Abstract
Success of epidermal growth factor receptor (EGFR) targeting agents in different cancer types is related to EGFR gene mutations and/or copy number gains. We investigated the EGFR gene status and protein expression by DNA mutational analysis, fluorescence in situ hybridization (FISH), and immunohistochemistry in tumor tissues from 80 patients with primary and corresponding recurrent ovarian serous carcinomas. The patients were classified into six groups with ascending EGFR gene copy numbers. EGFR amplification and high polysomy (FISH+) was present in a significant fraction of the primary (20%) and recurrent (22%) ovarian carcinomas. On mutational analysis, only one tumor with a silent EGFR mutation was observed, and this was the only carcinoma with high-level amplification. EGFR protein immunoexpression was seen in 28% of primary and 33% of recurrent carcinomas and correlated to amplification in the primary tumors (P=0.003). In recurrent carcinoma, moderate and strong EGFR expression was associated with amplification (P=0.034). These molecular events potentially have impact on the responsiveness to EGFR targeting agents in ovarian cancer.
Similar content being viewed by others
Main
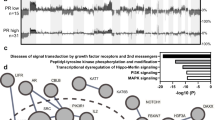
Ovarian cancer is usually diagnosed in advanced stages, which cannot be cured by surgery and adjuvant platinum-based chemotherapy alone. Therefore, recurrences are a major therapeutic challenge and second-line drugs are needed. Antineoplastic drugs that selectively inhibit the epidermal growth factor receptor (EGFR) are currently undergoing clinical trials. Preliminary data on therapy success in ovarian carcinoma are so far controversial. Schilder et al1 reported that one out of 27 patients with the EGFR mutation experienced a long-lasting response to the selective EGFR inhibitor gefitinib, whereas patients without mutations failed to respond. In contrast, Lacroix et al2 reported that the response of patients with platinum-resistant ovarian cancers to gefitinib was independent from EGFR mutational status. Their study revealed a high rate of overall therapy response (63%) associated with the combination of gefitinib and paclitaxel–carboplatin. In other cancers, EGFR-activating mutations were associated with enhanced sensitivity to EGFR inhibitors.3, 4, 5 Some studies showed that gene copy number gains detected by fluorescence in situ hybridization (FISH) were as well important for the success of EGFR targeting therapy.6, 7, 8, 9 In particular, one recent study by Cappuzzo et al9 in lung cancer stratified groups according to gene copy number changes and observed a significant impact on gefitinib response. EGFR-FISH data in ovarian carcinoma are so far not available. We therefore performed a thorough investigation of EGFR in a series of primary ovarian cancers and their corresponding recurrences. This included an analysis of EGFR gene copy number changes according to the analysis method by Cappuzzo et al,9 EGFR mutational analysis and EGFR immunohistochemistry.
Materials and methods
Patients
Primary tumors from 80 patients with advanced ovarian serous carcinoma from the years 1985 to 2003 were collected for this study. All patients had recurrences after receiving primary surgery and at least three cycles of platinum-based adjuvant chemotherapy. We investigated tissue from the primary tumors and the matched recurrences. All tumors were diagnosed at the Institute of Pathology, University of Basel and the Cantonal Hospitals of Baden, Liestal and St Gallen, Switzerland, and were re-examined for this study. The study was performed according to the guidelines of the institutional review boards of the participating institutions. Patient's age ranged from 20 to 77 years (median age of 59 years). The clinical data, including surgical and systemic treatment characteristics, were collected from the medical records. Chemoresistance was defined as recurrence occurring within 6 months after completion of platinum-based chemotherapy.10
Tissue Microarray (TMA)
Tissue samples were fixed in buffered 4% formalin, embedded in paraffin, and used to construct a TMA.11 Briefly, hematoxylin–eosin-stained sections were made from each selected primary block (donor blocks) to define representative tissue regions. Tissue cylinders (0.6 mm in diameter) were then punched from the region of the donor block with the use of a custom-made precision instrument (Beecher Instruments, Silver Spring, MD, USA). Tissue cylinders were transferred to a 25 mm × 35 mm paraffin block to produce the TMA block used for the study. The resulting TMA block was cut into 3 μm sections, which was transferred to glass slides by use of the Paraffin Sectioning Aid System (Instrumedics, Hackensack, NJ, USA). Sections from the TMA block were used for the different analyses.
