Abstract
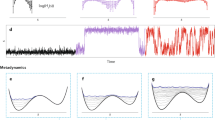
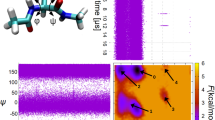
Molecular dynamics—the science of simulating the motions of a system of particles—applied to biological macromolecules gives the fluctuations in the relative positions of the atoms in a protein or in DNA as a function of time. Knowledge of these motions provides insights into biological phenomena such as the role of flexibility in ligand binding and the rapid solvation of the electron transfer state in photosynthesis. Molecular dynamics is also being used to determine protein structures from NMR, to refine protein X-ray crystal structures faster from poorer starting models, and to calculate the free energy changes resulting from mutations in proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Brooks, C. L., III, Karplus, M. & Pettitt, B. M. Proteins: A Theoretical Perspective of Dynamics, Structure and Thermodynamics (Wiley, New York, 1988).
McCammon, J. A. & Harvey, S. Dynamics of Proteins and Nucleic Acids (Cambridge University Press, Cambridge, 1987).
Barnes, J. E. Nature 338, 123–126 (1989).
Brooks, B. R. et al. J. comput. Chem. 4, 187–217 (1983).
Stillinger, F. H. & Rahman, A. J. chem. Phys. 60, 1545–1557 (1974).
McCammon, J. A., Gelin, B. R. & Karplus, M. Nature 267, 585–590 (1977).
Aqvist, J., van Gunsteren, W. F., Leijonmarck, M. & Tapia, O. J. molec. Biol. 183, 461 (1985).
Dobson, C. M. & Karplus, M. Meth. Enzym. 131, 362–389 (1986).
Gelin, B. R. & Karplus, M. Proc. natn. Acad. Sci. U.S.A. 74, 801–805 (1977).
Ringe, D. et al. Trans. Am. cryst. Ass. 20, 109–122 (1984).
Doster, W., Cusack, S. & Petry, W. Nature 337, 754–755 (1989).
Smith, J., Kuczera, K. & Karplus, M. Proc. natn. Acad. Sci. U.S.A. 87, 1601–1605 (1990).
Elber, R. & Karplus, M. Science 235, 318–321 (1987).
Chothia, C. & Lesk, A. M. Trends biochem. Sci. 10, 116 (1985).
Lesk, A. M. & Chothia, C. J. molec. Biol. 136, 225 (1980).
Case, D. A. & Karplus, M. J. molec. Biol. 132, 343–368 (1979).
Elber, R. & Karplus, M. J. Am. chem. Soc. (in the press).
Teutlein, H. et al. in The Photosynthetic Bacterial Reaction Center (ed. Vermeglio, A.) 101–118 (Plenum, London, 1988).
Fleming, G. R., Martin, J.-L. & Breton, J. Nature 333, 190–192 (1988).
Brünger, A. T., Clore, G. M., Gronenborn, A. M. & Karplus, M. Proc. natn. Acad. Sci. U.S.A. 83, 3801–3805 (1986).
Wüthrich, K. Accts chem. Res. 22, 36–44 (1989).
Brünger, A. T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1987).
Brünger, A. T. & Karplus, M. Accts chem. Res. (in the press).
Beveridge, D. L. & Di Capua, F. M. A. Rev. Biophys. biophys. Chem. 181, 431–492 (1989).
Gao, J., Kuczera, K., Tidor, B. & Karplus M. Science 244, 1069–1072 (1989).
Harvey, S. Proteins 5, 78–92 (1989).
Deisenhofer, J. & Michel, H. Science 245, 1463–1473 (1989).
Post, C. B. et al. J. molec. Biol. 190, 455–479 (1986).
Hoch, J. C., Dobson, C. M. & Karplus, M. Biochemistry 24, 3831–3841 (1985).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Karplus, M., Petsko, G. Molecular dynamics simulations in biology. Nature 347, 631–639 (1990). https://doi.org/10.1038/347631a0
Issue Date:
DOI: https://doi.org/10.1038/347631a0
This article is cited by
-
Identification of Potential Hits against Fungal Lysine Deacetylase Rpd3 via Molecular Docking, Molecular Dynamics Simulation, DFT, In-Silico ADMET and Drug-Likeness Assessment
Chemistry Africa (2024)
-
Drawing a materials map with an autoencoder for lithium ionic conductors
Scientific Reports (2023)
-
High-throughput single-molecule quantification of individual base stacking energies in nucleic acids
Nature Communications (2023)
-
A State-of-the-Art Review on Machine Learning-Based Multiscale Modeling, Simulation, Homogenization and Design of Materials
Archives of Computational Methods in Engineering (2023)
-
Design of short peptides and peptide amphiphiles as collagen mimics and an investigation of their interactions with collagen using molecular dynamics simulations and docking studies
Journal of Molecular Modeling (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.