosphorylation of Mdm2 and MdmX also enhances the degradation of these proteins by reducing their association with HAUSP – further releasing p53 from this level of negative regulation (Cummins and Vogelstein, 2004; Li et al., 2004; Meulmeester et al., 2005a, 2005b; Tang et al., 2006). The COP1 ubiquitin ligase is also phosphorylated in an ATM-dependent fashion, leading to a dissociation of COP1 from p53 and so promoting p53 stabilization (Dornan et al., 2006). In addition to the stabilization of p53, DNA damage-induced kinases may also play a role in activating p53 as a transcription factor, in part by promoting acetylation within the C-terminus of p53 (Lambert et al., 1998; Dumaz and Meek, 1999).
Oncogene activation
One of the key mediators of oncogene-induced activation of p53 is p14ARF(ARF) (recently reviewed in Sherr, 2006). ARF, the alternate reading frame product of the p16/INK4A locus, has been shown to have a variety of functions, with one of its key roles being the binding and inhibition of Mdm2. Loss of ARF can, to a large degree, substitute for loss of p53 in tumour development, suggesting that disabling this pathway of p53 activation is critical for cancer progression (Sherr, 2006). Interestingly, loss of ARF does not prevent the activation of p53 in response to DNA damage, indicating that these are independent – although clearly complementary and cooperative – signals to p53 (Stott et al., 1998; Kamijo et al., 1999).
Ribosomal stress
A number of ribosomal proteins, in particular L5, L11 and L23, have been shown to bind and inhibit Mdm2, thereby stabilizing and activating p53 (Lohrum et al., 2003; Zhang et al., 2003; Dai and Lu, 2004; Dai et al., 2004). Indeed, treating cells with siRNA for L5 or L11 abrogates the induction of p53 in response to ribosomal stress, such as treatment with the RNA PolI inhibitor Actinomycin D (Bhat et al., 2004; Dai and Lu, 2004). Physiological stimuli that have been shown to signal through the ribosomal stress pathway include serum starvation and cell–cell contact growth inhibition (Bhat et al., 2004). Furthermore, inactivation of the transcription initiation factor 1A (TIF-IA), which is responsible for regulating the transcription of ribosomal RNA, induced binding of ribosomal proteins, including L11, to Mdm2, and the activation of p53 (Yuan et al., 2005). Interestingly, the DNA damage signal to p53 also depends on nucleolar disruption (Rubbi and Milner, 2003), suggesting a cooperation between kinases such as ATM/ATR and the ribosomal proteins. Ribosomal stress also enhances Mdm2-mediated degradation of MdmX, thereby releasing p53 from the negative regulation exerted by both proteins (Gilkes et al., 2006). While somewhat surprising, a clear role for ribosomal proteins in tumour suppressor pathways has also been established in zebrafish (Amsterdam et al., 2004). The importance of these interactions in humans is also strongly suggested by the observation that cancer associated mutations in Mdm2 can prevent the interaction with L11 and L5 (Lindstrom et al., 2006).
Cell–cell contacts
Another DNA-damage independent stimulus that leads to the activation of p53 is the loss of cell–matrix adhesion (reviewed in Grossmann (2002)). Loss of cell–cell or cell–matrix contact occurs in migrating cells, especially when they enter the blood or lymph systems. In non-transformed cells, the loss of adhesion can trigger a form of apoptosis known as anoikis, which in some cell types has been shown to be p53 dependent. The ability to overcome anoikis may be an important step in tumour progression towards metastasis, and the loss of p53 is critical for this step in some cell types. The PTEN phosphatase, a target of p53, appears to play an important role, as the expression of PTEN in a p53 null background can restore the sensitivity to anoikis (Lu et al., 1999). While the pathways leading to the activation of p53 in response to loss of adhesion are not clear, signalling through integrins can provide adhesion-associated survival signals to prevent the activation of p53 and cell death (Wang et al., 2002).
Hypoxia
Cells at the centre of expanding tumours often experience hypoxic conditions, due to the lack of angiogenesis to provide the ever increasing mass of cells with oxygen. Hypoxia activates p53 and stimulates p53-dependent apoptosis through mechanisms that – although not clear – appear to involve ATR, HIF 1α (hypoxia inducible factor) and VHL (von Hippel-Lindau) (reviewed in Hammond and Giaccia, 2005). Under normoxic conditions, HIF 1α is degraded through its interaction with VHL (Maxwell et al., 1999; Semenza, 2001). Hypoxic or anoxic conditions promote the stabilization of HIF 1α by disrupting its interaction with VHL, and HIF 1α is able to bind p53 and promote its stabilization. Furthermore, VHL can also interact with p53 directly and promote p53 phosphorylation and acetylation, leading to the activation (Roe et al., 2006). Interestingly, p53 induced by hypoxia shows a different pattern of target gene activation than seen following DNA damage (reviewed in Hammond and Giaccia (2005), Krieg et al. (2006)), suggesting that p53 may mediate a quite different response to hypoxia compared to other stress signals.
All signals are not equal
The realization that different stress signals can activate p53 through different pathways raises the question – are all these pathways equally important for tumour suppression? The DNA damage pathway has been shown to be constitutively active in tumours (DiTullio et al., 2002), with evidence that the DNA damage response pathways are activated even before cells become tumorigenic. The resultant activation of p53 in early hyperplastic and pre-cancerous lesions appears to prevent the formation of tumours (Bartkova et al., 2005; Gorgoulis et al., 2005), highlighting the importance of DNA damage pathways.
