Abstract
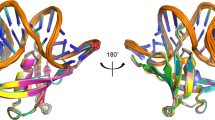
FOR life to be sustained, mistakes in DNA repair must be tolerated when damage obscures the genetic information. In bacteria such as Escherichia coli, DNA damage elicits the well regulated 'SOS response' (reviewed in ref. 1). For the extreme case of damage that cannot be repaired by conventional enzymes, there are proteins that allow the replication of DNA through such lesions, but with a reduction in the fidelity of replication2. Essential proteins in this mutagenic process are RecA, DNA polymerase III, UmuD, UmuD′ and UmuC (umu: UV mutagen-esis)1–3. Regulation of this response involves a RecA-mediated self-cleavage of UmuD to produce UmuD′ To understand this system in more detail, we have determined the crystal structure of the E. coli UmuD′ mutagenesis protein at 2.5 Å resolution. Globular heads folded in an unusual β-structure associate to form molecular dimers, and extended ammo-terminal tails associate to produce crystallized filaments. The structure provides insight into the mechanism of the self-cleavage reaction that UmuD-like proteins undergo as part of the global SOS response4–8.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Friedberg, E. C., Walker, G. C. & Siede, W. DNA Repair ana Mutagenesis (ASM, Washington DC, 1995).
Rajagopalan, M. et al. Proc. natn. Acad. Sci. U.S.A. 89, 10777–10781 (1992).
Woodgate, R. & Sedgwick, S. G. Molec. Microbiol. 6, 2213–2218 (1992).
Little, J. W. Proc. natn. Acad. Sci. U.S.A. 81, 1375–1379 (1984).
Gimble, F. S. & Sauer, R. T. J. molec. Biol. 192, 39–47 (1986).
Shinagawa, H., Iwasaki, H., Kato, T. & Nakata, A. Proc. natn. Acad. Sci. U.S.A. 85, 1806–1810 (1988).
Burckhardt, S. E., Woodgate, R., Scheuermann, R. H. & Echols, H. Proc. natn. Acad. Sci. U.S.A. 85, 1811–1815 (1988).
Nohmi, T., Battista, J. R., Dodson, L. A. & Walker, G. C. Proc. natn. Acad. Sci. U.S.A. 85, 1816–1820 (1988).
Peat, T. S., Frank, E. G., Woodgate, R. & Hendrickson, W. A. Proteins (in the press).
Hendrickson, W. A. Science 254, 51–58 (1991).
Hendrickson, W. A., Horton, J. R. & LeMaster, D. M. EMBO J. 9, 1665–1672 (1990).
Lin, L.-L. & Little, J. W. J. molec. Biol. 210, 439–452 (1989).
Battista, J. R., Ohta, T., Nohmi, T., Sun, W. & Walker, G. C. Proc. natn. Acad. Sci. U.S.A. 87, 7190–7194 (1990).
Roland, K. L., Smith, M. H., Rupley, J. A. & Little, J. W. J. molec. Biol. 228, 395–408 (1992).
Gimble, F. S. & Sauer, R. T. J. molec. Biol. 206, 29–39 (1989).
Little, J. W. J. Bact. 175, 4939–4950 (1993).
Strynadka, N. C. J. et al. Nature 359, 700–705 (1992).
Story, R. M., Weber, I. T. & Steitz, T. A. Nature 355, 318–325 (1992).
Shapiro, L. et al. Nature 374, 327–337 (1995).
Studier, F. W., Rosenberg, A. H., Dunn, J. J. & Dubendorf, J. W. Meth. Enzym. 185, 60–89 (1990).
Otwinowski, Z. in DENZO (eds Sawyer, L, Isaacs, N. & Bailey, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK, 1993).
Collaborative Computational Project no.4 Acta crystallogr. D50, 760–763 (1994).
Debaerdemaeker, T. et al. MULTAN 88 Acta crystallogr. A44, 353–357 (1988).
Brunger, A. T. XPLOR, Version 3.1, A System for X-ray Crystallography and NMR (Yale Univ. Press, New Haven, 1992).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Acta crystallogr. A47, 110–119 (1991).
Kleywegt, G. J. & Jones, T. A. From first Map to Final Model (eds Bailey, S. H. R. & Walker, R.) 59–66 (SERC Daresbury Laboratory, Warrington, UK, 1994).
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. J. appl. Crystallogr. 26, 283–291 (1993).
Evans, S. V. J. molec. Graphics 11, 134–138 (1990).
Kraulis, P. J. J. appl. Crystallogr. 24, 946–950 (1991).
Nicholls, A., Sharp, K. A. & Honig, B. Proteins 11, 281–296 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Peat, T., Frank, E., McDonald, J. et al. Structure of the UmuD′ protein and its regulation in response to DNA damage. Nature 380, 727–730 (1996). https://doi.org/10.1038/380727a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/380727a0
This article is cited by
-
Purification and interactions of the MucA’ and MucB proteins constituting the DNA polymerase RI
Genes and Environment (2019)
-
A tale of two polymers: new insights into helical filaments
Nature Reviews Molecular Cell Biology (2003)
-
Crystal structure of a bacterial signal peptidase in complex with a β-lactam inhibitor
Nature (1998)
-
Dimerization of the UmuD' protein in solution and its implications for regulation of SOS mutagenesis
Nature Structural Biology (1997)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.