Abstract
Salicylic acid (SA) mediates plant defences against pathogens, accumulating in both infected and distal leaves in response to pathogen attack1,2,3,4,5. Pathogenesis-related gene expression and the synthesis of defensive compounds associated with both local and systemic acquired resistance (LAR and SAR) in plants require SA. In Arabidopsis, exogenous application of SA suffices to establish SAR, resulting in enhanced resistance to a variety of pathogens. However, despite its importance in plant defence against pathogens, SA biosynthesis is not well defined. Previous work has suggested that plants synthesize SA from phenylalanine6,7,8,9,10; however, SA could still be produced when this pathway was inhibited6,8, and the specific activity of radiolabelled SA in feeding experiments was often lower than expected7,8. Some bacteria such as Pseudomonas aeruginosa synthesize SA using isochorismate synthase (ICS) and pyruvate lyase11. Here we show, by cloning and characterizing an Arabidopsis defence-related gene (SID2) defined by mutation, that SA is synthesized from chorismate by means of ICS, and that SA made by this pathway is required for LAR and SAR responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dempsey, D. M. A., Shah, J. & Klessig, D. F. Salicylic acid and disease resistance in plants. Crit. Rev. Plant Sci. 18, 547–575 (1999).
Ryals, J. A. et al. Systemic acquired resistance. Plant Cell 8, 1809–1819 (1996).
Malamy, J., Carr, J. P., Klessig, D. F. & Raskin, I. Salicylic acid: a likely endogenous signal in the resistance response of tobacco to viral infection. Science 250, 1002–1004 (1990).
Metraux, J.-P. et al. Increase in salicylic acid at the onset of systemic acquired resistance in cucumber. Science 250, 1004–1006 (1990).
Dorey, S. et al. Spatial and temporal induction of cell death, defense genes, and accumulation of salicylic acid in tobacco leaves reacting hypersensitively to a fungal glycoprotein elicitor. Mol. Plant Microbe Interact. 10, 646–655 (1997).
Mauch-Mani, B. & Slusarenko, A. J. Production of salicylic acid precursors is a major function of phenylalanine ammonia-lyase in the resistance of Arabidopsis to Peronospora parasitica. Plant Cell 8, 203–212 (1996).
Yalpani, N., Leon, J., Lawton, M. A. & Raskin, I. Pathway of salicylic acid biosynthesis in healthy and virus-inoculated tobacco. Plant Physiol. 103, 315–321 (1993).
Coquoz, J.-L., Buchala, A. & Metraux, J.-P. The biosynthesis of salicylic acid in potato plants. Plant Physiol. 117, 1095–1101 (1998).
Ribnicky, D. M., Shulaev, V. & Raskin, I. Intermediates of salicylic acid biosynthesis in tobacco. Plant Physiol. 118, 565–572 (1998).
Leon, J., Shulaev, V., Yalpani, N., Lawton, M. A. & Raskin, I. Benzoic acid 2-hydroxylase, a soluble oxygenase from tobacco, catalyzes salicylic acid biosynthesis. Proc. Natl Acad. Sci. USA 92, 10413–10417 (1995).
Serino, L. et al. Structural genes for salicylate biosynthesis from chorismate in Pseudomonas aeruginosa. Mol. Gen. Genet. 249, 217–228 (1995).
Uknes, S. et al. Biological induction of systemic acquired resistance in Arabidopsis. Mol. Plant Microbe Interact. 6, 692–698 (1993).
Maleck, K. et al. The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nature Genet. 26, 403–410 (2000).
Nawrath, C. & Metraux, J.-P. Salicylic acid induction-deficient mutants of Arabidopsis express PR-2 and PR-5 and accumulate high levels of camalexin after pathogen inoculation. Plant Cell 11, 1393–1404 (1999).
Dewdney, J. et al. Three unique mutants of Arabidopsis identify eds loci required for limiting growth of a biotrophic fungal pathogen. Plant J. 24, 205–218 (2000).
van Tegelen, L. J. P., Moreno, P. R. H., Croes, A. F., Verpoorte, R. & Wullems, G. J. Purification and cDNA cloning of isochorismate synthase from elicited cell cultures of Catharanthus roseus. Plant Physiol. 119, 705–712 (1999).
Cao, H., Bowling, S. A., Gordon, A. S. & Dong, X. Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 6, 1583–1592 (1994).
Delaney, T., Friedrich, L. & Ryals, J. Arabidopsis signal transduction mutant defective in chemically and biologically induced disease resistance. Proc. Natl Acad. Sci. USA 92, 6602–6606 (1995).