However, recent studies have also suggested the signalling through the ARF pathway – which is required to respond to oncogene activation but not DNA damage – is the response that is critical for tumour suppression. These studies suggest that while the DNA damage-induced activation of p53 certainly produces a strong response, this is not necessary for tumour suppression, which requires ARF and is therefore more likely to reflect the induction of p53 in response to oncogene activation (Christophorou et al., 2006; Efeyan et al., 2006) (Figure 2). Is it possible that activation of p53 by acute genotoxic stress is only deleterious to health, leading to radiation sickness that might be avoided in the absence of functional p53? p53 may become beneficial only once most of the damage has been resolved, to drive elimination of the rare cells that have acquired an oncogenic mutation. Further studies of the relative contribution of each of the p53 activating signals will be very revealing.
Activation and response to p53. p53 is activated by many stress signals that can differ in type and severity, which can lead to the activation of different responses, not all of which are clearly beneficial to the organism. Induction of cell death or senescence in response to severe DNA damage or oncogene activation are thought to be amongst the most important mechanisms of p53-mediated tumour suppression. While there is evidence that DNA damage signalling in premalignancies activates p53 and contributes to tumour suppression, other work has suggested that the response to acute genotoxic stress results predominantly in toxicity, and that the response to oncogene activation plays the most important role in tumour suppression. In addition to these two acute insults, p53 responds to a variety of milder, constitutive stresses – such as the generation of ROS by normal metabolism – and under these conditions p53 functions to allow survival and repair of the damage, and so may even contribute to the longevity of the organism.
Regulation of p53 stability and cancer therapy
A large number of human tumours retain wild-type p53, and many of these show evidence of perturbations in the pathways that allow stabilization and activation of p53 in response to stress. These pathways present attractive targets for developing treatments that could restore p53 function in these tumours. Examples of these include drugs that might inhibit the interaction between p53 and Mdm2, such as Nutlin-3 (Vassilev et al., 2004) or RITA (Issaeva et al., 2004). While there is some controversy concerning the mechanisms of function of RITA (Krajewski et al., 2005), both compounds allow for the stabilization of active p53 and show promising results in animal tumour models. Interestingly, Nutlin-3 does not efficiently block the interaction between p53 and MdmX, resulting in inefficient activation of p53 in tumours overexpressing MdmX (Hu et al., 2006; Patton et al., 2006). However, other studies have suggested that in some cells, Nutlin-3 treatment can result in the degradation of MdmX (Wade et al., 2006) – possibly through the action of Mdm2 released from p53 – and so the utility of Nutlin-3 as a therapy is likely to be strongly context dependent.
Other approaches to reactivating p53 have included directly targeting the Mdm2 E3 ligase activity (Yang et al., 2005); and the direct targeting of MdmX or HAUSP may also prove fruitful. It is clear that as the secrets of p53 regulation are uncovered, more potential targets for the design of novel therapies will be revealed. It seems likely that the next 40 000 reports on p53 will be as exciting as the first.
References
Abida WM, Nikolaev A, Zhao W, Zhang W, Gu W . (2006). FBXO11 promotes the neddylation of p53 and inhibits its transcriptional activity. J Biol Chem [Epub ahead of print].
Adhikary S, Marinoni F, Hock A, Hulleman E, Popov N, Beier R et al. (2005). The ubiquitin ligase HectH9 regulates transcriptional activation by Myc and is essential for tumor cell proliferation. Cell 123: 409–421.
Amsterdam A, Sadler KC, Lai K, Farrington S, Bronson RT, Lees JA et al. (2004). Many ribosomal protein genes are cancer genes in zebrafish. PLoS Biol 2: E139.
Andrews P, He YJ, Xiong Y . (2006). Cytoplasmic localized ubiquitin ligase cullin 7 binds to p53 and promotes cell growth by antagonizing p53 function. Oncogene 25: 4534–4548.
Asher G, Lotem J, Cohen B, Sachs L, Shaul Y . (2001). Regulation of p53 stability and p53-dependent apoptosis by NADH quinone oxidoreductase 1. Proc Natl Acad Sci USA 98: 1188–1193.
Asher G, Shaul Y . (2005). p53 proteasomal degradation: poly-ubiquitination is not the whole story. Cell Cycle 4: 1015–1018.
Banin S, Moyal L, Shieh S-Y, Taya Y, Anderson CW, Chessa L et al. (1998). Enhanced phosphorylation of p53 by ATM in response to DNA damage. Science 281: 1674–1677.
Bartkova J, Horejsi Z, Koed K, Kramer A, Tort F, Zieger K et al. (2005). DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434: 864–870.
Bensaad K, Tsuruta A, Selak MA, Vidal MN, Nakano K, Bartrons R et al. (2006). TIGAR, a p53-inducible regulator of glycolysis and apoptosis. Cell 126: 107–120.
Bensaad K, Vousden KH . (2005). Savior and slayer: the two faces of p53. Nat Med 11: 1278–1279.
Bhat KP, Itahana K, Jin A, Zhang Y . (2004). Essential role of ribosomal protein L11 in mediating growth inhibition-induced p53 activation. EMBO J 23: 2402–2412.
Bischof O, Schwamborn K, Martin N, Werner A, Sustmann C, Grosschedl R et al. (2006). The E3 SUMO ligase PIASy is a regulator of cellular senescence and apoptosis. Mol Cell 22: 783–794.
