Abstract
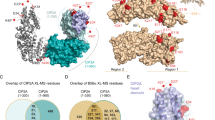
The cyclin-dependent kinases 4 and 6 (Cdk4/6) that control the G1 phase of the cell cycle and their inhibitor, the p16INK4a tumour suppressor, have a central role in cell proliferation and in tumorigenesis. The structures of Cdk6 bound to p16INK4a and to the related p19INK4d reveal that the INK4 inhibitors bind next to the ATP-binding site of the catalytic cleft, opposite where the activating cyclin subunit binds. They prevent cyclin binding indirectly by causing structural changes that propagate to the cyclin-binding site. The INK4 inhibitors also distort the kinase catalytic cleft and interfere with ATP binding, which explains how they can inhibit the preassembled Cdk4/6–cyclin D complexes as well. Tumour-derived mutations in INK4a and Cdk4 map to interface contacts, solidifying the role of CDK binding and inhibition in the tumour suppressor activity of p16INK4a.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Sherr, C. J. Cancer cell cycles. Science 274, 1672–1677 (1996).
Hall, M. & Peters, G. Genetic alterations of cyclin, cyclin-dependent kinases, and cdk inhibitors in human cancer. Adv. Cancer Res. 68, 68–108 (1996).
Weinber, R. A. The retinoblastoma protein and cell cycle control. Cell 81, 323–330 (1995).
Sherr, C. J. G1 phase progression cycling on cue. Cell 79, 551–555 (1994).
Hunter, T. & Pines, J. Cyclins and cancer II: cyclin D and cdk inhibitors come of age. Cell 79, 573–582 (1994).
Serrano, M., Hannon, G. J. & Beach, D. Anew regulatory motif in cell-cycle control causing specific inhibition of cyclin D/CDK4. Nature 366, 704–707 (1993).
Kamb, A. et al. Acell cycle regulator potentially involved in genesis of many tumor types. Science 264, 436–440 (1994).
Nobori, T. et al. Deletions of the cyclin-dependent kinase-4 inhibitor gene in multiple human cancers. Nature 368, 753–756 (1994).
Caldas, C. et al. Frequent somatic mutations and homozygous deletions of the p16 (MTS1) gene in pancreatic adenocarcinoma. Nature Genet. 8, 27–32 (1994).
Hussussian, C. J. et al. Germline p16 mutations in familial melanoma. Nature Genet. 8, 15–21 (1994).
Kamb, A. et al. Analysis of the p16 gene (CDKN2) as a candidate for the chromosome 9p melanoma susceptibility locus. Nature Genet. 8, 23–26 (1994).
Serrano, M. et al. Role of the INK4a locus in tumor suppression and cell mortality. Cell 85, 27–37 (1996).
Koh, J., Enders, G. H., Dynlacht, B. D. & Harlow, E. Tumour-derived p16 alleles encoding proteins defective in cell-cycle inhibition. Nature 375, 506–510 (1995).
Serrano, M., Gomez-Lahoz, E., DePinho, R. A., Beach, D. & Bar-Sagi, D. Inhibition of ras-induced proliferation and cellular transformation by p16INK4. Science 267, 249–252 (1995).
Yang, R., Gombart, A. F., Serrano, M. & Koeffler, H. P. Mutational effects on the p16INK4a tumor supressor protein. Cancer Res. 55, 2503–2506 (1995).
Ranade, K. et al. Mutations associated with familial melanoma impair p16INK4 function. Nature Genet. 10, 114–116 (1995).
Parry, D. & Peters, G. Temperature-sensitive mutants of p16CDKN2 associated with familial melanoma. Mol. Cell. Biol. 16, 3844–3852 (1996).
Zhang, B. & Peng, Z. Defective folding of mutant p16(INK4) proteins encoded by tumor-derived alleles. J. Biol. Chem. 271, 28734–28737 (1996).
