Abstract
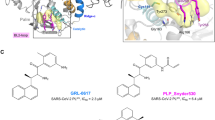
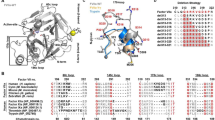
The serpins have evolved to be the predominant family of serine-protease inhibitors in man1,2. Their unique mechanism of inhibition involves a profound change in conformation3, although the nature and significance of this change has been controversial. Here we report the crystallographic structure of a typical serpin–protease complex and show the mechanism of inhibition. The conformational change is initiated by reaction of the active serine of the protease with the reactive centre of the serpin. This cleaves the reactive centre, which then moves 71 Å to the opposite pole of the serpin, taking the tethered protease with it. The tight linkage of the two molecules and resulting overlap of their structures does not affect the hyperstable serpin, but causes a surprising 37% loss of structure in the protease. This is induced by the plucking of the serine from its active site, together with breakage of interactions formed during zymogen activation4. The disruption of the catalytic site prevents the release of the protease from the complex, and the structural disorder allows its proteolytic destruction5,6. It is this ability of the conformational mechanism to crush as well as inhibit proteases that provides the serpins with their selective advantage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Potempa, J., Korzus, E. & Travis, J. The serpin superfamily of proteinase inhibitors: structure, function, and regulation. J. Biol. Chem. 269, 15957–15960 (1994).
Laskowski, M. & Qasim, M. A. What can the structures of enzyme-inhibitor complexes tell us about the structures of enzyme substrate complexes? Biochim. Biophys. Acta 1477, 324–337 (2000).
Loebermann, H., Tokuoka, R., Deisenhofer, J. & Huber, R. Human alpha 1-proteinase inhibitor. Crystal structure analysis of two crystal modifications, molecular model and preliminary analysis of the implications for function. J. Mol. Biol. 177, 531– 557 (1984).
Huber, R. & Bode, W. Structural basis of the activation and action of trypsin. Acc. Chem. Res. 11, 114–122 (1978).
Kaslik, G., Patthy, A., Balint, M. & Graf, L. Trypsin complexed with alpha 1-proteinase inhibitor has an increased structural flexibility. FEBS Lett. 370, 179–183 (1995).
Stavridi, E. S. et al. Structural change in alpha-chymotrypsin induced by complexation with alpha 1-antichymotrypsin as seen by enhanced sensitivity to proteolysis. Biochemistry 35, 10608– 10615 (1996).
Huber, R. & Carrell, R. W. Implications of the three-dimensional structure of alpha 1-antitrypsin for structure and function of serpins. Biochemistry 28, 8951–8966 (1989).
Elliott, P. R., Abrahams, J. P. & Lomas, D. A. Wild-type alpha 1-antitrypsin is in the canonical inhibitory conformation. J. Mol. Biol. 275, 419–425 (1998).
Wright, H. T. & Scarsdale, J. N. Structural basis for serpin inhibitor activity. Proteins 22, 210– 225 (1995).
Stratikos, E. & Gettins, P. G. Formation of the covalent serpin–proteinase complex involves translocation of the proteinase by more than 70 A and full insertion of the reactive center loop into beta-sheet A. Proc. Natl Acad. Sci. USA 96, 4808–4813 (1999).
Fa, M. et al. The structure of a serpin-proteinase complex revealed by intramolecular distance measurements using donor-donor energy migration and mapping of interaction sites. Structure 8, 397– 405 (2000).
Picard, V., Marque, P. E., Paolucci, F., Aiach, M. & Le Bonniec, B. F. Topology of the stable serpin–protease complexes revealed by an autoantibody that fails to react with the monomeric conformers of antithrombin. J. Biol. Chem. 274, 4586–4593 (1999).
Carrell, R. W. & Owen, M. C. Plakalbumin, alpha 1-antitrypsin, antithrombin and the mechanism of inflammatory thrombosis. Nature 317, 730–732 (1985).
Stein, P. E. et al. Crystal structure of ovalbumin as a model for the reactive centre of serpins. Nature 347, 99– 102 (1990).
Engh, R. A., Huber, R., Bode, W. & Schulze, A. J. Divining the serpin inhibition mechanism: a suicide substrate ‘spring’? Trends Biotechnol. 13, 503–510 (1995).
Gooptu, B. et al. Inactive conformation of the serpin alpha(1)-antichymotrypsin indicates two-stage insertion of the reactive loop: implications for inhibitory function and conformational disease. Proc. Natl Acad. Sci. USA 97, 67–72 ( 2000).
