Abstract
Early signaling in camptothecin-treated MCF-7 cells followed an intrinsic pathway, but death was delayed and late events exhibited few hallmarks of apoptosis. BH3-only proteins, such as Noxa, Puma and BimEL, were activated and localized to mitochondrial sites within 24 h following drug exposure. However, caspase activity was low and death was unaffected by caspase inhibition. Transmission electron micrographs showed the presence of large vacuoles in drug-treated cells. An autophagic survival response has been attributed to MCF-7 cells following nutrient starvation or exposure to tamoxifen. Here, we show that autophagy also plays an important role in the delayed DNA damage response. Confocal microscopy revealed colocalization of mitochondria with large autophagic vacuoles and inhibitors of autophagy increased mitochondrial depolarization and caspase-9 activity, and accelerated cell death. Furthermore, downregulation of autophagy proteins, Beclin 1 and Atg7, unmasked a caspase-dependent, apoptotic response to DNA damage. We propose that a post-mitochondrial caspase cascade is delayed as a result of early disposal of damaged mitochondria within autophagosomes. Our data also suggest that the use of autophagy as a means of delaying apoptosis or prolonging survival may be characteristic of noninvasive breast tumor cells. These studies underscore a potential role for autophagy inhibitors in combination with conventional chemotherapeutic drugs in early breast cancer therapy.
Similar content being viewed by others
Main
A variety of stimuli, including growth factor deprivation, UV or gamma radiation and DNA-damaging agents activate an intrinsic pathway of apoptosis involving the release of cytochrome c (cyt c) and other mitochondrial apoptogenic factors.1 This release promotes the assembly of a large cytoplasmic complex called the ‘apoptosome’ and induces caspase-9 to initiate an apoptotic caspase cascade beginning with the activation of effector caspase-3 and culminating in the death of the cell.2 The term ‘apoptosis’ has been used synonymously with programmed cell death (PCD), but it has become evident in recent years that alternative, nonapoptotic forms of cell suicide are also ‘programmed’. Apoptosis, an ATP dependent process characterized by cell shrinkage, activation of caspases and DNA fragmentation, corresponds to Type I PCD, while autophagic cell death is classified as Type II PCD.3 During autophagy, the cell encapsulates bulk cytoplasm and whole organelles in double-membrane-bound vacuoles or autophagosomes, degrading and recycling the contents by metabolic catabolism.4
Bcl-2 protein family proteins are strategic regulators of the intrinsic pathway exerting their influence on the outer mitochondrial membrane.5 The proapoptotic BH3-only proteins of the Bcl-2 family are components of signal transduction pathways that convey the ‘death’ message to mitochondria. These proteins can be activated in response to a variety of death stimuli, including DNA-damaging compounds.6, 7 Drugs such as camptothecin (CPT) and etoposide, typically activate the mitochondrial pathway by upregulating p53 expression. Apoptosis mediated by p53 involves both transcriptional activation of a number of targets, including Bcl-2 family genes encoding Puma, Noxa, Bim, Bid and Bax, as well as protein–protein interactions with Bcl-2 family members at mitochondrial sites.6, 8, 9
We were interested in identifying intermediates involved in transducing proapoptotic signals following DNA damage in breast cancer cells using the MCF-7 cell line as a model. MCF-7 cells do not express caspase-3 and, although the cells die in response to a variety of apoptotic stimuli such as UV radiation, paclitaxel, tamoxifen and etoposide, studies have reported little detectable processing or activation of initiator caspase-9 during the process.10, 11 The observation that CPT-treated MCF-7 cells exhibit a delayed death response with few phenotypic features of apoptosis prompted further investigation into the underlying mechanisms.
Results
MCF-7 cells mount a delayed death response to CPT that is caspase-independent
MCF-7 cells exposed to 2 μM CPT were analyzed for viability at 24 h intervals by flow cytometry. Annexin V (Ann V) single positive, propidium iodide (PI) single positive, and double-positive populations were collectively counted as ‘dead’, whereas double-negative cells were considered ‘viable’. Cells exhibited a lag period lasting over 24 h in their response to CPT, with significant decrease in viability in 48 h, represented by a strong single-positive PI component indicative of necrosis. The release of mitochondrial cyt c within 24 h of exposure to CPT (Figure 1B), however, suggested that dying cells were engaging the intrinsic pathway. Confocal microscopy of Mitotracker Red-labeled cells (Figure 1C) show a distinct change in mitochondrial morphology over 48 h of CPT exposure from an evenly distributed punctate pattern to one of aggregates in the cytosol. Other hallmarks of apoptosis, however, such as blebbing, discrete DNA fragmentation (not shown) or caspase activation, were absent. The broad-spectrum caspase inhibitor, z-VAD-fmk, did not inhibit CPT-induced death and, in fact, modestly accelerated the death response (Figure 1D). The Western blot in Figure 1E suggests that activation of initiator caspases 8 and 9 and effector caspases 6 and 7 as determined by cleavage of their zymogen forms (MCF-7 cells do not express caspase-3), was low to undetectable 48 h following CPT exposure.
