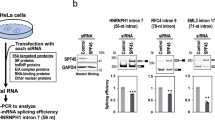
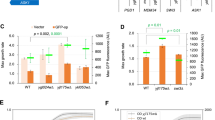
Abstract
FIVE small nuclear RNAs (snRNAs) are required for nuclear pre-messenger RNA splicing: Ul, U2, U4, U5 and U61,2. The yeast Ul and U2 snRNAs base-pair to the 5′ splice site and branch-point sequences of introns respectively1. The role of the U5 and U4/U6 small nuclear ribonucleoprotein particles (snRNPs) in splicing is not clear, though a catalytic role for the U6 snRNA has been proposed3. Less is known about yeast splicing factors, but the availability of genetic techniques in Saccharomyces cerevisiae has led to the identification of mutants deficient in nuclear pre-mRNA splicing (prp2-prp27)4,5. Several PRP genes have now been cloned and their protein products characterized. The PRP8 protein is a component of the US snRNP and associates with the U4/U6 snRNAs/snRNP to form a multi-snRNP particle believed to be important for spliceosome assembly6. We have isolated extragenic suppressors of the prp8–l mutation of S. cerevisiae and present here the preliminary characterization of one of these suppressors, spp81. The predicted amino-acid sequence of the SPP81protein shows extensive similarity to a recently identified family of proteins thought to possess ATP–dependent RNA helicase activity. The possible role of this putative helicase in nuclear pre-mRNA splicing is discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Krainer, A. R. & Maniatis, T. in Transcription and Splicing 131–206 (IRL, Oxford, 1988).
Guthrie, C. & Patterson, B. A. Rev. Genet. 22, 387–419 (1988).
Brow, D. A. & Guthrie, C. Nature 334, 213–218 (1988).
Hartwell, L. H. J. Bacteriol. 93, 1662–1670 (1967).
Vijayraghaven, U., Company, M. & Abelson, J. Genes Dev. 3, 1206–1216 (1989).
Lossky, M., Anderson, G. J., Jackson, S. P. & Beggs, J. D. Cell 51, 1019–1026 (1987).
Jackson, S. P., Lossky, M. & Beggs, J. D. Molec. cell. Biol. 8, 1067–1075 (1988).
Pearson, N. J., Thorburn, P. C. & Haber, J. E. Molec. cell. Biol. 2, 571–577 (1982).
Struhl, K. Nucleic. Acids Res. 13, 8587–8601 (1985).
Struhl, K., Strinchcombe, D. T., Scherer, S. & Davis, R. R. Proc. natn. Acad. Sci. U.S.A. 76, 1035–1039 (1979).
Struhl, K. & Davis, R. R. J. molec. Biol. 136, 309–322 (1980).
Ford, M. J., Anton, I. A. & Lane, D. P. Nature 332, 736–738 (1988).
Linder, P. et al. Nature 337, 121–122 (1989).
Iggo, R. D. & Lane, D. P. EMBO J. 8, 1827–1831 (1989).
Hirling, H., Scheffner, M., Restle, T. & Stahl, M. Nature 339, 562–564 (1989).
Leroy, P., Alzari, P., Sassoon, D., Wolgemuth, D. & Fellous, M. Cell 57, 549–559 (1989).
Seraphin, B., Simon, M., Boulet, Z. & Faye, G. Nature 334, 84–87 (1989).
Siliciano, P. G., Brow, D. A., Roiha, H. & Guthrie, C. Cell 50, 585–592 (1988).
Lamond, A. I., Konarska, M. M., Grabowski, P. J. & Sharp, P. A. Proc. natn. Acad. Sci. U.S.A. 85, 411–415 (1988).
Dalbadie-McFarland, G. & Abelson, J. Proc. natnl. Acad. Sci. U.S.A. 87, 4236–4240 (1990).
Gallwitz, D. & Sures, I. Proc. natn. Acad. Sci. U.S.A. 77, 2546–2550 (1980).
Feinberg, A. P. & Vogelstein, B. Anal. Biochem. 132, 6–13 (1983).
Hopper, A. K., Banks, F. & Evangelidis, V. Cell 14, 211–219 (1978).
Beltzer, J. P., Chang, L.-F., Hinkkenen, A. E. & Kohlhaw, G. B. J. biol. Chem. 261, 5160–5167 (1986).
Henikoff, S. Gene 28, 351–359 (1984).
Higgins, D. G. & Sharp, P. M. CABIOS 5, 151–153 (1989).
Sherman, F., Fink, G. R. & Hicks, J. B. Methods in Yeast Genetics (Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, 1982).
Chen, W. & Struhl, K. EMBO J. 4, 3273–3280 (1985).
Ito, H., Fukuda, Y., Murata, K. & Kimura, A. J. Bact. 153, 163–168 (1983).
Jamieson, D. J. & Beggs, J. D. Molec. Microbiol. (in the press).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Jamieson, D., Rahe, B., Pringle, J. et al. A suppressor of a yeast splicing mutation (prp8-l) encodes a putative ATP-dependent RNA helicase. Nature 349, 715–717 (1991). https://doi.org/10.1038/349715a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/349715a0
This article is cited by
-
The zinc finger protein Zn72D and DEAD box helicase Belle interact and control maleless mRNA and protein levels
BMC Molecular Biology (2009)
-
Exploring functional relationships between components of the gene expression machinery
Nature Structural & Molecular Biology (2005)
-
Identification of four genes involved in suppression of the pre-mRNA splicing defect in thesng1-1/rhp6 - mutant of fission yeast
Journal of Genetics (2000)
-
An extragenic suppressor ofprp24-1 defines genetic interaction betweenPRP24 andPRP21 gene products ofSaccharomyces cerevisiae
Molecular and General Genetics MGG (1996)
-
The S. cerevisiae nuclear gene SUV3 encoding a putative RNA helicase is necessary for the stability of mitochondrial transcripts containing multiple introns
Current Genetics (1995)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.