Bode AM, Dong Z . (2004). Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer 4: 793–805.
Bond GL, Hu W, Bond EE, Robins H, Lutzker SG, Arva NC et al. (2004). A single nucleotide polymorphism in the MDM2 promoter attenuates the p53 tumor suppressor pathway and accelerates tumor formation in humans. Cell 119: 591–602.
Boyd SD, Tsai KY, Jacks T . (2000). An intact HDM2 RING-finger domain is required for nuclear exclusion of p53. Nature Cell Biol 2: 563–568.
Brady M, Vlatkovic N, Boyd MT . (2005). Regulation of p53 and MDM2 activity by MTBP. Mol Cell Biol 25: 545–553.
Budanov AV, Sablina AA, Feinstein E, Koonin EV, Chumakov PM . (2004). Regeneration of peroxiredoxins by p53-regulated sestrins, homologs of bacterial AhpD. Science 304: 596–600.
Bykov VJ, Wiman KG . (2003). Novel cancer therapy by reactivation of the p53 apoptosis pathway. Ann Med 35: 458–465.
Chen C, Sun X, Guo P, Dong XY, Sethi P, Cheng X et al. (2005a). Human Kruppel-like factor 5 is a target of the E3 ubiquitin ligase WWP1 for proteolysis in epithelial cells. J Biol Chem 280: 41553–41561.
Chen D, Kon N, Li M, Zhang W, Qin J, Gu W . (2005b). ARF-BP1/Mule is a critical mediator of the ARF tumor suppressor. Cell 121: 1071–1083.
Cheng TH, Cohen SN . (2007). Human MDM2 isoforms translated differentially on constitutive vs p53-regulated transcripts have distinct functions in the p53/MDM2 and TSG101/MDM2 feedback control loops. Mol Cell Biol 27: 111–119.
Chipuk JE, Bouchier-Hayes L, Kuwana T, Newmeyer DD, Green DR . (2005). PUMA couples the nuclear and cytoplasmic proapoptotic function of p53. Science 309: 1732–1735.
Chipuk JE, Kuwana T, Bouchier-Hayes L, Droin NM, Newmeyer DD, Schuler M et al. (2004). Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science 303: 1010–1014.
Christophorou MA, Ringhausen I, Finch AJ, Brown Swigart L, Evan GI . (2006). The pathological response to DNA damage does not contribute to p53-mediated tumour suppression. Nature 14: 214–217.
Crighton D, Wilkinson S, O'Prey J, Syed N, Harrison PR, Gasco M et al. (2006). DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell 14: 121–134.
Cummins JM, Vogelstein B . (2004). HAUSP is required for p53 destabilization. Cell Cycle 3: 689–692.
Dai MS, Lu H . (2004). Inhibition of MDM2-mediated p53 ubiquitination and degradation by ribosomal protein L5. J Biol Chem 279: 44475–44482.
Dai MS, Zeng SX, Jin Y, Sun XX, David L, Lu H . (2004). Ribosomal protein L23 activates p53 by inhibiting MDM2 function in response to ribosomal perturbation but not to translation inhibition. Mol Biol Cell 24: 7654–7668.
Danial NN, Korsmeyer SJ . (2004). Cell death: critical control points. Cell 116: 205–219.
Dawson S, Apcher S, Mee M, Higashitsuji H, Baker R, Uhle S et al. (2002). Gankyrin is an ankyrin-repeat oncoprotein that interacts with CDK4 kinase and the S6 ATPase of the 26 S proteasome. J Biol Chem 277: 10893–10902.
de Graaf P, Little NA, Ramos YF, Meulmeester E, Letteboer SJ, Jochemsen AG . (2003). Hdmx protein stability is regulated by the ubiquitin ligase activity of Mdm2. J Biol Chem 278: 38315–38324.
de Toledo SM, Azzam EI, Dahlberg WK, Gooding TB, Little JB . (2000). ATM complexes with HDM2 and promotes its rapid phosphorylation in a p53-independent manner in normal and tumor human cells exposed to ionizing radiation. Oncogene 19: 6185–6193.
DiTullio Jr RA, Mochan TA, Venere M, Bartkova J, Sehested M, Bartek J et al. (2002). 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer. Nat Cell Biol 4: 998–1002.
Donehower LA, Harvey M, Slagle BL, McArthur MJ, Montgomery Jr CA, Butel JS et al. (1992). Mice deficient for p53 are developmentally normal but susceptible to spontaneous tumours. Nature 356: 215–221.
Dornan D, Shimizu H, Mah A, Dudhela T, Eby M, O'Rourke K et al. (2006). ATM engages autodegradation of the E3 ubiquitin ligase COP1 after DNA damage. Science 313: 1122–1126.
Dornan D, Wertz I, Shimizu H, Arnott D, Frantz GD, Dowd P et al. (2004). The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 429: 86–92.
Dumaz N, Meek DW . (1999). Serine15 phosphorylation stimulates p53 transactivation but does not directly influence interaction with HDM2. EMBO J 18: 7002–7010.
Efeyan A, Garcia-Cao I, Herranz D, Velasco-Miguel S, Serrano M . (2006). Tumour biology: policing of oncogene activity by p53. Nature 443: 159.
