Abstract
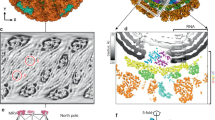
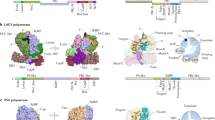
The reovirus core is an assembly with a relative molecular mass of 52 million that synthesizes, modifies and exports viral messenger RNA. Analysis of its structure by X-ray crystallography shows that there are alternative, specific and completely non-equivalent contacts made by several surfaces of two of its proteins; that the RNA capping and export apparatus is a hollow cylinder, which probably sequesters its substrate to ensure completion of the capping reactions; that the genomic double-stranded RNA is coiled into concentric layers within the particle; and that there is a protein shell that appears to be common to all groups of double-stranded RNA viruses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fields,B. N. in Fields Virology (eds Fields, B. N., Knipe, D. M. & Howley, P. M.) 1553–1555 (Lippincott-Raven, Philadelphia, 1996).
Nibert,M. L., Schiff,L. A. & Fields, B. N. in Fields Virology (eds Fields, B. N., Knipe, D. M. & Howley, P. M.) 1557–1596 (Lippincott-Raven, Philadelphia, 1996).
Grimes,J. M. et al. The atomic structure of the bluetongue virus core. Nature 395, 470–478 ( 1998).
Furuichi,Y., Muthukrishnan,S., Tomasz, J. & Shatkin,A. Mechanism and formation of reovirus mRNA 5′-terminal blocked and methylated sequence m7GpppGmpC. J. Biol. Chem. 251, 5043–5053 ( 1976).
Earnshaw,W. C. & Harrison,S. C. DNA arrangement in isometric phage heads. Nature 268, 598– 602 (1977).
Dryden,K. A. et al. Internal structures containing transcriptase-related proteins in top component particles of mammalian orthoreovirus. Virology 245, 33–46 ( 1998).
Gouet,P. et al. The highly ordered double-stranded RNA genome of bluetongue virus revealed by crystallography. Cell 97, 481 –490 (1999).
Hill,C. L. et al. The structure of cypovirus and the functional organization of dsRNA viruses. Nature Struct. Biol. 6, 565 –568 (1999).
Dryden,K. A. et al. Early steps in reovirus infection are associated with dramatic changes in supramolecular structure and protein conformation: analysis of virions and subviral particles by cryoelectron microscopy and image reconstruction. J. Cell Biol. 122, 1023– 1041 (1993).
Kohlstaedt,L. A., Wang,J., Friedman,J. M., Rice,P. A. & Steitz,T. A. Crystal structure of 3.5?Å resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 256, 1783–1790 ( 1992).
Stehle,T., Yan,Y., Benjamin,T. L. & Harrison,S. C. Structure of murine polyomavirus complexed with an oligosaccharide receptor fragment. Nature 369, 160–163 ( 1994).
Liddington,R. C. et al. Structure of simian virus 40 at 3.8?Å resolution. Nature 354, 278–284 (1991).
Labbe,M., Charpilienne,A., Crawford, S. E., Estes,M. K. & Cohen,J. Expression of rotavirus VP2 produces empty corelike particles. J. Virol. 65, 2946–2952 (1991).
Moss,S. R. & Nuttall,P. A. Subcore- and core-like particles of Broadhaven virus (BRDV), a tick-bourne orbivirus, synthesized from baculovirus expressed VP2 and VP7, the major core proteins of BRVD. Virus Res. 32, 401–407 ( 1994).
Xu,P., Miller,S. & Joklik,W. K. Generation of reovirus core-like particles in cells infected with hybrid vaccinia viruses that express genome segments L1, L2, L3, and S2. Virology 197, 726– 731 (1993).
Mao,Z. & Joklik,W. K. Isolation and enzymatic characterization of protein λ2, the reovirus guanylyltransferase. Virology 185, 377–386 ( 1991).
Luongo,C. L., Reinisch,K. M., Harrison, S. C. & Nibert,M. L. Identification of the guanylyltransferase region and active site in reovirus mRNA capping protein l2. J. Biol. Chem. 275, 2804–2810 (2000).
Håkansson,K., Doherty,A. J., Shuman,S. & Wigley,D. B. X-ray crystallography reveals a large conformational change during guanyl transfer by mRNA capping enzymes. Cell 89, 545–553 (1997).
Schluckebier,G., O'Gara,M., Saenger,W. & Cheng,X. Universal catalytic domain structure of AdoMet-dependent methyltransferases. J. Mol. Biol. 247, 16–20 ( 1995).
Hodel,A. E., Gershon,P. D., Shi,X. & Quiocho,F. A. The 1.85?Å structure of the vaccinia protein VP39: a bifunctional enzyme that participates in the modification of both mRNA ends. Cell 85, 247–256 (1996).
Hu,G., Gershon,P. D., Hodel,A. E. & Quiocho,F. A. mRNA cap recognition: dominant role of enhanced stacking interactions between methylated bases and protein aromatic side chains. Proc. Natl Acad. Sci. USA 96, 7149–7154 ( 1999).
Wickner,S., Maurizi,M. R. & Gottesman, S. Posttranslational quality control: Folding, refolding, and degrading proteins. Science 286, 1888 –1893 (1999).
Earnshaw,W. C., King,J., Harrison,S. C. & Eiserling,F. A. The structural organization of DNA packaged within the heads of T4 wild-type, isometric and giant bacteriophages. Cell 14, 559– 568 (1978).
Harvey,J. D., Bellamy,A. R., Earnshaw,W. C. & Schutt,C. Biophysical studies of reovirus type 3: iv low-angle x-ray diffraction studies. Virology 112, 240–249 (1981).
