Abstract
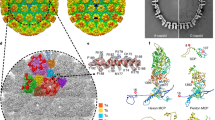
Despite the development of vaccines, the hepatitis B virus remains a major cause of human liver disease1. The virion consists of a lipoprotein envelope surrounding an icosahedral capsid composed of dimers of a 183-residue protein, 'core antigen' (HBcAg)2. Knowledge of its structure is important for the design of antiviral drugs, but it has yet to be determined. Residues 150–183 are known to form a protamine-like domain required for packaging RNA, and residues 1–149 form the 'assembly domain' that polymerizes into capsids2 and, unusually for a capsid protein, is highly α-helical3. Density maps calculated from cryo-electron micrographs4–6 show that the assembly domain dimer is T-shaped: its stem constitutes the dimer interface and the tips of its arms make the polymerization contacts. By refining the procedures used to calculate the map, we have extended the resolution to 9 Å, revealing major elements of secondary structure. In particular, the stem, which protrudes as a spike on the capsid's outer surface, is a 4-helix bundle, formed by the pairing of α-helical hairpins from both subunits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hollinger, F. B. Hepatitis B virus. In Fields Virology 3rd edn (eds Fields, B. N. et al.) 2738–2308 (Lippincott-Raven, Philadelphia, 1996).
Nassal, M. & Schaller, H. Hepatitis B virus nucleocapsid assembly. In Virus Strategies (eds Doefler, W. & Bohm, P.) 41–75 (VCH, Weinheim, 1993).
Wingfield, P. T., Stahl, S. J., Williams, R. W. & Steven, A. C. Hepatitis core antigen produced in Escherichia coli: subunit composition, conformational analysis, and in vitro capsid assembly. Biochemistry 34, 4919–4932 (1995).
Crowther, R. A. et al. Three-dimensional structure of hepatitis B virus core particles determined by electron cryomicroscopy. Cell 77, 943–950 (1994).
Kenney, J. M., von Bonsdorff, C. H., Nassal, M. & Fuller, S. D. Evolutionary conservation in the hepatitis B virus core structure: comparison of human and duck cores. Structure 3, 1009–1019 (1995).
Zlotnick, A. et al. Dimorphism of hepatitis B virus capsids is strongly influenced by the C-terminus of the capsid protein. Biochemistry 35, 7412–7421 (1996).
Zemlin, F. Expected contribution of the field-emission gun to high-resolution transmission electron microscopy. Micron 25, 223–226 (1994).
Conway, J. F. et al. The effects of radiation damage on the structure of frozen hydrated HSV-1 capsids. J. Struct. Biol. 111, 222–233 (1993).
van Heel, M. Similarity measures between images. Ultra microscopy 21, 95–100 (1987).
Winkelmann, D. A., Baker, T. S. & Rayment, I. Three-dimensional structure of myosin subfragment-1 from electron microscopy of sectioned crystals. J. Cell Biol. 114, 701–713 (1991).
Chothia, C. Principles that determine the structure of proteins. Annu. Rev. Biochem. 53, 537–572 (1984).
Presnell, S. R. & Cohen, F. E. Topological distribution of four-alpha-helix bundles. Proc. Natl Acad. Sci. USA 86, 6592–6596 (1989).
Yu, M. S., Miller, R. H., Emerson, S. & Purcell, R. H. A. A hydrophobic heptad repeat of the core protein of woodchuck hepatitis-virus is required for capsid assembly. J. Virol. 70, 7085–7091 (1996).
Salfeld, J., Pfaff, E., Noah, M. & Schaller, H. Antigenic determinants and functional domains in core antigen and e antigen from hepatitis B virus. J. Virol. 63, 798–808 (1989).
Bringas, R. Folding and assembly of hepatitis B virus core protein. A new model proposal. J. Struct. Biol. (in the press).
Rossmann, M. G. & Johnson, J. E. Icosahedral RNA virus structure. Annu. Rev. Biochem. 58, 533–573 (1989).
Bamford, J. K., Bamford, D. H., Li, T. & Thomas, G. J. Jr. Structural studies of the enveloped dsRNA bacteriophage phi 6 of Pseudomonas syringae by Raman spectrosocpy. II. Nucleocapsid structure and thermostability of the virion, nucleocapsid and polymerase complex. J. Mol. Biol. 230, 473–482 (1993).
Momany, C. et al. Crystal structure of dimeric HIV-1 capsid protein. Nature Struct. Biol. 3, 763–770 (1996).
Rao, Z. et al. Crystal structure of SIV matrix antigen and implications for virus assembly. Nature 378, 743–747 (1995).
Arents, G., Burlingame, R. W., Wang, B. C., Love, W. E. & Moudrianakis, E. N. The nucleosomal core histone octamer at 3.1 Å resolution: a tripartite protein assembly and a left-handed superhelix. Proc. Natl Acad. Sci. USA 88, 10148–10152 (1991).
Xie, X. et al. Structural similarity between TAFs and the heterotetrameric core of the histone octamer. Nature 380, 316–322 (1996).
Banner, D. W., Kokkinidis, M. & Tsernoglou, D. Structure of the ColEl rop protein at 1.7 Å resolution. J. Mol. Biol. 196, 657–675 (1987).
Milburn, M. V. et al. A novel dimer configuration revealed by the crystal structure at 2.4 Å resolution of human interleukin-5. Nature 363, 172–176 (1993).
Unwin, P. N. & Henderson, R. Molecular structure determination by electron microscopy of unstained crystalline specimens. J. Mol. Biol. 94, 425–440 (1975).
Jeng, T. W., Crowther, R. A., Stubbs, G. & Chiu, W. Visualization of alpha-helices in tobacco mosaic virus by cryo-electron microscopy. J. Mol. Biol. 205, 251–257 (1989).
Morgan, D. G., Owen, C., Melanson, L. A. & DeRosier, D. J. Structure of bacterial flagellar filaments at 11 Å resolution: packing of the alpha-helices. J. Mol. Biol. 249, 88–110 (1995).
Booy, F. P. et al. High resolution studies of papillomavirus and herpes simplex virus (Abstract). Proc. Xlth Eur. Congr. Microscopy, Dublin (1996).
Baker, T. S. & Cheng, R. H. A model-based approach for determining orientations of biological macromolecules imaged by cryoelectron microscopy. J. Struct. Biol. 116, 120–130 (1996).
Baker, T. S., Drak, J. & Bina, M. The capsid of small papova viruses contains 72 pentameric capsomeres: direct evidence from cryo-electron-microscopy of simian virus 40. Biophys. J. 55, 243–253 (1989).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Conway, J., Cheng, N., Zlotnick, A. et al. Visualization of a 4-helix bundle in the hepatitis B virus capsid by cryo-electron microscopy. Nature 386, 91–94 (1997). https://doi.org/10.1038/386091a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/386091a0
This article is cited by
-
Hepatitis B virus virion secretion is a CRM1-spike-mediated late event
Journal of Biomedical Science (2022)
-
Asymmetrizing an icosahedral virus capsid by hierarchical assembly of subunits with designed asymmetry
Nature Communications (2021)
-
Enhanced stability of a chimeric hepatitis B core antigen virus-like-particle (HBcAg-VLP) by a C-terminal linker-hexahistidine-peptide
Journal of Nanobiotechnology (2018)
-
Solid-state [13C–15N] NMR resonance assignment of hepatitis B virus core protein
Biomolecular NMR Assignments (2018)
-
Cryo-EM reconstruction of the Cafeteria roenbergensis virus capsid suggests novel assembly pathway for giant viruses
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.