Arap, W., Knudsen, E. S., Wang, J. Y., Cavenee, W. K. & Huang, H. J. Point mutations can inactivate in vitro and in vivo activities of p16(INK4a)/CDKN2A in human glioma. Oncogene 14, 603–609 (1997).
Wolfel, T. et al. Ap16INK4a-insensitive CDK4 mutant targeted by cytolytic T lymphocytes in a human melanoma. Science 269, 1281–1284 (1995).
Zuo, L. et al. Germline mutations in the p16INK4a binding domain of CDK4 in familial melanoma. Nature Genet. 12, 97–99 (1996).
Hannon, G. J. & Beach, D. p15INK4B is a potential effector of TGF β-induced cell cycle arrest. Nature 371, 257–261 (1994).
Hirai, H., Roussel, M. F., Kato, J. Y., Ashmun, R. A. & Sherr, C. J. Novel INK4 proteins, p19 and p18, are specific inhibitors of the cyclin D-dependent kinases CDK4 and CDK6. Mol. Cell. Biol. 15, 2672–2681 (1995).
Guan, K. L. et al. Isolation and characterization of p19INK4d, a p16-related inhibitor specific to CDK6 and CDK4. Mol. Biol. Cell 7, 57–70 (1996).
Xiong, Y., Zhang, H. & Beach, D. Subunit rearrangement of the cyclin-dependent kinases is associated with cellular transformation. Genes Dev. 7, 1572–1583 (1993).
Li, Y., Nichols, M. A., Shay, J. W. & Xiong, Y. Transcriptional repression of the D-type cyclin-dependent kinase inhibitor p16 by the retinoblastoma susceptibility gene product pRb. Cancer Res. 54, 6078–6082 (1994).
Parry, D., Bates, S., Mann, D. J. & Peters, G. Lack of cyclin D–Cdk complexes in Rb-negative cells correlates with high levels of p16INK4/MTS1 tumour suppressor gene product. EMBO J. 14, 503–511 (1995).
Lukas, J. et al. Retinoblastoma-protein-dependent cell-cycle inhibition by the tumour suppressor p16. Nature 375, 503–506 (1995).
Russo, A. A., Jeffrey, P. D., Patten, A. K., Massagué, J. & Pavletich, N. P. Crystal structure of the p27Kip1 cyclin-dependent-kinase inhibitor bound to the cyclin A–Cdk2 complex. Nature 382, 325–331 (1996).
De Bondt, H. L. et al. Crystal structure of cyclin-dependent kinase 2. Nature 363, 595–602 (1993).
Knighton, D. R. et al. Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 407–413 (1991).
Gorina, S. & Pavletich, N. P. Structure of the p53 tumor suppressor bound to the ankyrin and SH3 domains of 53BP2. Science 274, 1001–1005 (1996).
Luh, F. Y. et al. Structure of the cyclin-dependent kinase inhibitor p19Ink4d. Nature 389, 999–1003 (1997).
Byeon, I. et al. Tumor suppressor p16INK4A: determination of solution structure and analyses of its interaction with cyclin-dependent kinase 4. Mol. Cell 1, 421–431 (1998).
Venkataramani, R., Swaminathan, K. & Marmostein, R. Crystal structure of the CDK4/6 inhibitory protein p18INK4c provides insights into ankyrin-like repeat structure/function and tumor-derived p16INK4 mutations. Nature Struct. Biol. 5, 74–81 (1998).
Jeffrey, P. D. et al. Mechanism of CDK activation revealed by the structure of cyclin A–CDK2 ocmplex. Nature 376, 313–320 (1995).
Russo, A., Jeffrey, P. D. & Pavletich, N. P. Structural basis of cyclin-dependent kinase activation by phosphorylation. Nature Struct. Biol. 3, 696–700 (1996).
Coleman, K. G. et al. Identification of CDK4 sequences involved in cyclin D1 and p16 binding. J. Biol. Chem. 272, 18869–18874 (1997).
Brotherton, D. H. et al. Crystal structure of the complex of the cyclin D-dependent kinase Cdk6 bound to the cell cycle inhibitor p19INK4d. Nature 395, 244–250 (1998).