El-Deiry WS, Harper JW, Oconnor PM, Velculescu VE, Canman CE, Jackman J et al. (1994). WAF1/CIP1 is induced in p53-mediated G1 arrest and apoptosis. Cancer Research 54: 1169–1174.
Esser C, Scheffner M, Hohfeld J . (2005). The chaperone-associated ubiquitin ligase CHIP is able to target p53 for proteasomal degradation. J Biol Chem 280: 27443–27448.
Fang S, Jensen JP, Ludwig RL, Vousden KH, Weissman AM . (2000). Mdm2 is a RING finger-dependent ubiquitin protein ligase for itself and p53. J Biol Chem 275: 8945–8951.
Francoz S, Froment P, Bogaerts S, De Clercq S, Maetens M, Doumont G et al. (2006). Mdm4 and Mdm2 cooperate to inhibit p53 activity in proliferating and quiescent cells in vivo. Proc Natl Acad Sci USA 103: 3232–3237.
Fu XY, Wang HY, Tan L, Liu SQ, Cao HF, Wu MC . (2002). Overexpression of p28/gankyrin in human hepatocellular carcinoma and its clinical significance. World J Gastroenterol 8: 638–643.
Garcia-Cao I, Garcia-Cao M, Martin-Caballero J, Criado LM, Klatt P, Flores JM et al. (2002). ‘Super p53’ mice exhibit enhanced DNA damage response, are tumor resistant and age normally. EMBO J 21: 6225–6235.
Garcia-Cao I, Garcia-Cao M, Tomas-Loba A, Martin-Caballero J, Flores JM, Klatt P et al. (2006). Increased p53 activity does not accelerate telomere-driven ageing. EMBO Rep 7: 546–552.
Gatz SA, Wiesmuller L . (2006). p53 in recombination and repair. Cell Death Diff 13: 1003–1016.
Gilkes DM, Chen L, Chen J . (2006). MDMX regulation of p53 response to ribosomal stress. EMBO J 25: 5614–5625.
Gorgoulis VG, Vassiliou LV, Karakaidos P, Zacharatos P, Kotsinas A, Liloglou T et al. (2005). Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434: 907–913.
Gronroos E, Terentiev AA, Punga T, Ericsson J . (2004). YY1 inhibits the activation of the p53 tumor suppressor in response to genotoxic stress. Proc Natl Acad Sci USA 101: 12165–12170.
Grossman SR . (2001). p300/CBP/p53 interaction and regulation of the p53 response. Eur J Biochem 268: 2773–2778.
Grossman SR, Deato ME, Brignone C, Chan HM, Kung AL, Tagami H et al. (2003). Polyubiquitination of p53 by a ubiquitin ligase activity of p300. Science 300: 342–344.
Grossmann J . (2002). Molecular mechanisms of ‘detachment-induced apoptosis – Anoikis’. Apoptosis 7: 247–260.
Gu J, Kawai H, Nie L, Kitao H, Wiederschain D, Jochemsen AG et al. (2002). Mutual dependence of MDM2 and MDMX in their functional inactivation of p53. J Biol Chem 277: 19251–19254.
Gu J, Nie N, Wiederschain D, Yuan Z-M . (2001). Identification of p53 sequence elements that are required for MDM2-mediated nuclear export. Mol Cell Biol 21: 8533–8546.
Hainaut P, Hollstein M . (2000). p53 and human cancer: the first ten thousand mutations. Adv Cancer Res 77: 81–137.
Hammond EM, Giaccia AJ . (2005). The role of p53 in hypoxia-induced apoptosis. Biochem Biophys Res Commun 331: 718–725.
Higashitsuji H, Higashitsuji H, Itoh K, Sakurai T, Nagao T, Sumitomo Y et al. (2005a). The oncoprotein gankyrin binds to MDM2/HDM2, enhancing ubiquitylation and degradation of p53. Cancer Cell 8: 75–87.
Higashitsuji H, Liu Y, Mayer RJ, Fujita J . (2005b). The oncoprotein gankyrin negatively regulates both p53 and RB by enhancing proteasomal degradation. Cell Cycle 4: 1335–1337.
Hu B, Gilkes DM, Farooqi B, Sebti SM, Chen J . (2006). MDMX overexpression prevents p53 activation by the MDM2 inhibitor Nutlin. J Biol Chem 281: 33030–33035.
Issaeva N, Bozko P, Enge M, Protopopova M, Verhoef LG, Masucci M et al. (2004). Small molecule RITA binds to p53, blocks p53-HDM-2 interaction and activates p53 function in tumors. Nat Med 10: 1321–1328.
Iwakuma T, Lozano G . (2003). MDM2, an introduction. Mol Cancer Res 1: 933–1000.
Jacks T, Remington L, Williams BO, Schmitt EM, Halachmi S, Bronson RT et al. (1994). Tumor spectrum analysis in p53-mutant mice. Current Biol 4: 1–7.
Jackson MW, Berberich SJ . (2000). MdmX protects p53 from Mdm2-mediated degradation. Mol Cell Biol 20: 1001–1007.
Jeffers JR, Parganas E, Lee Y, Yang C, Wang J, Brennan J et al. (2003). Puma is an essential mediator of p53-dependent and -independent apoptotic pathways. Cancer Cell 4: 321–328.
Jones SN, Roe AE, Donehower LA, Bradley A . (1995). Rescue of embryonic lethality in Mdm2-deficient mice by absence of p53. Nature 378: 206–208.