Harrison,S. C. Packaging of DNA into bacteriophage heads: a model. J. Mol. Biol. 171, 577–580 ( 1983).
Cerritelli,M. E. et al. Encapsidated conformations of bacteriophage T7 DNA. Cell 91, 271–280 ( 1997).
Booy,F. P. et al. Liquid crystalline, phage-like packing of encapsidated DNA in herpes simplex virus. Cell 64, 1007– 1015 (1991).
Butcher,S. J., Dokland,T., Ojala,P. M., Bamford,D. H. & Fuller, S. D. Intermediates in the assembly pathway of the double-stranded RNA virus φ6. EMBO J. 16, 4477– 4487 (1997).
Cheng,R. H. et al. Fungal virus capsids, cytoplasmic compartments for the replication of double-stranded RNA, formed as icosahedral shells of asymmetric Gag dimers. J. Mol. Biol. 244, 255– 258 (1994).
Shaw,A. L., Samal,S. K., Subramanian, K. & Prasad,B. V. V. The structure of aquareovirus shows how the different geometries of the two layers of capsid are reconciled to provide symmetrical interactions and stabilization. Structure 4, 957– 967 (1996).
Zhang,H. et al. Visualization of protein–RNA interactions in cytoplasmic polyhedrosis virus. J. Virol. 73, 1624– 1629 (1999).
Prasad,B. V., Wang,G. J., Clerx,J. P. & Chiu,W. Three-dimensional structure of rotavirus. J. Mol. Biol. 199, 269–275 (1988).
Coombs,K. M., Fields,B. N. & Harrison, S. C. Crystallization of the reovirus type 3 Dearing core. J. Mol. Biol. 215, 1–5 (1990).
Otwinowski,Z. & Minor,W. in Macromolecular Crystallography A (eds Carter, C. W. & Sweet, R. M.) 307–326 (Academic, New York, 1997).
CCP4. The CCP4 suite: programs for X-ray crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jones,T. A. in Molecular Replacement (eds Dodson, E. J., Gover, S. & Wolf, W.) 91–105 (SERC Daresbury Laboratory, Warrington, 1992).
Kleywegt,G. J. & Jones,T. A. in From First Map to Final Model (eds Bailey, S., Hubbard, R. & Waller, D.) 59–66 (SERC Daresbury Laboratory, Warrington, 1994).
Jones,T. A., Zou,J. Y., Cowan,S. W. & Kjelgaard,M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger,A. T. et al. Crystallography & NMR System: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jacobsen,D. H., Hogle,J. M. & Filman, D. J. A pseudo-cell based approach to efficient crystallographic refinement of viruses. Acta Crystallogr. D 52, 693–711 (1996).
Laskowski,R. A., MacArthur,M. W., Moss,D. S. & Thornton,J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283– 291 (1993).
Carson,M. in Macromolecular Enzymology (eds Carter, C. W. Jr & Sweet, R. M.) 493–505 (Academic, New York, 1996).
Esnouf,R. M. An extensively modified version of Molscript that includes greatly enhanced coloring capabilities. J. Mol. Graphics 15, 133–138 (1997).
Merrit,E. A. & Murphy,M. E. P. Raster3D version 2.0. A program for photorealistic molecular graphics. Acta Crsytallogr. D 50, 869–873 (1994).
Nicholls,A., Sharp,K. A. & Honig,B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Harrison,S. J. et al. Mammalian reovirus L3 gene sequences and evidence for a distinct amino-terminal region of the lambda1 protein. Virology 258, 54–64 (1999).
Acknowledgements
We thank T. F. Severson, R. L. Margraf and S. J. Harrison for growing reovirus-infected cells; T. S. Baker and S. B. Walker for providing us with an unpublished cryoEM reconstruction of cores from reovirus reassortant F18; D. Thiel and the staff of CHESS and McCHESS and R. Pahl and the staff of the BioCars beamline 14-ID-B for assistance; members of the Harrison laboratory, especially J. Cruzan, for scanning image plates; W. Minor for help with data collected with an early version of the double image plate cassette holder; B. Miller for constructing an improved vacuum-based version; S. Ray for advice on computing strategies; P. Adams and R. Grosse-Kunstleve for advice on refinement; and R. Grosse-Kunstleve for implementing refinement in the smaller rhombohedral cell. K.M.R. thanks D. W. Rodgers for help in the initial cryopreservation attempts, for suggesting a vacuum-based design for the double image plate cassette holder and for advice. We acknowledge support from the NIH (to S.C.H. and M.L.N.). S.C.H. is an investigator in the Howard Hughes Medical Institute. M.L.N. receives support from a grant of the Lucille P. Markey Charitable Trust to the Institute for Molecular Virology and, as a Shaw Scientist, from the Milwaukee Foundation. This work was started with the stimulus and collaboration of the late B. N. Fields.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reinisch, K., Nibert, M. & Harrison, S. Structure of the reovirus core at 3.6?Å resolution. Nature 404, 960–967 (2000). https://doi.org/10.1038/35010041
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35010041
This article is cited by
-
Reovirus directly engages integrin to recruit clathrin for entry into host cells
Nature Communications (2021)
-
Assembly intermediates of orthoreovirus captured in the cell
Nature Communications (2020)
-
Hierarchical structure assembly model of rice dwarf virus particle formation
Biophysical Reviews (2018)
-
Structure and function of S9 segment of grass carp reovirus Anhui strain
VirusDisease (2017)
-
In situ structures of the segmented genome and RNA polymerase complex inside a dsRNA virus
Nature (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.