Abstract
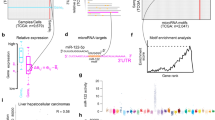
MicroRNAs (miRNAs) are ∼22 nucleotide-long noncoding RNAs involved in several biological processes including development, differentiation and proliferation. Recent studies suggest that knowledge of miRNA expression patterns in cancer may have substantial value for diagnostic and prognostic determinations as well as for eventual therapeutic intervention. We performed comprehensive analysis of miRNA expression profiles of 27 sarcomas, 5 normal smooth muscle and 2 normal skeletal muscle tissues using microarray technology and/or small RNA cloning approaches. The miRNA expression profiles are distinct among the tumor types as demonstrated by an unsupervised hierarchical clustering, and unique miRNA expression signatures were identified in each tumor class. Remarkably, the miRNA expression patterns suggested that two of the sarcomas had been misdiagnosed and this was confirmed by reevaluation of the tumors using histopathologic and molecular analyses. Using the cloning approach, we also identified 31 novel miRNAs or other small RNA effectors in the sarcomas and normal skeletal muscle tissues examined. Our data show that different histological types of sarcoma have distinct miRNA expression patterns, reflecting the apparent lineage and differentiation status of the tumors. The identification of unique miRNA signatures in each tumor type may indicate their role in tumorigenesis and may aid in diagnosis of soft tissue sarcomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ambros V, Bartel B, Bartel DP, Burge CB, Carrington JC, Chen X et al. (2003). A uniform system for microRNA annotation. RNA 9: 277–279.
Baer C, Nees M, Breit S, Selle B, Kulozik AE, Schaefer KL et al. (2004). Profiling and functional annotation of mRNA gene expression in pediatric rhabdomyosarcoma and Ewing's sarcoma. Int J Cancer 110: 687–694.
Binh MB, Sastre-Garau X, Guillou L, de Pinieux G, Terrier P, Lagace R et al. (2005). MDM2 and CDK4 immunostainings are useful adjuncts in diagnosing well-differentiated and dedifferentiated liposarcoma subtypes: a comparative analysis of 559 soft tissue neoplasms with genetic data. Am J Surg Pathol 29: 1340–1347.
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE et al. (2005). A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353: 1793–1801.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al. (2004). Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci USA 101: 2999–3004.
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM et al. (2006). The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 38: 228–233.
Doench JG, Sharp PA . (2004). Specificity of microRNA target selection in translational repression. Genes Dev 18: 504–511.
Esau C, Kang X, Peralta E, Hanson E, Marcusson EG, Ravichandran LV et al. (2004). MicroRNA-143 regulates adipocyte differentiation. J Biol Chem 279: 52361–52365.
Espinosa I, Lee CH, Kim M, Rouse B, Subramanian S, Montgomery K et al. (2007). A novel monoclonal antibody against DOG1 is a sensitive and specific marker for gastrointestinal stromal tumors. Am J Surg Path (in press).
Esquela-Kerscher A, Slack FJ . (2006). Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6: 259–269.
Felli N, Fontana L, Pelosi E, Botta R, Bonci D, Facchiano F et al. (2005). MicroRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc Natl Acad Sci USA 102: 18081–18086.
Fletcher JA, Rubin BP . (2007). KIT Mutations in GIST. Curr Opin Genet Dev 17: 3–7.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ . (2006). miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34: D140–D144.
He L, Hannon GJ . (2004). MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5: 522–531.
Heinrich MC, Corless CL, Duensing A, McGreevey L, Chen CJ, Joseph N et al. (2003). PDGFRA activating mutations in gastrointestinal stromal tumors. Science 299: 708–710.
Kim HK, Lee YS, Sivaprasad U, Malhotra A, Dutta A . (2006). Muscle-specific microRNA miR-206 promotes muscle differentiation. J Cell Biol 174: 677–687.
Kim VN . (2005). MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6: 376–385.
Lau NC, Lim LP, Weinstein EG, Bartel DP . (2001). An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294: 858–862.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. (2005). MicroRNA expression profiles classify human cancers. Nature 435: 834–838.
Lui WO, Pourmand N, Patterson BK, Fire A . (2007). Patterns of known and novel small RNAs in human cervical cancer. Cancer Res 67: 6031–6043.
Mayanil CS, George D, Freilich L, Miljan EJ, Mania-Farnell B, McLone DG et al. (2001). Microarray analysis detects novel Pax3 downstream target genes. J Biol Chem 276: 49299–49309.
Michael MZ, O'Connor SM, van Holst Pellekaan NG, Young GP, James RJ . (2003). Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1: 882–891.
Nielsen TO, West RB, Linn SC, Alter O, Knowling MA, O'Connell JX et al. (2002). Molecular characterisation of soft tissue tumours: a gene expression study. Lancet 359: 1301–1307.
Politz JC, Zhang F, Pederson T . (2006). MicroRNA-206 colocalizes with ribosome-rich regions in both the nucleolus and cytoplasm of rat myogenic cells. Proc Natl Acad Sci USA 103: 18957–18962.
Pretto D, Barco R, Rivera J, Neel N, Gustavson MD, Eid JE . (2006). The synovial sarcoma translocation protein SYT-SSX2 recruits beta-catenin to the nucleus and associates with it in an active complex. Oncogene 25: 3661–3669.
Shimada S, Ishizawa T, Ishizawa K, Matsumura T, Hasegawa T, Hirose T . (2006). The value of MDM2 and CDK4 amplification levels using real-time polymerase chain reaction for the differential diagnosis of liposarcomas and their histologic mimickers. Hum Pathol 37: 1123–1129.
Subramanian S, West RB, Corless CL, Ou W, Rubin BP, Chu KM et al. (2004). Gastrointestinal stromal tumors (GISTs) with KIT and PDGFRA mutations have distinct gene expression profiles. Oncogene 23: 7780–7790.
Weiss SW, Goldbum JR . (2001). Soft tissue tumors. Mosby: St Louis.
West RB, Corless CL, Chen X, Rubin BP, Subramanian S, Montgomery K et al. (2004). The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am J Pathol 165: 107–113.
Yan Z, Choi S, Liu X, Zhang M, Schageman JJ, Lee SY et al. (2003). Highly coordinated gene regulation in mouse skeletal muscle regeneration. J Biol Chem 278: 8826–8836.
Zuker M . (2003). Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31: 3406–3415.
Acknowledgements
This work was supported by NIH grant CA112270, a grant from LifeRaft and the Department of Pathology, Stanford University. SS was supported by a postdoctoral fellowship from the US National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Rights and permissions
About this article
Cite this article
Subramanian, S., Lui, W., Lee, C. et al. MicroRNA expression signature of human sarcomas. Oncogene 27, 2015–2026 (2008). https://doi.org/10.1038/sj.onc.1210836
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210836
Keywords
This article is cited by
-
Neoplastic Transformation of Human Mesenchymal Stromal Cells Mediated via LIN28B
Scientific Reports (2019)
-
PAX3-FOXO1 drives miR-486-5p and represses miR-221 contributing to pathogenesis of alveolar rhabdomyosarcoma
Oncogene (2018)
-
miR-34a exerts as a key regulator in the dedifferentiation of osteosarcoma via PAI-1–Sox2 axis
Cell Death & Disease (2018)
-
Circulating MicroRNA-92b-3p as a Novel Biomarker for Monitoring of Synovial Sarcoma
Scientific Reports (2017)
-
miR-152 down-regulation is associated with MET up-regulation in leiomyosarcoma and undifferentiated pleomorphic sarcoma
Cellular Oncology (2017)