Abstract
Sex chromosomes have evolved independently in several different groups of organisms, but they share common features, including genetic degeneration of the Y chromosome1,2. Suppression of recombination between ancestral proto-X and proto-Y chromosomes is thought to have led to their gradual divergence, and to degeneration of the Y chromosome2, but the evolutionary forces responsible are unknown. In non-recombining Y chromosomes, deleterious mutations may be carried to fixation by linked advantageous mutations (“selective sweeps”)3. Occurrence of deleterious mutations may drive “Muller's ratchet” (stochastic loss of chromosomes with the fewest mutations)2,4. Selective elimination of deleterious mutations, causing “background selection”5,6 may accelerate stochastic fixation of mildly detrimental mutations2. All these processes lower effective population sizes, and therefore reduce variability of genes in evolving Y chromosomes. We have studied DNA diversity and divergence in a recently described X- and Y-linked gene pair7 (SLX-1 and SLY-1) of the plant Silene latifolia to obtain evidence about the early stages of Y degeneration. Here we show that DNA polymorphism in SLY-1 is 20-fold lower than in SLX-1, but the pattern of polymorphism does not suggest a selective sweep.
Similar content being viewed by others
Main
White campion Silene latifolia (Caryophyllaceae) is a short-lived perennial plant species that is dioecious (individuals are male or female) and has a mammal-like chromosomal sex-determination system8,9 (XX female and XY male). The Y chromosome is important in the suppression of female and promotion of male flower characters9. S. latifolia X and Y chromosomes, like those of mammals, have a small recombining pseudo-autosomal region9, estimated to be about 10% of the Y chromosome. The ancestor of S. latifolia seems to have diverged from the closest non-dioecious species about 10–20 million years ago10, and the morphologically distinct sex chromosomes probably evolved more recently. The relatively recent origin of the S. latifolia sex chromosomes provides an opportunity to study the evolutionary forces that operate in the early stages of sex chromosome evolution. Animal sex chromosome systems (such as those in humans11, hominids12, rodents13,14 and Drosophila15) are ancient, and their Y chromosomes are almost completely degenerated. The only data about the early stages of genetic degeneration have been obtained from estimates of DNA polymorphism on a Drosophila americana neo-Y, a recent translocation onto the Y chromosome that occurred about one million years ago. These data show only a small decrease in DNA polymorphism relative to the neo-X (ref. 16), which indicates that genetic diversity may be lost slowly from the Y chromosome. Despite its relatively recent origin, the S. latifolia Y chromosome shows some signs of degeneration8,9, and plants that have a Y but no X chromosome are usually inviable17,18. The S. latifolia Y chromosome is bigger than the X, probably owing to the accumulation of repetitive DNA; several attempts to isolate active Y-linked genes from S. latifolia yielded only repetitive sequences19. The first X-linked gene identified in this species seems to have a degenerated Y-linked homologue20.
Only one active Y-linked gene (SLY-1) has so far been characterized in S. latifolia7. Its non-Y-linked homologue was believed to be located on the X chromosome and was named SLX-1, although its X-linkage had not been proved. We tested this by observing the segregation of a molecular genetic marker (an HpaI restriction site) in the progeny of parents with the restriction site present (homozygous mother) or absent (father). All ten male progeny inherited the maternal variant, and five females were all heterozygous for the maternal and paternal alleles. We therefore conclude that SLX-1 is X-linked.
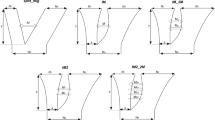
The coding regions of SLX-1 and SLY-1 have very similar sequences7, which may indicate that they are located in the recombining pseudo-autosomal region of the sex chromosomes. Although segregation tests suggest complete linkage of SLY-1 to sex7, these data are from a family of moderate size, and the probability of detecting rare recombination is low. However, rare recombination can be detected by examining sequence variants. If recombination occurs, some variant sites should be polymorphic in both genes in natural populations, and there should be no fixed differences between the genes. We compared sequences of the same 2-kilobase (kb) region of both SLX-1 and SLY-1 (Fig. 1) from 12 males: 3 were from independent laboratory strains, and the remainder were from US (n = 2), Scottish (n = 5) and Portuguese (n = 2) natural populations.
The entire SLlY-1 gene (C. Delichére, personal communication) is shown, to specify the intron numbers. For the region studied, we determined the intron positions (thin blocks) in both SLX-1 and SLY-1; they are in the same positions in both genes. Exon numbers are shown below the SLY-1 cDNA. Non-coding 3′ regions of the genes are shown in grey. The positions and directions of primers are shown by arrows. Primers ‘+11’, ‘+9’, ‘+6’ and ‘-10’ are homologous to both SLX-1 and SLY-1. Primers ‘-7’ and ‘-8’ are gene-specific.
