Abstract
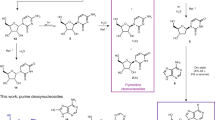
The ‘RNA world’ hypothesis proposes that early life developed by making use of RNA molecules, rather than proteins, to catalyse the synthesis of important biological molecules1. It is thought, however, that the nucleotides constituting RNA were scarce on early Earth1,2,3,4. RNA-based life must therefore have acquired the ability to synthesize RNA nucleotides from simpler and more readily available precursors, such as sugars and bases. Plausible prebiotic synthesis routes have been proposed for sugars5, sugar phosphates6 and the four RNA bases7,8,9,10,11, but the coupling of these molecules into nucleotides, specifically pyrimidine nucleotides, poses a challenge to the RNA world hypothesis1,2,3. Here we report the application of in vitro selection to isolate RNA molecules that catalyse the synthesis of a pyrimidine nucleotide at their 3′ terminus. The finding that RNA can catalyse this type of reaction, which is modelled after pyrimidine synthesis in contemporary metabolism, supports the idea of an RNA world that included nucleotide synthesis and other metabolic pathways mediated by ribozymes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Joyce, G. F. & Orgel, L. E. in The RNA World (eds Gesteland, R. F. & Atkins, J. F.) 1–25 (Cold Spring Harbor Lab., Cold Spring Harbor, NY, 1993).
Fuller, W. D., Sanchez, R. A. & Orgel, L. E. Studies in prebiotic synthesis VI. Synthesis of purine nucleosides. J. Mol. Biol. 67, 25–33 (1972).
Fuller, W. D., Sanchez, R. A. & Orgel, L. E. Studies in prebiotic synthesis VII. Solid-state synthesis of purine nucleosides. J. Mol. Evol. 1, 249–257 (1972).
Larralde, R., Robertson, M. P. & Miller, S. L. Rates of decomposition of ribose and other sugars: implications for chemical evoution. Proc. Natl Acad. Sci. USA 92, 8158–8160 (1995).
Mizuno, T. & Weiss, A. H. Synthesis and utilization of formose sugars. Adv. Carbohyd. Chem. Biochem. 29, 173–227 (1974).
Pitsch, S., Eschenmoser, A., Gedulin, B., Hui, S. & Arrhenius, G. Mineral induced formation of sugar phosphates. Origin Life Evol. Biosphere 25, 297–334 (1995).
Oro, J. Mechanism of synthesis of adenine from hdyrogen cyanide under possible primitive earth conditions. Nature 191, 1193–1194 (1961).
Sanchez, R. A., Ferris, J. P. & Orgel, L. E. Studies in prebiotic synthesis. II. Synthesis of purine precursors and amino acids from aqueous hydrogen cyanide. J. Mol. Biol. 30, 223–253 (1967).
Ferris, J. P., Sanchez, R. A. & Orgel, L. E. Studies in prebiotic synthesis. 3. Synthesis of pyrimidines from cyanoacetylene and cyanate. J. Mol. Biol. 33, 693–704 (1968).
Stoks, P. G. & Schwartz, A. W. Uracil in carbonaceous meteorites. Nature 282, 709–710 (1979).
Robertson, M. P. & Miller, S. L. An efficient prebiotic synthesis of cytosine and uracil. Nature 375, 772–774 (1995).
Bhatia, M. B., Vinitsky, A. & Grubmeyer, C. Kinetic mechanism of orotate phosphoribosyltransferase from Salmonella typhimurium. Biochemistry 29, 10480–10487 (1990).
Tao, W., Grubmeyer, C. & Blanchard, J. S. Transition state structure of Salmonella typhimurium orotate phosphoribosyltransferase. Biochemistry 35, 14–21 (1996).
Igloi, G. L. Interaction of tRNAs and of phosphorothioate-substituted nucleic acids with an organomercurial. Probing the chemical environment of thiolated residues by affinity electrophoresis. Biochemistry 27, 3842–3849 (1988).
Wierzchowski, K. L., Litonska, E. & Shugar, D. Infrared and ultraviolet studies on the tautomeric equilibria in aqueous medium between monoanionic species of uracil, thymine, 5-fluorouracil, and other 2,4-diketopyrimidines. J. Am. Chem. Soc. 87, 4621–4629 (1965).