Immunohistochemistry
Standard indirect immunoperoxidase procedures were used for immunohistochemistry (ABC-Elite, Vectra Laboratories) on TMAs. Immunohistochemistry was performed with a monoclonal antibody against EGFR (clone 2-18C9; PharmDx kit®K 1494, DAKO, Glostrup, Denmark). TMA was analyzed by light microscopy by use of a × 10 objective by two independent pathologists (GSI and SS). Membranous staining in ≥1% of cancer cells was considered positive, according to the EGFR PharmDx scoring guidelines. Appropriate positive and negative controls were used. EGFR immunostaining could be evaluated in 78 primary and 73 recurrent tumors. Analysis failures were owing to either lack of tissue on the TMA section or lack of vital tumor cells in the array samples.
FISH
FISH was carried out as described previously.12 Sections (5 μm) were made from TMA blocks using an adhesive-coated slide system (Instrumedics Inc., Hackensack, NJ, USA). Sections were treated according to the Paraffin Pretreatment Reagent Kit protocol before hybridization (Vysis, IL, USA). FISH was performed with a Spectrum Orange-labeled EGFR gene probe, each in combination with a Spectrum Green-labeled centromere probe for chromosome 7 as reference (Vysis). Hybridization and posthybridization washes were performed according to the ‘LSI procedure’ (Vysis). Slides were then counterstained with 125 ng/ml 4′,6-diamino-2-phenylindole in antifade solution. The signal enumeration was performed under × 400 magnification. The green and red signals were counted in 20 tumor nuclei per tissue sample. FISH could be evaluated in 64 primary and 49 recurrent tumors. Analysis failures were owing to either lack of tissue on the TMA section or lack of vital tumor cells in the array samples.
According to Cappuzzo et al,9 patients were classified into six groups with ascending EGFR gene copy numbers (Table 1). Briefly, disomy was defined as ≤2 copies in >90% of cells, low trisomy as ≤2 copies in ≥40% of cells, 3 copies in 10–40% of the cells, ≥4 copies in <10% of cells, high trisomy as ≤2 copies in ≥40% of cells, 3 copies in ≥40% of cells, ≥4 copies in <10% of cells, low polysomy as ≥4 copies in 10–40% of cells, high polysomy as ≥4 copies in ≥40% of cells, and gene amplification as the presence of EGFR gene clusters and a ratio of EGFR gene to chromosome of ≥2 copies of EGFR per cell in ≥10% of analyzed cells.
Polymerase Chain Reaction (PCR) Analysis
DNA was isolated from paraffin-embedded tissue as described previously.13 Genotyping of EGFR was carried out in exons 18–21 by multiplex PCR analysis with the following primers: forward—EGFRexon18, GCTGAGGTGACCCTTGTCTC, EGFRexon19, TGCCAGTTAACGTCTTCCTTC, EGFRexon20, CATGCGAAGCCACACTGAC and EGFRexon21, CCTCACAGCAGGGTCTTCTC; and reverse—EGFRexon18, GGCCTGTGCCAGGGACCTTA, EGFRexon19, CCACACAGCAAAGCAGAAAC, EGFRexon20, CCGTATCTCCCTTCCCTGAT and EGFRexon21, GGAAAATGCTGGCTGACCTA. For seminested PCR analysis, the following primer sequences were used: forward—EGFRexon18, GAGGTGACCCTTGTCTCTGT, EGFRexon19, AACGTCTTCCTTCTCTCTCTG, EGFRexon20, GCGAAGCCACACTGACGTG and EGFRexon21, CCTCACAGCAGGGTCTTCTC; and reverse—EGFRexon18, GGCCTGTGCCAGGGACCTTA, EGFRexon19, CCACACAGCAAAGCAGAAAC, EGFRexon20, CCGTATCTCCCTTCCCTGAT and EGFRexon21, TGCTGGCTGACCTAAAGCCA.
