Abstract
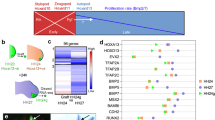
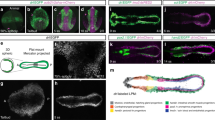
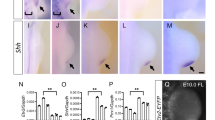
THE positional cues that govern the fate of cells along the dorsoven-tral axis of the developing vertebrate limb are established in the mesoderm before outgrowth of limb buds. In Drosophila, a LIM/-homeodomain gene, apterous, expressed in the dorsal compartment of the wing disc, specifies dorsal cell fate1,2. Here we report the isolation of a vertebrate LIM-homeodomain containing gene, Chick Lmxl (C-Lmxl). Transcripts for C-Lmxl are detected in the presumptive dorsal limb mesoderm and are restricted thereafter to the dorsal mesoderm of the developing chick bud. C-Lmxl expression is regulated by the overlying ectoderm where Wnt7a messenger RNA is localized3. Wnt7a, required for normal development of the dorsoventral axis in mouse limbs4, can induce ectopic expression of C-Lmxl in ventral mesoderm. Misexpression of C-Lmxl during limb outgrowth causes ventral to dorsal transformations of limb mesoderm. We propose that C-Lmxl specifies dorsal cell fate during chick limb development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Cohen, B., McGuffin, M. E., Pfeifle, C., Segal, D. & Cohen, S. M. Genes Dev. 6, 715–729 (1992).
Diaz-Benjumea, F. J. & Cohen, S. M. Cell 75, 741–752 (1993).
Dealy, C. N., Roth, A., Ferrari, D., Brown, A. M. C. & Kosher, R. A. Mech. Dev. 43, 175–186 (1993).
Parr, B. A. & McMahon, A. P. Nature 374, 350–353 (1995).
German, M. S., Wang, J., Chadwick, R. B. & Rutter, W. J. Genes Dev. 6, 2165–2176 (1992).
Logan, C. et al. Devl. Genet. 13, 345–358 (1992).
Hamburger, V. J. exp. Zool. 77, 379–400 (1938).
Dhouailly, D. & Kieny, M. Devl Biol. 28, 162–175 (1972).
Saunders, J. W. Jr & Reuss, C. Devl Biol. 38, 41–50 (1974).
Kieny, M. J. Embryol. exp. Morph. 8, 457–467 (1960).
Pautou, M. P. & Kieny, M. C.r. hebd. Séanc. Acad. Sci., Paris D277, 1225–1228 (1973).
MacCabe, J. A., Errick, J. & Saunders, J. W. Jr Devl Biol. 39, 69–82 (1974).
MacCabe, J. A., Saunders, J. W. Jr & Pickett, M. Devl Biol. 31, 323–335 (1973).
Klingensmith, J. & Nusse, R. Devl Biol. 166, 396–414 (1994).
Morgan, B. A., Izpisúa Belmonte, J. C., Duboule, D. & Tabin, C. J. Nature 358, 236–239 (1992).
Saunders, G. W., Gasseling, M. T. & Errick, J. E. Devl Biol. 50, 16–25 (1976).
Tickle, C. Curr. Biol. 5, 478–484 (1995).
Duboule, D. Curr. Opin. Genet. Dev. 1, 211–216 (1991).
Duboule, D. & Morata, G. Trends Genet. 10, 358–364 (1994).
Hamburger, V. & Hamilton, H. J. Morph. 88, 49–92 (1951).
Izpisúa Belmonte, J. C., De Robertis, E. M., Storey, K. G. & Stern, C. Cell 74, 645–659 (1993).
Niswander, L., Tickle, C., Vogel, A., Booth, I. & Martin, G. R. Cell 75, 579–587 (1993).
Yang, Y. Z. & Niswander, L. Cell 80, 939–947 (1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vogel, A., Rodriguez, C., Warnken, W. et al. Dorsal cell fate specified by chick Lmxl during vertebrate limb development. Nature 378, 716–720 (1995). https://doi.org/10.1038/378716a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/378716a0
This article is cited by
-
Hnrnpk is essential for embryonic limb bud development as a transcription activator and a collaborator of insulator protein Ctcf
Cell Death & Differentiation (2023)
-
Embryonic muscle splitting patterns reveal homologies of amniote forelimb muscles
Nature Ecology & Evolution (2022)
-
Gene expression changes during the evolution of the tetrapod limb
Biologia Futura (2022)
-
Identification of limb-specific Lmx1b auto-regulatory modules with Nail-patella syndrome pathogenicity
Nature Communications (2021)
-
Nail-patella syndrome
Pflügers Archiv - European Journal of Physiology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.