Abstract
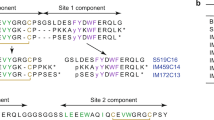
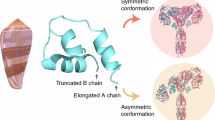
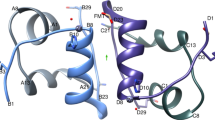
CRYSTAL structures of insulin have been determined in various distinct forms1–5, the relevance of which to receptor recognition has long been the subject of speculation2,6,7. Recently the crystal structure of an inactive insulin analogue has been determined and, surprisingly, found to have a conformation identical to native insulin8,9. On this basis Dodson and colleagues have suggested that the known insulin crystal structures reflect an inactive conformation, and that a change in conformation is required for activity—specifically, the carboxy terminal residues of the B-chain are proposed to separate from the amino terminal residues of the A-chain8. Here we report the solution structure of an active insulin mutant7,10, determined by two-dimensional NMR, which supports this hypothesis. In the mutant, the carboxy terminal β-turn and β-strand of the B-chain are destabilized and do not pack across the rest of the molecule. We suggest that analogous detachment of the carboxy terminal region of the B-chain occurs in native insulin on binding to its receptor. Our finding that partial unfolding of the B-chain exposes an alternative protein surface rationalizes the receptor-binding properties of a series of anomalous insulin analogues7,11–13, including a mutant insulin associated with diabetes mellitus in man14,15.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chothia, C., Lesk, A. M., Dodson, G. G. & Hodgkin, D. C. Nature 302, 500–505 (1983).
Baker, E. N. et al. Phil. Trans. R. Soc. B319, 389–456 (1988).
Bentley, G., Dodson, E., Dodson, G., Hodgkin, D. & Mercola, D. Nature 261, 166–168 (1976).
Badger, J. et al. Acta crystallogr. B47, 127–136 (1991).
Derewenda, U. et al. Nature 338, 594–596 (1989).
Dodson, G. G., Hubbard, R. E. & Reynolds, C. D. Biopolymers 22, 281–292 (1983).
Mirmira, R. & Tager, H. S. J. biol. Chem. 264, 6349–6354 (1989).
Derewenda, U. et al. J. molec. Biol. 220, 425–433 (1991).
Markussen, J. Int. J. Pept. Protein Res. 26, 70–77 (1985).
Shoelson, S. E., Lu, Z-X., Parlautan, L., Lynch, C. S. & Weiss, M. A. Biochemistry (submitted).
Kobayashi, M. et al. Biochem. biophys. Res. Commun. 107, 329–336 (1982).
Nakagawa, S. H. & Tager, H. S. J. biol. Chem. 261, 7332–7341 (1986).
Casaretto, M. et al. Biol. Chem. Hoppe-Seyler 368, 709–716 (1987).
Shoelson, S. E. et al. Nature 302, 540–542 (1983).
Nanjo, A. C. T. et al. J. clin. Invest. 77, 514–519 (1986).
Brown, H., Sanger, F. & Kitai, R. Biochem. J. 60, 556–565 (1955).
Brange, J. et al. Nature 333, 679–682 (1988).
Steiner, D. F., Bell, G. I. & Tager, H. S. in Endocrinology (ed. DeGroot, L. J.) 1263–1289 (Saunders, Philadelphia, 1990).
Hua, Q. X. & Weiss, M. A. Biochemistry 30, 5505–5515 (1991).
Pullen, R. A. et al. Nature 259, 369–373 (1976).
Wuthrich, K. Meth. Enzym. 177, 125–136 (1989).
Clore, G. M. & Gronenborn, A. M. Crit. Rev. Biochem. molec. Biol. 24, 479–563 (1989).
Bi, R. C. et al. Biopolymers 23, 391–395 (1984).
Dai, J.-B., Lou, M.-Z., You, J.-M. & Liang, D.-C. Sci. China 30, 55–65 (1987).
Wang, S.-h., Hu, S.-q., Burke, G. T. & Katsoyannis, P. J. Protein Chem. 10, 313–324 (1991).
Weiss, M. A., Hua, Q. X., Lynch, C. L., Frank, B. H. & Shoelson, S. E. Biochemistry 30, 7373–7389 (1991).
Hua, Q. X., Shoelson, S. E. & Weiss, M. A. J. molec. Biol. (submitted).
Kline, A. D. & Justice, R. M. Jr Biochemistry 29, 2906–2913 (1990).
Boelens, R., Ganadu, M. L., Verheyden, P. & Kaptein, R. Eur. J. Biochem. 191, 147–153 (1990).
Kristensen, S. M., Jorgensen, A. M., Led, J. J., Balschmidt, P. & Hansen, F. B. J. molec. Biol. 218, 221–231 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hua, Q., Shoelson, S., Kochoyan, M. et al. Receptor binding redefined by a structural switch in a mutant human insulin. Nature 354, 238–241 (1991). https://doi.org/10.1038/354238a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/354238a0
This article is cited by
-
Molecular Docking, Dynamics, and WaterSwap Analysis to Identify Anti-aggregating Agents of Insulin and IFN-β
Applied Biochemistry and Biotechnology (2022)
-
Insulin und sein Rezeptor – Spezifität durch Kombinatorik?
BIOspektrum (2020)
-
Insulin in motion: The A6-A11 disulfide bond allosterically modulates structural transitions required for insulin activity
Scientific Reports (2017)
-
A molecular modeling study on full-length insulin: insight into initial events of amyloid formation
Structural Chemistry (2014)
-
How insulin engages its primary binding site on the insulin receptor
Nature (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.