Kahyo T, Nishida T, Yasuda H . (2001). Involvement of PIAS1 in the sumoylation of tumor suppressor p53. Mol Cell 8: 713–718.
Kamijo T, van de Kamp E, Chong MJ, Zindy F, Diehl JA, Sherr CJ et al. (1999). Loss of the ARF tumor suppressor reverses premature replicative arrest but not radiation hypersensitivity arising from disabled atm function. Cancer Res 59: 2464–2469.
Kawai H, Wiederschain D, Kitao H, Stuart J, Tsai KK, Yuan ZM . (2003). DNA damage-induced MDMX degradation is mediated by MDM2. J Biol Chem 278: 45946–45953.
Khosravi R, Maya R, Gottlieb T, Oren M, Shiloh Y, Shkedy D . (1999). Rapid ATM-dependent phosphorylation of MDM2 precedes p53 accumulation in response to DNA damage. Proc Natl Acad Sci USA 96: 14973–14977.
Krajewski M, Ozdowy P, D'Silva L, Rothweiler U, Holak TA . (2005). NMR indicates that the small molecule RITA does not block p53-MDM2 binding in vitro. Nat Med 11: 1135–1136; author reply 1136–1137.
Krieg AJ, Hammond EM, Giaccia AJ . (2006). Functional analysis of p53 binding under differential stresses. Mol Cell Biol 26: 7030–7045.
Laine A, Ronai Z . (2006). Regulation of p53 localization and transcription by the HECT domain E3 ligase WWP1. Oncogene [Epub ahead of print].
Laine A, Topisirovic I, Zhai D, Reed JC, Borden KL, Ronai Z . (2006). Regulation of p53 localization and activity by Ubc13. Mol Cell Biol 26: 8901–8913.
Lambert PF, Kashanchi F, Radonovich MF, Shiekhattar R, Brady JN . (1998). Phosphorylation of p53 serine 15 increases interaction with CBP. J Biol Chem 273: 33048–33053.
Le Cam L, Linares LK, Paul C, Julien E, Lacroix M, Hatchi E et al. (2006). E4F1 is an atypical ubiquitin ligase that modulates p53 effector functions independently of degradation. Cell 127: 775–788.
Lee MH, Lee SW, Lee EJ, Choi SJ, Chung SS, Lee JI et al. (2006). SUMO-specific protease SUSP4 positively regulates p53 by promoting Mdm2 self-ubiquitination. Nat Cell Biol 8: 1424–1431.
Leng RP, Lin Y, Ma W, Wu H, Lemmers B, Chung S et al. (2003). Pirh2, a p53-induced ubiquitin-protein ligase, promotes p53 degradation. Cell 112: 779–791.
Leu JI, Dumont P, Hafey M, Murphy MP, George DL . (2004). Mitochondrial p53 activates Bak and causes disruption of a Bak-Mcl1 complex. Nature Cell Biol 6: 443–450.
Li M, Brooks CL, Kon N, Gu W . (2004). A dynamic role of HAUSP in the p53-Mdm2 pathway. Mol Cell 13: 879–886.
Li M, Brooks CL, Wu-Baer F, Chen D, Baer R, Gu W . (2003). Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 302: 1972–1975.
Li M, Chen D, Shiloh A, Luo J, Nikolaev AY, Qin J et al. (2002). Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature 416: 648–652.
Li T, Santockyte R, Shen RF, Tekle E, Wang G, Yang DC et al. (2006). Expression of SUMO-2/3 induced senescence through p53- and pRB-mediated pathways. J Biol Chem 281: 36221–36227.
Lindstrom MS, Jin A, Deisenroth C, Wolf GW, Zhang Y . (2006). Cancer-associated mutations in MDM2 zinc finger domain disrupt ribosomal protein interaction and attenuate MDM2-Induced p53 degradation. Mol Cell Biol [Epub ahead of print].
Logan IR, Gaughan L, McCracken SR, Sapountzi V, Leung HY, Robson CN . (2006). Human PIRH2 enhances androgen receptor signaling through inhibition of histone deacetylase 1 and is overexpressed in prostate cancer. Mol Cell Biol 26: 6502–6510.
Lohrum MA, Ludwig RL, Kubbutat MHG, Hanlon M, Vousden KH . (2003). Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 3: 577–587.
Lohrum MAE, Woods DB, Ludwig RL, Bálint E, Vousden KH . (2001). C-terminal ubiquitination of p53 contributes to nuclear export. Mol Cell Biol 21: 8521–8532.
Longworth MS, Laimins LA . (2004). Pathogenesis of human papillomaviruses in differentiating epithelia. Microbiol Mol Biol Rev 68: 362–372.
Lu Y, Lin YZ, LaPushin R, Cuevas B, Fang X, Yu SX et al. (1999). The PTEN/MMAC1/TEP tumor suppressor gene decreases cell growth and induces apoptosis and anoikis in breast cancer cells. Oncogene 18: 7034–7045.
Maier B, Gluba W, Bernier B, Turner T, Mohammad K, Guise T et al. (2004). Modulation of mammalian life span by the short isoform of p53. Genes Dev 18: 306–319.
Malkin D, Li FP, Strong LC, Fraumini Jr JF, Nelson CE, Kim DH et al. (1990). Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science 250: 1233–1238.