The SLY-1 sequences showed five segregating nucleotide substitutions (four in introns and a Pro/Leu amino-acid replacement at position 349 in exon 13 of the SLY-1 protein) and one seven-nucleotide insertion–deletion (indel) polymorphism in intron 13 (Fig. 1). To quantify the diversity values21 we estimated two commonly used measures of variability, θ = 0.00082 ± 0.00046, and π = 0.0010 ± 0.0002, excluding the indel variant. When the indel was treated as a single variant, θ = 0.00098 ± 0.00053 and π = 0.00122 ± 0.00023. In contrast, the same 2-kb region of the SLX-1 alleles revealed 95 segregating nucleotide substitutions (5 replacement substitutions and 90 silent variants) and 33 indel polymorphisms (all in introns). Excluding indel regions, θ = 0.016 ± 0.0016 and π = 0.016 ± 0.0013.
Between SLX-1 and SLY-1, there are 54 fixed nucleotide differences and no shared polymorphic sites. The SLX-1 sequences, but not those of SLY-1, show evidence of recombination (for the 12 SLX-1 allele sequences the “four gamete test”22 detects a minimum of 16 recombination events, and the estimate of recombination per nucleotide23 is 0.015). The contrast between SLX-1 and SLY-1 shows that SLY-1 is located in the differential segment of the Y chromosome.
The net silent site divergence, Ks (ref. 21), between the SLX-1 and SLY-1 sequences is 3%. Assuming a synonymous site molecular clock with a rate of about 0.6% per million years, the best current estimate for plant nuclear genes24, this corresponds to a total divergence of about 5 million years between the X and Y genes, and thus about 2.5 million years since they stopped recombining. This is less than the putative age of the sex chromosomes10 and much less than the divergence between the MROS3-X and MROS3-Y genes20. This indicates that cessation of recombination may have occurred at different times for different loci, possibly owing to migration of the boundary between the pseudo-autosomal region and non-recombining sections of the Y chromosome.
The apparent recent halting of recombination provides an opportunity to analyse variation in SLY-1 and to infer the evolutionary forces involved in the degeneration of the Y chromosome. If no selective factors (genetic hitch-hiking, background selection and so on) were operating, the effective population size of a Y-linked gene would be one-quarter of that for an autosomal gene, and one-third of that for an X-linked gene. Thus, in the neutral case, SLY-1 should have roughly one-third of the DNA variation observed in SLX-1. However, SLY-1 DNA polymorphism is reduced by a factor of 20 compared with that of SLX-1 (Fig. 2). Neutral coalescent simulations25, based on 32 segregating sites (one-third of the number in SLX-1), show that the diversity in SLY-1 is significantly (P < 0.0001) reduced.
What process decreased SLY-1 DNA diversity so greatly in this short time? Sexual selection can decrease the Y effective population size compared with that of X chromosomal loci. This cannot be excluded, but is unlikely to have had such an extreme and rapid effect in an insect-pollinated plant2. The difference in SLX-1 and SLY-1 diversity could be due to a bottleneck in effective population size for Y chromosomes. However, this is not consistent with the data because the extent of SLX-1/SLY-1 divergence implies that there was sufficient time for SLY-1 diversity to recover after recombination ceased. A recent selective sweep at a locus on the S. latifolia Y chromosome, eliminating diversity throughout the species, also appears unlikely, because existing SLY-1 polymorphisms should then be variants accumulated since the selective sweep and should thus mostly be singletons. We found, however, that none of the six SLY-1 polymorphic sites (including the indel variant) are singletons, and no significant deviation from neutrality was detected in these data by Tajima's D (ref. 26) and Fu and Li's D*and F* statistics27, which were positive, not negative as would be expected after a recent selective sweep (data not shown). Finally, we generated samples of 12 sequences with 32 segregating sites (one-third of the number in the SLX-1 sample) by coalescent simulations, assuming selective sweeps at different times in the sample's history. The simulations assumed no recombination and used several different strengths of selection. Given the estimated reduction in the number of segregating sites in the Y-linked gene, the number of generations since the selective sweep probably does not exceed the effective population size21 of the species; longer times yielded numbers of segregating sites greater than were seen in less than 5% of simulations. Assuming this length of time, or longer, the simulations almost invariably yielded frequencies of single substitutions that were much higher than those observed (Table 1). Thus a simple, classical selective sweep cannot explain the low diversity of the SLY-1 gene. This differs from the interpretation of low sequence variability in a Y-linked dynein gene in Drosophila15, which may be attributable to selective sweeps, rather than deleterious mutations, because D. melanogaster Y chromosomes carry few genes and thus a high deleterious mutation rate seems unlikely. However, our results do not rule out background selection, which does not greatly affect the shape of the gene tree5. An explanation involving Muller's ratchet may also be compatible with our data, but no detailed study has yet been done of the ratchet's effects on diversity at neutral sites in a non-recombining chromosome, and more theoretical work is needed.