Zoller, M. J., Nelson, N. C. & Taylor, S. S. Affinity labeling of cAMP-dependent protein kinase with p-Fluorosulfonylbenzoyl adenosine. J. Biol. Chem. 256, 10837–10842 (1981).
Atherton-Fessler, S., Parker, L. L., Geahlen, R. L. & Piwnica-Worms, H. Mechanisms of p34cdc2 regualtion. Mol. Cell. Biol. 13, 1675–1685 (1993).
Fahraeus, R., Paramio, J. M., Ball, K. L., Lain, S. & Lane, D. P. Inhibition of pRb phosphorylation and cell-cycle progression by a 20-residue peptide derived from p16CDKN2/INK4A. Curr. Biol. 6, 84–91 (1996).
Gerber, M. R., Farrell, A., Deshaies, R. J., Herskowitz, I. & Morgan, D. O. Cdc37 is required for association of the protein kinase Cdc28 with G1 and mitotic cyclins. Proc. Natl Acad. Sci. USA 92, 4651–4655 (1995).
Stepanova, L., Leng, X., Parker, S. B. & Harper, J. W. Mammalian p50Cdc37 is a protein kinase-targeting subunit of Hsp90 that binds and stabilizes Cdk4. Genes Dev. 10, 1491–1502 (1996).
Reynisdottir, I., Polyak, K., Iavarone, A. & Massagué, J. Kip/Cip and Ink4 Cdk inhibitors cooperate to induce cell cycle arrest in response to TGF-beta. Genes Dev. 9, 1831–1845 (1995).
Meyerson, M. & Harlow, E. Identification of G1 kinase activity for cdk6, a novel cyclin D partner. Mol. Cell. Biol. 14, 2077–2086 (1994).
Collaborative Computational Project, Number4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A. T. X-PLOR, a System for Crystallography and NMR(Yale Univ. Press, New Haven, CT, 1991).
Kraulis, P. J. Molscript: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merrit, E. A. & Murphy, M. E. Raster3D Version 2.0: a program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Acknowledgements
We thank S. Geromanos and H. Erdjument-Bromage of the Sloan-Kettering Microchemistry Facility for N-terminal sequence and mass spectroscopic analyses; H. Chou and E. D. Harlow for baculovirus vectors expressing Cdk6, GST–Cdk6 and cyclin D1; R. Marmorstein for the crystallographic coordinates of p18; C. Ogata of the National Synchrotron Light Source X4A beam line and the staff of the Cornell High Energy Synchrotron Source MacChess for help with data collection; and C. Murray for administrative help. Supported by the Howard Hughes Medical Institute, the NIH, the Pew Charitable Trusts, the Dewitt Wallace Foundation, and the Samuel and May Rudin Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Russo, A., Tong, L., Lee, JO. et al. Structural basis for inhibition of the cyclin-dependent kinase Cdk6 by the tumour suppressor p16INK4a. Nature 395, 237–243 (1998). https://doi.org/10.1038/26155
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/26155
This article is cited by
-
Rejuvenating aged microglia by p16ink4a-siRNA-loaded nanoparticles increases amyloid-β clearance in animal models of Alzheimer’s disease
Molecular Neurodegeneration (2024)
-
ANKRD29, as a new prognostic and immunological biomarker of non–small cell lung cancer, inhibits cell growth and migration by regulating MAPK signaling pathway
Biology Direct (2023)
-
Disruptor: Computational identification of oncogenic mutants disrupting protein-protein and protein-DNA interactions
Communications Biology (2023)
-
Therapeutic potential of CDK4/6 inhibitors in renal cell carcinoma
Nature Reviews Urology (2022)
-
Mechanisms of Resistance to CDK4/6 Inhibitors in Hormone Receptor-Positive (HR +) Breast Cancer: Spotlight on Convergent CDK6 Upregulation and Immune Signaling
Current Breast Cancer Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.