Mani A, Oh AS, Bowden ET, Lahusen T, Lorick KL, Weissman AM et al. (2006). E6AP mediates regulated proteasomal degradation of the nuclear receptor coactivator amplified in breast cancer 1 in immortalized cells. Cancer Res 66: 8680–8686.
Marine JC, Jochemsen AG . (2005). Mdmx as an essential regulator of p53 activity. Biochem Biophys Res Commun 331: 750–760.
Maxwell PH, Wiesener MS, Chang GW, Clifford SC, Vaux EC, Cockman ME et al. (1999). The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399: 271–275.
Maya R, Balass M, Kim ST, Shkedy D, Leal JF, Shifman O et al. (2001). ATM-dependent phosphorylation of Mdm2 on serine 395: role in p53 activation by DNA damage. Genes Dev 15: 1067–1077.
McDonald III ER, El-Deiry WS . (2004). Suppression of caspase-8- and -10-associated RING proteins results in sensitization to death ligands and inhibition of tumor cell growth. Proc Natl Acad Sci USA 101: 6170–6175.
McDonough H, Patterson C . (2003). CHIP: a link between the chaperone and proteasome systems. Cell Stress Chaperones 8: 303–308.
Megidish T, Xu JH, Xu CW . (2002). Activation of p53 by protein inhibitor of activated Stat1 (PIAS1). J Biol Chem 277: 8255–8259.
Melchior F, Hengst L . (2002). SUMO-1 and p53. Cell Cycle 1: 245–249.
Mendrysa SM, O'Leary KA, McElwee MK, Michalowski J, Eisenman RN, Powell DA et al. (2006). Tumor suppression and normal aging in mice with constitutively high p53 activity. Genes Dev 20: 16–21.
Mendrysa SM, Perry ME . (2006). Tumor suppression by p53 without accelerated aging: just enough of a good thing? Cell Cycle 5: 714–717.
Meulmeester E, Maurice MM, Boutell C, Teunisse AFAS, Ovaa H, Abraham TE et al. (2005a). Loss of HAUSP-mediated deubiquitination contributes to DNA damage-induced destabilization of Hdmx and Hdm2. Mol Cell 18: 565–576.
Meulmeester E, Pereg Y, Shiloh Y, Jochemsen AG . (2005b). ATM-mediated phosphorylations inhibit Mdmx/Mdm2 stabilization by HAUSP in favor of p53 activation. Cell Cycle 4: 1166–1170.
Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P et al. (2003). p53 has a direct apoptogenic role at the mitochondria. Mol Cell 11: 577–590.
Moll UM, LaQuaglia M, Benard J, Riou G . (1995). Wild-type p53 protein undergoes cytoplasmic sequestration in undifferentiated neuroblastomas but not in differentiated tumors. Proc Natl Acad Sci USA 92: 4407–4411.
Moll UM, Marchenko N, Zhang XK . (2006). p53 and Nur77/TR3 – transcription factors that directly target mitochondria for cell death induction. Oncogene 25: 4725–4743.
Moll UM, Riou G, Levine AJ . (1992). Two distinct mechanisms alter p53 in breast cancer: mutation and nuclear exclusion. Proc Natl Acad Sci USA 89: 7262–7266.
Montes de Oca Luna R, Wagner DS, Lozano G . (1995). Rescue of early embryonic lethality in mdm2-deficient mice by deletion of p53. Nature 378: 203–206.
Moren A, Imamura T, Miyazono K, Heldin CH, Moustakas A . (2005). Degradation of the tumor suppressor Smad4 by WW and HECT domain ubiquitin ligases. J Biol Chem 280: 22115–22123.
Nagao T, Higashitsuji H, Nonoguchi K, Sakurai T, Dawson S, Mayer RJ et al. (2003). MAGE-A4 interacts with the liver oncoprotein gankyrin and suppresses its tumorigenic activity. J Biol Chem 278: 10668–10674.
Nakano K, Vousden KH . (2001). PUMA, a novel proapoptotic gene, is induced by p53. Mol Cell 7: 683–694.
Nelson V, Davis GE, Maxwell SA . (2001). A putative protein inhibitor of activated STAT (PIASy) interacts with p53 and inhibits p53-mediated transactivation but not apoptosis. Apoptosis 6: 221–234.
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T et al. (2000). Noxa, a BH3-only member of the Bcl-2 family and candidate mediator of p53-induced apoptosis. Science 288: 1053–1058.
Pan Y, Chen J . (2003). MDM2 promotes ubiquitination and degradation of MDMX. Mol Cell Biol 23: 5113–5121.
Patton JT, Mayo LD, Singhi AD, Gudkov AV, Stark GR, Jackson MW . (2006). Levels of HdmX expression dictate the sensitivity of normal and transformed cells to Nutlin-3. Cancer Res 66: 3169–3176.
Pietsch EC, Humbey O, Murphey ME . (2006). Polymorphisms in the p53 pathway. Oncogene 25: 1602–1611.
Pim D, Banks L . (2004). p53 polymorphic variants at codon 72 exert different effects on cell cycle progression. Int J Cancer 108: 196–199.
Poyurovsky MV, Priest C, Kentsis A, Borden KL, Pan ZQ, Pavletich N et al. (2006). EMBO J [Epub ahead of print].
Poyurovsky MV, Prives C . (2006). Unleashing the power of p53: lessons from mice and men. Genes Dev 20: 125–131.