Methods
We used the sequences of SLX-1 and SLY-1 to design the following primers for the polymerase chain reaction (PCR) and sequencing (Fig. 1): ‘-7’, 5′-ACTTGCAACG ACTTCACTTTGAG-3′; ‘-8’, 5′-ATCGAATTCCAGTGGAAGTCCA-3′; ‘+11’, 5′-AAGCTCACAATGCTGATCTTC-3′; ‘+9’, 5′-GCTGAAGATGGCTTGCTAAAC-3′; ‘+6’, 5′-TGGACTTCCACTGGAATTCGAT-3′; ‘-10’, 5′-TCCAGCAGAGCTTGAACAGTC-3′.
A standard cetyltrimethylammonium bromide plant miniprep method with several modifications28 was used to isolate the total genomic DNA from 12 S. latifolia males and from 17 plants (2 parents, 10 sons and 5 daughters) used in the SLX-1 segregation analysis. The 2-kb 3′ region in 12 S. latifolia males was amplified using the following primer pairs: ‘+11’ and ‘-7’ for SLX-1; ‘+11’ and ‘-8’ for SLY-1. Because we studied sequences from the X and Y chromosomes of males, direct sequencing was possible for all individuals. Amplification products were passed through 1% agarose gels, column-purified (Qiagen gel extraction kit) and sequenced with the primers listed above, using an ABI Prism 377 automatic sequencer (Perkin Elmer).
The primer pair ‘+6’ and ‘-7’ was used to amplify the short (0.6 kb) 3′ SLX-1 region used in the SLX-1 segregation analysis. PCR products of both parents were directly sequenced and these sequences were used to choose a restriction enzyme (HpaI) to cut the PCR product of only one of the parents. The restriction site was used as a molecular genetic marker in the SLX-1 segregation analysis.
For the coalescent simulations, we used the program ProSeq v.2.4 (D. Filatov, unpublished, available at http://helios.bto.ed.ac.uk/evolgen/filatov/proseq.html). All simulations assumed zero recombination. Simulations to estimate the significance of the lack of diversity in the SLY-1 gene were conditioned on the observed numbers of segregating sites. Simulations with a selective sweep were conducted according to ref. 29. The neighbour-joining tree was constructed using MEGA software30, based on pairwise divergence values, with Jukes-Cantor correction.
References
Bull, J. J. Evolution of Sex Determining Mechanisms. (Benjamin/Cummings, Menlo Park, 1993).
Charlesworth, B. The evolution of chromosomal sex determination and dosage compensation. Curr. Biol. 6, 149–162 (1996).
Rice, W. R. Genetic hitchhiking and the evolution of reduced genetic activity of the Y sex chromosome. Genetics 116, 161–167 (1987).
Manning, J. T. & Thompson, D. J. Muller's ratchet and the accumulation of favourable mutations. Acta Biotheoretica 33, 219–225 (1984).
Charlesworth, D., Charlesworth, B. & Morgan, M. T. The pattern of neutral molecular variation under the background selection model. Genetics 141, 1619–1632 (1995).
Orr, H. A. & Kim, Y. An adaptive hypothesis for the evolution of the Y chromosome. Genetics 150, 1693–1698 (1998).
Delichère, C. et al. SlY1, the first active gene cloned from a plant Y chromosome encodes a WD-repeat protein. EMBO J. 18, 4169–4179 (1999).
Grant, S. et al. Genetics of sex determination in flowering plants. Dev. Genet. 15, 214–230 (1994).
Westergaard, M. The mechanism of sex determination in dioecious flowering plants. Adv. Genet. 9, 217–281 (1959).
Desfeux, C. et al. Evolution of reproductive systems in genus Silene. Proc. R. Soc. Lond. B. 263, 409–414 (1996).