Cao, H., Glazebrook, J., Clarke, J. D., Volko, S. & Dong, X. The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 88, 57–63 (1997).
Ryals, J. et al. The Arabidopsis NIM1 protein shows homology to the mammalian transcription factor inhibitor I kappa B. Plant Cell 9, 425–439 (1997).
Despres, J., DeLong, C., Glaze, S., Liu, E. & Fobert, P. R. The Arabidopsis NPR1/NIM1 protein enhances the DNA binding activity of a subgroup of the TGA family of bZIP transcription factors. Plant Cell 12, 279–290 (2000).
Bowling, S. A. et al. A mutation in Arabidopsis that leads to constitutive expression of systemic acquired resistance. Plant Cell 6, 1845–1857 (1994).
Bowling, S. A., Clarke, J. D., Liu, Y., Klessig, D. F. & Dong, X. The cpr5 mutant of Arabidopsis expresses both NPR1-dependent and NPR1-independent resistance. Plant Cell 9, 1573–1584 (1997).
Clarke, J. D., Liu, Y., Klessig, D. F. & Dong, X. Uncoupling PR gene expression from NPR1 and bacterial resistance: characterization of the dominant Arabidopsis cpr6-1 mutant. Plant Cell 10, 557–569 (1998).
Eulgem, T., Rushton, P. J., Robatzek, S. & Somssich, I. E. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 5, 199–206 (2000).
Lebel, E. et al. Functional analysis of regulatory sequences controlling PR-1 gene expression in Arabidopsis. Plant J. 16, 223–233 (1998).
Yang, Y. & Klessig, D. F. Isolation and characterization of a tobacco mosaic virus-inducible myb oncogene homolog from tobacco. Proc. Natl Acad. Sci. USA 93, 14972–14977 (1996).
Bender, J. & Fink, G. R. A Myb homologue, ATR1, activates tryptophan gene expression in Arabidopsis. Proc. Natl Acad. Sci. USA 95, 5655–5660 (1998).
Niyogi, K. K. & Fink, G. R. Two anthranilate synthase genes in Arabidopsis: defense-related regulation of tryptophan pathway. Plant Cell 4, 721–733 (1992).
Reuber, T. L. et al. Correlation of defense gene induction defects with powdery mildew susceptibility in Arabidopsis enhanced disease susceptibility mutants. Plant J. 16, 473–485 (1998).
Acknowledgements
We thank S. Volko for RNA from leaves infected with Pseudomonas pathovar maculicola strain ES4326, C. Nawrath for the sid2-1 seeds, and D. Lee, J. Stone, J. Dangl and E. Meyerowitz for comments on drafts of the manuscript. This work was supported by a National Institutes of Health grant to F.M.A. and a National Research Initiative Competitive Grants Program/USDA postdoctoral fellowship to M.C.W.
Author information
Authors and Affiliations
Corresponding author
Supplementary information

Supplemental Figure 5
(JPG 23.48 KB)
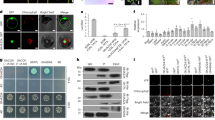
Northern blot analysis of total RNA from Erysiphe-infected and uninfected leaves of wild type or mutant Arabidopsis plants. Duplicate samples are shown from a representative experiment. Each experiment was performed at least twice. Experiments were performed as described in Methods. ICS1/UBQ for npr1 averaged 1.5x of wild type for the experiment shown. In additional experiments, ICS1/UBQ for npr1 averaged >2x above wild type for each experiment. Note that PR1 was not detected in Erysiphe-infected leaves of sid2 or nahG plants, but low levels of PR1 were consistently observed in the npr1 mutant.
Rights and permissions
About this article
Cite this article
Wildermuth, M., Dewdney, J., Wu, G. et al. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414, 562–565 (2001). https://doi.org/10.1038/35107108
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/35107108
This article is cited by
-
Decoding early stress signaling waves in living plants using nanosensor multiplexing
Nature Communications (2024)
-
The OsICS1 is directly regulated by OsWRKY6 and increases resistance against Xanthomonas oryzae pv. oryzae
Planta (2024)
-
The expression of the NPR1-dependent defense response pathway genes in Persea americana (Mill.) following infection with Phytophthora cinnamomi
BMC Plant Biology (2023)
-
Jasmonate signaling drives defense responses against Alternaria alternata in chrysanthemum
BMC Genomics (2023)
-
Cotton microbiome profiling and Cotton Leaf Curl Disease (CLCuD) suppression through microbial consortia associated with Gossypium arboreum
npj Biofilms and Microbiomes (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.