Purdie CA, Harrison DJ, Peter A, Dobbie L, White S, Howie SE et al. (1994). Tumour incidence, spectrum and ploidy in mice with a large deletion in the p53 gene. Oncogene 9: 603–609.
Qi L, Heredia JE, Altarejos JY, Screaton R, Goebel N, Niessen S et al. (2006). TRB3 links the E3 ubiquitin ligase COP1 to lipid metabolism. Science 312: 1763–1766.
Rajendra R, Malegaonkar D, Pungaliya P, Marshall H, Rasheed Z, Brownell J et al. (2004). Topors functions as an E3 ubiquitin ligase with specific E2 enzymes and ubiquitinates p53. J Biol Chem 279: 36440–36444.
Roe JS, Kim H, Lee SM, Kim ST, Cho EJ, Youn HD . (2006). p53 stabilization and transactivation by a von Hippel-Lindau protein. Mol Cell 22: 395–405.
Rubbi CP, Milner J . (2003). Disruption of the nucleolus mediates stabilization of p53 in response to DNA damage and other stresses. EMBO J 22: 6068–6077.
Sablina AA, Budanov AV, Ilyinskaya GV, Agapova LS, Kravchenko JE, Chumakov PM . (2005). The antioxidant function of the p53 tumor suppressor. Nat Med 11: 1306–1313.
Sakaguchi K, Saito S, Higashimoto Y, Roy S, Anderson CW, Appella E . (2000). Damage-mediated phosphorylation of human p53 threonine 18 through a cascade mediated by a casein 1-like kinase. Effect on Mdm2 binding. J Biol Chem 275: 9278–9283.
Salcedo A, Mayor Jr F, Penela P . (2006). Mdm2 is involved in the ubiquitination and degradation of G-protein-coupled receptor kinase 2. EMBO J 25: 4752–4762.
Schmidt D, Muller S . (2002). Members of the PIAS family act as SUMO ligases for c-Jun and p53 and repress p53 activity. Proc Natl Acad Sci USA 99: 2872–2877.
Secombe J, Parkhurst SM . (2004). Drosophila Topors is a RING finger-containing protein that functions as a ubiquitin-protein isopeptide ligase for the hairy basic helix-loop-helix repressor protein. J Biol Chem 279: 17126–17133.
Semenza GL . (2001). HIF-1, O(2), and the 3 PHDs: how animal cells signal hypoxia to the nucleus. Cell 107: 1–3.
Sharp DA, Kratowicz SA, Sank MJ, George DL . (1999). Stabilization of the MDM2 oncoprotein by interaction with the structurally related MDMX protein. J Biol Chem 274: 38189–38196.
Sherr CJ . (2006). Divorcing ARF and p53: an unsettled case. Nat Rev Cancer 6: 663–673.
Sherr CJ, Roberts JM . (1999). CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev 13: 1501–1512.
Shieh S-Y, Ikeda M, Taya Y, Prives C . (1997). DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell 91: 325–334.
Shvarts A, Steegenga WT, Riteco N, van Laar T, Dekker P, Bazuine M et al. (1996). MDMX: a novel p53-binding protein with some functional properties of MDM2. EMBO J 15: 5349–5357.
Siliciano JD, Canman CE, Taya Y, Sakaguchi K, Appella E, Kastan MB . (1997). DNA damage induces phosphorylation of the amino terminus of p53. Genes Dev 11: 3471–3481.
Srivastava S, Zou Z, Pirollo K, Blattner W, Chang EH . (1990). Germ-line transmission of a mutated p53 gene in a cancer-prone family with Li-Fraumeni syndrome. Nature 348: 747–749.
Stad R, Little NA, Xirodimas DP, Frenk R, van der Eb AJ, Lane DP et al. (2001). Mdmx stabilizes p53 and Mdm2 via two distinct mechanisms. EMBO Rep 2: 1029–1034.
Stott F, Bates SA, James M, McConnell BB, Starborg M, Brookes S et al. (1998). The alternative product from the human CDKN2A locus, p14(ARF), participates in a regulatory feedback loop with p53 and MDM2. EMBO J 17: 5001–5014.
Sui G, Affar el B, Shi Y, Brignone C, Wall NR, Yin P et al. (2004). Yin Yang 1 is a negative regulator of p53. Cell 117: 859–872.
Tang J, Qu LK, Zhang J, Wang W, Michaelson JS, Degenhardt YY et al. (2006). Critical role for Daxx in regulating Mdm2. Nat Cell Biol 8: 855–862.
Tanimura S, Ohtsuka S, Mitsui K, Shirouzu K, Yoshimura A, Ohtsubo M . (1999). MDM2 interacts with MDMX through their RING finger domains. FEBS Lett 447: 5–9.
Toledo F, Wahl GM . (2006). Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nat Rev Cancer 6: 909–923.
Tyner SD, Venkatachalam S, Choi J, Ghebranious N, Igelmann H, Lu X et al. (2002). p53 mutant mice that display early ageing-associated phenotypes. Nature 415: 45–53.
Uldrijan S, Pannekoek WJ, Vousden KH . (2006). EMBO J [Epub ahead of print].
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F, Filipovic Z et al. (2004). In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303: 844–848.
Villunger A, Michalak EM, Coultas L, Mullauer F, Bock G, Ausserlechner MJ et al. (2003). p53- and drug-induced apoptotic responses mediated by BH3-only proteins puma and noxa. Science 302: 1036–1038.