Herve, M. & Ghelis, C. Conformational stability of the covalent complex between elastase and alpha 1-proteinase inhibitor. Arch. Biochem. Biophys. 285, 142–146 (1991).
Olson, S. T. et al. Role of the catalytic serine in the interactions of serine proteinases with protein inhibitors of the serpin family. Contribution of a covalent interaction to the binding energy of serpin–proteinase complexes. J. Biol. Chem. 270, 30007– 30017 (1995).
Plotnick, M. I., Mayne, L., Schechter, N. M. & Rubin, H. Distortion of the active site of chymotrypsin complexed with a serpin. Biochemistry 35, 7586–7590 (1996).
Gils, A. & Declerck, P. J. Structure-function relationships in serpins: current concepts and controversies. Thromb. Haemost. 80, 531–541 ( 1998).
Harrop, S. J. et al. The crystal structure of plasminogen activator inhibitor 2 at 2. 0 Å resolution: implications for serpin function. Struct. Fold Des. 7, 43–54 (1999).
Holmes, W. E. et al. Alpha 2-antiplasmin Enschede: alanine insertion and abolition of plasmin inhibitory activity. Science 238, 209–211 (1987).
Bode, W. & Huber, R. Structural basis of the endoproteinase-protein inhibitor interaction. Biochim. Biophys. Acta 1477, 241–252 (2000).
Leslie, A. W. G. in Joint CCP4 and ESF-EACMB Newsletter on Protein Crystallography vol. 26 (Daresbury Laboratory, Warrington, 1992 ).
Evans, P. R. in Proceedings of the CCP4 Study Weekend. Data Collection and Processing (eds Sawyer, L., Isaacs, N. & Bailey, S.) 114–122 (Daresbury Laboratory, 1993).
Collaborative Computational Project Number 4. The CCP4 suite: programs for protein crystal- lography. Acta Crystallogr. 50, 760–763 (1994).
Navaza, J. AMORE - An automated package for molecular replacement. Acta Cryst. A 50, 157–163 ( 1994).
Engh, R. et al. The S variant of human alpha 1-antitrypsin, structure and implications for function and metabolism. Protein Eng. 2, 407–415 (1989).
Lee, S. L. New inhibitors of thrombin and other trypsin-like proteases: hydrogen bonding of an aromatic cyano group with a backbone amide of the P1 binding site replaces binding of a basic side chain. Biochemistry 36, 13180–13186 (1997).
Colman, P. M., Fehlhammer, H. & Bartles, K. in Crystallographic Computing Techniques (eds Ahmed, F. R., Huml, K. & Sedlacek, B.) 248–258 (Munksgaard, Copenhagen, 1976).
Read, R. J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Cryst. A 42, 140– 149 (1986).
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Cryst. D 54, 905–921 (1998).
Pannu, N. S. & Read, R. J. Improved structure refinement through maximum likelihood. Acta Cryst. A 52, 659 –668 (1996).
Kraulis, P. J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946 –950 (1991).
Esnouf, R. Further additions to MolScript version 1.4, including reading and contouring of electron-density maps. Acta Crystallogr. D 55, 938–940 (1999).
Merritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Acknowledgements
We thank our colleagues, N. Pannu for advice throughout; D. Lomas for reading the paper; A. Lesk and P. Stein for discussions; and K. Belzar for support. This work was supported by grants from the Wellcome Trust, the European Community and the National Institutes of Health (J.A.H.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huntington, J., Read, R. & Carrell, R. Structure of a serpin–protease complex shows inhibition by deformation . Nature 407, 923–926 (2000). https://doi.org/10.1038/35038119
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35038119
This article is cited by
-
Probing of the reactive center loop region of alpha-1-antitrypsin by mutagenesis predicts new type-2 dysfunctional variants
Cellular and Molecular Life Sciences (2024)
-
Plasminogen activator inhibitor 1 is not a major causative factor for exacerbation in a mouse model of SARS-CoV-2 infection
Scientific Reports (2023)
-
Neuroserpin, a crucial regulator for axogenesis, synaptic modelling and cell–cell interactions in the pathophysiology of neurological disease
Cellular and Molecular Life Sciences (2022)
-
Proteomic and molecular dynamic investigations of PTM-induced structural fluctuations in breast and ovarian cancer
Scientific Reports (2021)
-
Deep mutational scanning of the plasminogen activator inhibitor-1 functional landscape
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.