Abstract
Spliceosomal introns, one of the hallmarks of eukaryotic genomes, were thought to have originated late in evolution1,2 and were assumed not to exist in eukaryotes that diverged early — until the discovery of a single intron with an aberrant splice boundary in the primitive 'protozoan' Giardia3. Here we describe introns from a close relative of Giardia, Carpediemonas membranifera, that have boundary sequences of the normal eukaryotic type, indicating that canonical introns are likely to have arisen very early in eukaryotic evolution.
Similar content being viewed by others
Main
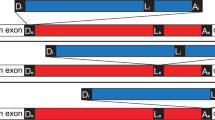
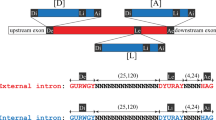
Carpediemonas membranifera is a poorly studied, free-living microbial eukaryote that is considered to be a relative of Giardia on the basis of its morphology4. Using the polymerase chain reaction (PCR) with Carpediemonas genomic DNA as template, we determined the partial sequences of two distinct carbamate kinase genes from this organism. In both genes, an insertion of 33 or 31 nucleotides interrupts the similar protein-coding sequence shared with carbamate kinase genes from other organisms (Fig. 1a). These insertions are bounded by guanine and thymine (GT) nucleotides at the 5′ end and adenine and guanine (AG) nucleotides at the 3′ end, which is a characteristic of most of the spliceosomal introns that interrupt protein-coding genes in other eukaryotes.
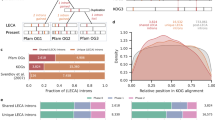
a, Portions of two Carpediemonas carbamate kinase genes, showing intron sequences (red) interrupting the protein-coding sequence (in blue). The introns have canonical splice boundaries (GT...AG; large red type). b, Maximum-likelihood evolutionary tree of eukaryotic cytosolic Hsp70 proteins ('Γ + invariable sites' model). Endoplasmic-reticulum Hsp70 ('BiP') is used as an outgroup. The grouping of Carpediemonas with Giardia and Spironucleus is shown in the blue box; statistical support (bootstrap percentages) for this grouping is assessed using likelihood (upper left of box), and likelihood distance (lower left). The higher percentages (right in each pair) apply when the outgroup is omitted. The basal placement of Trichomonas is weakly supported with likelihood (21%); green arrow shows a more plausible position on the basis of other evidence7,13. The intron splice boundaries for the relevant groups and the origin of canonical introns are shown in red. New sequences have been deposited at GenBank under accession numbers AY131204–AY131209.
We used PCR with reverse transcription to recover the messenger RNA sequence of one of the two Carpediemonas carbamate kinase genes. This sequence lacks the insertion, which is presumably removed (spliced) from the messenger RNA before translation. We conclude that the insertions in the Carpediemonas carbamate kinase genes are canonical 'GT...AG' spliceosomal introns, albeit comparatively small ones.
To determine the evolutionary affinities of Carpediemonas, we used PCR to amplify near-complete sequences for two genes that encode cytosolic heat-shock protein 70 (Hsp70). We also sequenced a cloned Hsp70 gene from Spironucleus barkhanus, a very close relative of Giardia. Maximum likelihood analysis of Hsp70 proteins reveals a specific evolutionary relation between Carpediemonas, Giardia and Spironucleus (Fig. 1b); three other molecular markers also support this relationship5.
The single intron found in a Giardia gene has a non-canonical CT dinucleotide at its 5′ splicing boundary3, which could be interpreted as a 'frozen' primitive eukaryotic condition: canonical 'GT...AG' spliceosomal introns might then be a later innovation in more modern cells. Our results indicate that this is not the case, however, as canonical introns seem to be an ancestral feature of the larger evolutionary grouping that includes Giardia and Carpediemonas. The aberrant Giardia intron probably represents a lineage-specific (or intron-specific) secondary alteration of the 5′ splice boundary.
The extremely early divergence attributed to Giardia is based on the absence or aberration of many typical eukaryotic features, such as mitochondria and introns, and on its arguably deep-branching position in many phylogenetic trees6,7,8,9. The grouping of Giardia with Carpediemonas (which, as well as canonical introns, has organelles that are probably derived from mitochondria4) weakens this argument for early divergence.
Irrespective of the true evolutionary position of Giardia, the only potentially 'early' eukaryotic group in which introns have not been found are the parabasalids, such as Trichomonas10,11. Trichomonas is already known to possess some of the cellular machinery for intron splicing12, however, and there is evidence to indicate that it is evolutionarily affiliated with Giardia and its relatives7,13 (and is slightly misplaced in many phylogenies, including that shown in Fig. 1b). An affiliation with Giardia implies a similar closeness to Carpediemonas, and it is likely that parabasalids have, or had, canonical introns. There is now every reason to assume that canonical introns were present in the most recent common ancestor of living eukaryotes.
References
Palmer, J. D. & Logsdon, J. M. Curr. Opin. Genet. Dev. 1, 470–477 (1991).
Logsdon, J. M. Curr. Opin. Genet. Dev. 8, 637–648 (1998).
Nixon, J. E. J. et al. Proc. Natl Acad. Sci. USA 99, 3701–3705 (2002).
Simpson, A. G. B. & Patterson, D. J. Eur. J. Protistol. 35, 353–370 (1999).
Simpson, A. G. B. et al. Mol. Biol. Evol. 19, 1782–1791 (2002).
Cavalier-Smith, T. Trends Genet. 7, 145–148 (1991).
Embley, T. M. & Hirt, R. P. Curr. Opin. Genet. Dev. 8, 624–629 (1998).
Roger, A. J. Am. Nat. 154 (suppl.), 146–163 (1999).
Sogin, M. L. Curr. Opin. Genet. Dev. 7, 792–799 (1997).
Johnson, P. J. Proc. Natl Acad. Sci. USA 99, 3359–3361 (2002).
Archibald, J. M., O'Kelly, C. J. & Doolittle, W. F. Mol. Biol. Evol. 19, 422–431 (2002).
Fast, N. M., Logsdon, J. M. & Doolittle, W. F. Mol. Biochem. Parasitol. 99, 514–522 (1999).
Dacks, J. B. & Roger, A. J. J. Mol. Evol. 48, 779–783 (1999).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
brief communications is intended to provide a forum for both brief, topical reports of general scientific interest and technical discussion of recently published material of particular interest to non-specialist readers. Priority will be given to contributions that have fewer than 500 words, 10 references and only one figure. Detailed guidelines are available on Nature's website (http://www.nature.com) or on request from nature@nature.com
Rights and permissions
About this article
Cite this article
Simpson, A., MacQuarrie, E. & Roger, A. Early origin of canonical introns. Nature 419, 270 (2002). https://doi.org/10.1038/419270a
Issue Date:
DOI: https://doi.org/10.1038/419270a
This article is cited by
-
Origin and evolution of spliceosomal introns
Biology Direct (2012)
-
A genomic survey of the fish parasite Spironucleus salmonicida indicates genomic plasticity among diplomonads and significant lateral gene transfer in eukaryote genome evolution
BMC Genomics (2007)
-
Patterns of intron gain and conservation in eukaryotic genes
BMC Evolutionary Biology (2007)
-
The evolution of spliceosomal introns: patterns, puzzles and progress
Nature Reviews Genetics (2006)
-
Introns and the origin of nucleus–cytosol compartmentalization
Nature (2006)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.