Data Analysis
Statistical tests included the Wilcoxon signed rank and the Pearson test, and were two-sided. A P-value of ≤0.05 was considered to be statistically significant. Statistical analyses were performed with SPSS 12.0 software (SPSS Inc., Chicago, IL, USA).
Results
EGFR Gene Copy Numbers in Serous Ovarian Carcinoma
Data on FISH analysis in primary and recurrent ovarian cancers are shown in Table 1. According to Cappuzzo et al,9 patients were subdivided into FISH+ (amplification or high polysomy) and FISH− (disomy, low trisomy, high trisomy, and low polysomy) neoplasms. FISH+ were 13 out of 64 (20.3%) primary and 11 out of 49 (22.4%) recurrent neoplasms. There was no significant difference in the frequency of FISH+ and FISH− tumors in the primary and recurrent carcinoma groups. Interestingly, six FISH− tumors out of 64 (9.4%) primary carcinomas were FISH+ in the corresponding recurrent tumors. Reversely, five FISH+ tumors out of 49 (10.2%) primary carcinomas were FISH− in their recurrences. Within the primary and recurrent tumor groups, no significant difference was observed between chemosensitive and chemoresistant neoplasms.
EGFR Mutations in Serous Ovarian Carcinoma
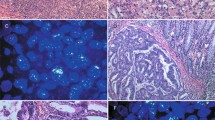
The EGFR gene mutation status was evaluated in 11 tumor specimens that were identified as amplified or as high polysomy, and in three randomly selected FISH− tumors. Only one tumor showed a silent point mutation in exon 20 (codon 787; ACC to ACT), which was also detectable in the recurrent tumor from the same patient. Remarkably, the primary tumor showed a ratio of EGFR gene copy number to chromosome 7 of between 7.5 and 15, with gene clusters, corresponding to the definition of amplification by Cappuzzo et al9 (Figure 1a). The recurrent tumor from the same patient showed >7 copies in >40% of cells, without clustering, corresponding to the definition of high polysomy9 (Figure 1b).
Dual-color FISH with probes for the epidermal growth factor receptor gene (red) and chromosome 7 centromere (green), and EGFR protein immunohistochemical analysis. (a) Primary ovarian carcinoma with high-level amplification, (b) corresponding recurrent tumor with high polysomy, and (c) strong membranous immunoexpression in ovarian carcinoma (magnification × 200).
EGFR Protein Expression in Serous Ovarian Carcinoma
The data on EGFR protein expression in primary and recurrent serous ovarian carcinomas are summarized in Table 1. EGFR protein immunoexpression in primary and recurrent neoplasms was nearly identical (Table 1 and Figure 1c). EGFR protein overexpression was associated with EGFR gene amplification in primary tumors (P=0.003), but was not correlated to high polysomy. In recurrent carcinoma, moderate and strong EGFR expression was associated with EGFR amplification (P=0.034). Regarding the frequency of EGFR protein expression, there was no significant difference between chemosensitive and chemoresistant tumors.
Discussion
The only two studies that investigated EGFR gene mutations in ovarian cancer showed that EGFR-activating mutations are very rare,1, 2 as was also demonstrated in a small number of carcinomas in this study. In the present study, reporting for the first time EGFR FISH data on ovarian carcinoma, tumors defined as FISH+ were observed in a remarkable proportion of over 20% of ovarian serous carcinomas. Remarkably, the only mutation—although silent—that was observed in this study was associated with a high-level gene amplification. EGFR mutations might therefore be a surrogate marker for high-level gene copy gains. Approximately 10% of FISH− and FISH+ primary tumors showed an inversed FISH status in their recurrences. This may be related to a true EGFR heterogeneity within the tumor specimens and/or to a bias related to the TMA technique. Thus, the total frequency of FISH+ tumors may be even higher. As shown in this study, immunohistochemistry is probably not a reliable tool for the detection of all substantial gene copy number changes, in particular high polysomy.
In conclusion, data in this study show that increased EGFR gene copy numbers are observed in a remarkable proportion of ovarian serous carcinomas, whereas mutations appear to be very rare. Based on the importance of copy number gains for EGFR targeting therapy success in other cancer types and their frequency in the present study, our results potentially have an important impact for ovarian carcinoma treatment.