Anagnostopoulos, T., Green, P. M., Rowley, G., Lewis, C. M. & Giannelli, F. DNA variation in a 5-Mb region of the X-chromosome and estimates of sex-specific/type-specific mutation rates. Am. J. Hum. Genet. 64, 508–517 (1999).
Burrows, W. & Ryder, O. A. Y-chromosome variation in great apes. Nature 385, 125–126 (1997).
Lundrigan, B. L. & Tucker, P. K. Tracing paternal ancestry in mice, using the Y-linked sex-determining locus Sry. Mol. Biol. Evol. 11, 483–492 (1994).
Nachman, M. W. & Aquadro, C. F. Polymorphism and divergence at the 5′ flanking region of the sex determining locus, Sry, in mice. Mol. Biol. Evol. 11, 539–547 (1994).
Zurovcova, M. & Eanes, W. F. Lack of nucleotide polymorphism in the Y-linked sperm flagellar dynein gene Dhc-Yh3 of Drosophila melanogaster and D. simulans. Genetics 153, 1709–1715 (1999).
McAllister, B. & Charlesworth, B. Reduced sequence variability on the neo-Y chromosome of Drosophila americana americana. Genetics 153, 221–233 (1999).
Ye, D. et al. Sex determination in the dioecious Melandrium. I. First lessons from androgenic haploids. Sex. Plant Reprod. 3, 179–186 (1990).
Vagera, J., Paulikova, D. & Dolezel, J. The development of male and female regenerants by in vitro Androgenesis in dioecious plant Melandrium album. Ann. Botany 73, 455–459 (1994).
Bùzek, J. et al. Isolation and characterization of X chromosome-derived DNA sequences from a dioecious plant Melandrium album. Chromosome Res. 5, 57–65 (1997).
Guttman, D. S. & Charlesworth, D. An X-linked gene with a degenerate Y-linked homologue in a dioecious plant. Nature 393, 263–265 (1998).
Nei, M. Molecular Evolutionary Genetics. (Columbia Univ. Press, New York, 1987).
Hudson, R. R. & Kaplan, N. L. Statistical properties of the number of recombination events in the history of a sample of DNA sequences. Genetics 111, 147–164 (1985).
Hudson, R. R. Estimating the recombination parameter of a finite population model without selection. Genet. Res. 50, 245–250 (1987).
Gaut, B. S. Molecular clocks and nucleotide substitution rates in higher plants. Evol. Biol. 30, 93–120 (1998).
Hudson, R. R. Gene genealogies and the coalescent process. Oxf. Surv. Evol. Biol. 7, 1–44 (1990).
Fu, Y. -X. & Li, W.-H. Statistical tests of neutrality of mutations. Genetics 133, 693–709 (1993).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Filatov., D. A. & Charlesworth, D. DNA polymorphism, haplotype structure and balancing selection in the Leavenworthia PgiC locus. Genetics 153, 1423–1434 (1999).
Braverman, J. M., Hudson, R. R., Kaplan, N. L., Langley, C. H. & Stephan, W. The hitchhiking effect on the site frequency spectrum of DNA polymorphism. Genetics 140, 783–796 (1995).
Kumar, S., Tamura, K. & Nei, M. MEGA: Molecular Evolutionary Genetics Analysis, version 1.0. (Pennsylvania State Univ., Univ. Park, 1993).
Acknowledgements
We thank M. Turelli and B. Charlesworth for discussions and advice. D.C. was supported by the Natural Environment Research Council of Great Britain, D.A.F. by a grant from the Leverhulme Trust, and F.M. and I.N. by research contracts from the Centre National de la Recherche Scientifique, the Institut National de la Recherche Agrononique and the Ecole Normale Supérieure of France.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Filatov, D., Monéger, F., Negrutiu, I. et al. Low variability in a Y-linked plant gene and its implications for Y-chromosome evolution. Nature 404, 388–390 (2000). https://doi.org/10.1038/35006057
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35006057
This article is cited by
-
Evolution of a plant sex chromosome driven by expanding pericentromeric recombination suppression
Scientific Reports (2024)
-
Gene flow signature in the S-allele region of cultivated buckwheat
BMC Plant Biology (2019)
-
The slowdown of Y chromosome expansion in dioecious Silene latifolia due to DNA loss and male-specific silencing of retrotransposons
BMC Genomics (2018)
-
Extremely low nucleotide diversity in the X-linked region of papaya caused by a strong selective sweep
Genome Biology (2016)
-
Neo-sex chromosomes of Ronderosia bergi: insight into the evolution of sex chromosomes in grasshoppers
Chromosoma (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.