Vogelstein B, Lane D, Levine AJ . (2000). Surfing the p53 network. Nature 408: 307–310.
Wade M, Wong ET, Tang M, Stommel JM, Wahl GM . (2006). Hdmx modulates the outcome of p53 activation in human tumor cells. J Biol Chem 281: 33036–33044.
Wang WJ, Kuo JC, Yao CC, Chen RH . (2002). DAP-kinase induces apoptosis by suppressing integrin activity and disrupting matrix survival signals. J Cell Biol 159: 169–179.
Watson IR, Irwin MS . (2006). Ubiquitin and ubiquitin-like modifications of the p53 family. Neoplasia 8: 655–666.
Weissman AM . (2001). Themes and variations on ubiquitylation. Nat Rev Mol Cell Biol 2: 169–178.
Xiong S, Van Pelt CS, Elizondo-Fraire AC, Liu G, Lozano G . (2006). Synergistic roles of Mdm2 and Mdm4 for p53 inhibition in central nervous system development. Proc Natl Acad Sci USA 103: 3226–3231.
Xirodimas DP, Saville MK, Bourdon JC, Hay RT, Lane DP . (2004). Mdm2-mediated NEDD8 conjugation of p53 inhibits its transcriptional activity. Cell 118: 83–97.
Yang JY, Zong CS, Xia W, Wei Y, Ali-Seyed M, Li Z et al. (2006a). MDM2 promotes cell motility and invasiveness by regulating E-cadherin degradation. Mol Cell Biol 26: 7269–7282.
Yang W, Rozan LM, McDonald III ER, Navaraj A, Liu JJ, Matthew EM et al. (2006b). Carps are ubiquitin ligases that promote MDM2-independent P53 and phospho-P53ser20 degradation. J Biol Chem [Epub ahead of print].
Yang WH, Kim JE, Nam HW, Ju JW, Kim HS, Kim YS et al. (2006c). Modification of p53 with O-linked N-acetylglucosamine regulates p53 activity and stability. Nat Cell Biol 8: 1074–1083.
Yang Y, Ludwig RL, Jensens JP, Pierre S, Medaglia MV, Davydov I et al. (2005). Small molecule inhibitors of HDM2 ubiquitin ligase activity stabilize and activate p53 in cells. Cancer Cell 7: 547–559.
Yee KS, Vousden KH . (2005). Complicating the complexity of p53. Carcinogenesis 26: 1317–1322.
Yoon KA, Nakamura Y, Arakawa H . (2004). Identification of ALDH4 as a p53-inducible gene and its protective role in cellular stresses. J Hum Genet 49: 134–140.
Yu J, Kinzler KW, Vogelstein B, Zhang L . (2003). PUMA mediates the apoptotic response to p53 in colorectal cancer cells. Proc Natl Acad Sci USA 100: 1931–1936.
Yu J, Zhang L, Hwang PM, Kinzler KW, Vogelstein B . (2001). PUMA induces the rapid apoptosis of colorectal cancer cells. Mol Cell 7: 673–682.
Yuan X, Zhou Y, Casanova E, Chai M, Kiss E, Grone HJ et al. (2005). Genetic inactivation of the transcription factor TIF-IA leads to nucleolar disruption, cell cycle arrest, and p53-mediated apoptosis. Mol Cell 19: 77–87.
Zauberman A, Barak Y, Ragimov N, Levy N, Oren M . (1993). Sequence-specific DNA binding by p53: identification of target sites and lack of binding to p53 – MDM2 complexes. EMBO J 12: 2799–2808.
Zhang X, Srinivasan SV, Lingrel JB . (2004). WWP1-dependent ubiquitination and degradation of the lung Kruppel-like factor, KLF2. Biochem Biophys Res Commun 316: 139–148.
Zhang Y, Wolf GW, Bhat K, Jin A, Allio T, Burkhart WA et al. (2003). Ribosomal protein L11 negatively regulates oncoprotein MDM2 and mediates a p53-dependent ribosomal-stress checkpoint pathway. Mol Cell Biol 23: 8902–8912.
Zhang Z, Zhang R . (2005). p53-independent activities of MDM2 and their relevance to cancer therapy. Curr Cancer Drug Targets 5: 9–20.
Zhong Q, Gao W, Du F, Wang X . (2005). Mule/ARF-BP1, a BH3-only E3 ubiquitin ligase, catalyzes the polyubiquitination of Mcl-1 and regulates apoptosis. Cell 121: 1085–1095.
Acknowledgements
We are grateful to Cancer Research UK for support and funding.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Horn, H., Vousden, K. Coping with stress: multiple ways to activate p53. Oncogene 26, 1306–1316 (2007). https://doi.org/10.1038/sj.onc.1210263
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210263
Keywords
This article is cited by
-
Molecular mechanisms behind ROS regulation in cancer: A balancing act between augmented tumorigenesis and cell apoptosis
Archives of Toxicology (2023)
-
Of the many cellular responses activated by TP53, which ones are critical for tumour suppression?
Cell Death & Differentiation (2022)
-
Development of a novel lipid metabolism-based risk score model in hepatocellular carcinoma patients
BMC Gastroenterology (2021)
-
FBXL6 degrades phosphorylated p53 to promote tumor growth
Cell Death & Differentiation (2021)
-
Single loss of a Trp53 allele triggers an increased oxidative, DNA damage and cytokine inflammatory responses through deregulation of IκBα expression
Cell Death & Disease (2021)