Psoda, A., Kazimierczuk, Z. & Shugar, D. Structure and tautomerism of the neutral and monoanionic forms of 4-thiouracil derivatives. J. Am. Chem. Soc. 96, 6832–6839 (1974).
Bartel, D. P. & Szostak, J. W. Isolation of new ribozymes from a large pool of random sequences. Science 261, 1411–1418 (1993).
Gray, M. W. The presence of 2′-O-methylpseudouridine in the 18S + 26S ribosomal ribonucleates of wheat embryo. Biochemistry 13, 5453–5463 (1974).
Jaffe, E. K. & Cohn, M. Diastereomers of the nucleoside phosphorothioates as probes of the structure of the metal nucleotide substrates and of the nucleotide binding site of yeast hexokinase. J. Biol. Chem. 254, 10839–10845 (1979).
Benner, S. A., Ellington, A. D. & Tauer, A. Modern metabolism as a palimpsest of the RNA world. Proc. Natl Acad. Sci. USA 86, 7054–7058 (1989).
Michel, F., Hanna, M., Green, R., Bartel, D. P. & Szostak, J. W. The guanosine binding site of the Tetrahymena ribozyme. Nature 342, 391–395 (1989).
Been, M. D. & Perrotta, A. T. Group I intron self-splicing with adenosine: evidence for a single nucleoside-binding site. Science 252, 434–437 (1991).
Gold, L., Polisky, B., Uhlenbeck, O. & Yarus, M. Diversity of oligonucleotide functions. Annu. Rev. Biochem. 64, 763–797 (1995).
Ekland, E. H. & Bartel, D. P. RNA-catalysed RNA polymerization using nucleoside triphosphates. Nature 382, 373–376 (1996).
Ekland, E. H. & Bartel, D. P. The secondary structure and sequence optimization of an RNA ligase ribozyme. Nucleic Acids Res. 23, 3231–3238 (1995).
England, T. E., Gumport, R. I. & Uhlenbeck, O. C. Dinucleoside pyrophosphates are substrates for T4-induced RNA ligase. Proc. Natl Acad. Sci. USA 74, 4839–4842 (1977).
Lohrmann, R. & Orgel, L. E. Preferential formation of (2′-5′)-linked internucleotide bonds in non-enzymatic reactions. Tetrahedron 34, 853–855 (1978).
Santoro, S. W. & Joyce, G. F. Ageneral purpose RNA-cleaving DNA enzyme. Proc. Natl Acad. Sci. USA 94, 4262–4266 (1997).
Mizuno, Y., Ikehara, M. & Watanabe, K. A. Potential antimetabolites. I. Selective thiation of uracil and 1,2,4-triazine-3,5(2H,4H)-dione (6-azauracil). Chem. Pharmac. Bull. 10, 647–652 (1962).
Wecker, M., Smith, D. & Gold, L. In vitro selection of a novel catalytic RNA: characterization of a sulfur alkylation reaction and interaction with a small peptide. RNA 2, 982–994 (1996).
Acknowledgements
We thank J. Stubbe, P. Zamore and members of the lab for helpful comments on the manuscript, and G. Joyce for providing the sequence of the RNA-cleaving DNA enzyme28 before publication. This work was supported by an MRC (Canada) postdoctoral fellowship to P.J.U. and a grant from the Searle Scholars Program/The Chicago Community Trust to D.P.B.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Unrau, P., Bartel, D. RNA-catalysed nucleotide synthesis. Nature 395, 260–263 (1998). https://doi.org/10.1038/26193
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/26193
This article is cited by
-
In vitro selection of aptamers and their applications
Nature Reviews Methods Primers (2023)
-
An RNA aptamer that shifts the reduction potential of metabolic cofactors
Nature Chemical Biology (2022)
-
Prebiotic chemical refugia: multifaceted scenario for the formation of biomolecules in primitive Earth
Theory in Biosciences (2022)
-
Cofactors are Remnants of Life’s Origin and Early Evolution
Journal of Molecular Evolution (2021)
-
From molecular to cellular form: modeling the first major transition during the arising of life
BMC Evolutionary Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.