References
Schilder RJ, Sill MW, Chen X, et al. Phase II study of gefitinib in patients with relapsed or persistent ovarian or primary peritoneal carcinoma and evaluation of epidermal growth factor receptor mutations and immunohistochemical expression: a Gynecologic Oncology Group Study. Clin Cancer Res 2005;11:5539–5548.
Lacroix L, Pautier P, Duvillard P, et al. Response of ovarian carcinomas to gefitinib–carboplatin–paclitaxel combination is not associated with EGFR kinase domain somatic mutations. Int J Cancer 2006;118:1068–1069.
Lynch TJ, Bell DW, Sordella R, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004;350:2129–2139.
Paez JG, Janne PA, Lee JC, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004;304:1497–1500.
Pao W, Miller V, Zakowski M, et al. EGF receptor gene mutations are common in lung cancers from ‘never smokers’ and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci USA 2004;101:13306–13311.
Amann J, Kalyankrishna S, Massion PP, et al. Aberrant epidermal growth factor receptor signaling and enhanced sensitivity to EGFR inhibitors in lung cancer. Cancer Res 2005;65:226–235.
Hirsch FR, Varella-Garcia M, Bunn Jr PA, et al. Epidermal growth factor receptor in non-small-cell lung carcinomas: correlation between gene copy number and protein expression and impact on prognosis. J Clin Oncol 2003;21:3798–3807.
Moroni M, Veronese S, Benvenuti S, et al. Gene copy number for epidermal growth factor receptor (EGFR) and clinical response to antiEGFR treatment in colorectal cancer: a cohort study. Lancet Oncol 2005;6:279–286.
Cappuzzo F, Hirsch FR, Rossi E, et al. Epidermal growth factor receptor gene and protein and gefitinib sensitivity in non-small-cell lung cancer. J Natl Cancer Inst 2005;97:643–655.
Jazaeri AA, Awtrey CS, Chandramouli GV, et al. Gene expression profiles associated with response to chemotherapy in epithelial ovarian cancers. Clin Cancer Res 2005;11:6300–6310.
Simon R, Mirlacher M, Sauter G . Tissue microarrays. Methods Mol Med 2005;114:257–268.
Moch H, Sauter G, Gasser TC, et al. EGF-r gene copy number changes in renal cell carcinoma detected by fluorescence in situ hybridization. J Pathol 1998;184:424–429.
Singer G, Schraml P, Belgard C, et al. KIT in ovarian carcinoma: disillusion about a potential therapeutic target. J Natl Cancer Inst 2003;95:1009–1010.
Acknowledgements
We thank PD Dr Beat Thürlimann (Kantonsspital St Gallen), Dr Regula Weber (Kantonsspital Baden), and Dr Ronja Müller (Kantonsspital Liestal) for support in obtaining the clinical data. We thank Mrs H Norotny, Mr A Rufle and Mrs B Stalder for excellent technical assistance. This study was supported by the Swiss Cancer League (Oncosuisse), grant number OCS 01506-02-2004.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stadlmann, S., Gueth, U., Reiser, U. et al. Epithelial growth factor receptor status in primary and recurrent ovarian cancer. Mod Pathol 19, 607–610 (2006). https://doi.org/10.1038/modpathol.3800575
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.3800575
Keywords
This article is cited by
-
High ratio of pCXCR4/CXCR4 tumor infiltrating immune cells in primary high grade ovarian cancer is indicative for response to chemotherapy
BMC Cancer (2022)
-
High density of CD66b in primary high-grade ovarian cancer independently predicts response to chemotherapy
Journal of Cancer Research and Clinical Oncology (2020)
-
Anti-cancer effects of Kaempferia parviflora on ovarian cancer SKOV3 cells
BMC Complementary and Alternative Medicine (2018)
-
High OX40 expression in recurrent ovarian carcinoma is indicative for response to repeated chemotherapy
BMC Cancer (2018)
-
A loop of cancer-stroma-cancer interaction promotes peritoneal metastasis of ovarian cancer via TNFα-TGFα-EGFR
Oncogene (2017)