Cell death induced by CPT in the MCF-7 breast cancer cell line is largely caspase-independent. (A) Cells were incubated with 2 μM CPT for the indicated time. Cell death was quantified using flow cytometry as described in Materials and Methods. Results (mean±S.E.) represent the average of three or more experiments. (B) Mitochondrial (top panel) and cytosolic (lower panel) fractions from control and 24 h CPT-treated MCF-7 cells were immunoblotted with cyt c, COX IV and actin antibodies. (C) Confocal images of control MCF-7 cells (a) and cells treated with CPT for 24 h (b) and 48 h (c) showing mitochondria labeled with the fluorescent dye Mitotracker Red observed at 60 × magnification using a Bio-Rad MRC 1024 Single Photon Confocal microscope. (D) Cells were incubated with 2 μM CPT in the presence or absence of 25 μM z-VAD-fmk for the indicated time. Viability was determined by flow cytometric analysis of Ann V and PI uptake. Results (mean±S.E.) represent the average of three experiments. (E) Lysates from control and 48 h CPT-treated MCF-7 cells were resolved by SDS-PAGE and sequentially immunoblotted with antibodies against caspase-9, -8, -7, and -6, followed by GAPDH to control for loading
A small subset of BH3-only transcripts is upregulated in MCF-7 cells following treatment with CPT
The release of mitochondrial cyt c pointed to an intrinsic death pathway. Therefore, the inability of these cells to activate a robust caspase cascade was intriguing. To further investigate this DNA damage response, we examined early signaling events. First, we determined whether BH3-only proteins of the Bcl-2 family were transcriptionally activated by carrying out RNase protection assays (RPAs) on total RNA isolated from cells exposed to 2 μM CPT using multiprobe BH3-only RPA templates.12 Figure 2a shows representative RPAs, and Figure 2b shows quantitative analyses of the observed changes in mRNA expression of some of the proapoptotic BH3-only members. CPT treatment resulted in the induction of at least five different BH3-only transcripts. Transcripts encoding Puma, the three isoforms of Bim (BimEL, BimL and BimS), Bik, Bid and Noxa (not shown) were upregulated; anti-apoptotic Mcl-1L also increased gradually whereas Mcl-1S levels remained constant. Among the transcripts included in probe set 2, only MAP-1 RNA showed any increase whereas expression of BNip3, Bmf, Rad9, Bcl-xL Bcl-xS, Bad and Bcl-GL remained constant.
Modulation of proapoptotic BH3-only member transcripts by CPT in MCF-7 cells. Cells were incubated with CPT for 6, 9, 12, 24, and 48 h before extraction and quantification of total RNA and protein. (a) Representative autoradiograph of an RPA gel from one of three experiments performed as described (10 μg total RNA, per time point) in Materials and Methods. RPAs using multiprobe sets 1 and 2 are shown in the left and right panels, respectively. Labeled multiprobes are in the left-most lane of each panel, and the identities of the probes and protected double-stranded RNAs are marked on the right of each autoradiograph. The protected fragments are shorter than the probes because of the nature of the assay system. An internal loading control template, GAPDH, was included with each template set. (b) Quantification of RNA levels from three RPA analyses. The optical density of each band was normalized relative to the optical density of GAPDH by the following equation: optical density of the mRNA band/optical density of the GAPDH band × 100 and relative RNA levels expressed as arbitrary units. Bars represent the mean (±S.E.) normalized optical densities of mRNA bands (P-value relative to control)
Bcl-2 family protein and p53 expression during the CPT response
Western blots of whole cell lysates from CPT-treated cells (Figure 3a) confirmed the early induction of BH3-only proteins, Puma and Noxa. Bim EL was processed to a shorter form due most likely to dephosphorylation. Phosphorylation of BimEL by ERK1/2 is known to dramatically reduce its ability to bind Bax and prime it for proteasomal degradation.13 Thus BimEL appears to be regulated both transcriptionally and posttranslationally in response to DNA damage. BimS was undetectable and BimL expression was unchanged (not shown). The expression of the other BH3-only proteins showing RNA induction, Bid, Bik and MAP-1, as well as that of the multi-domain antiapoptotic and proapoptotic Bcl-2 family members, Bcl-2, Bcl-xL, Bak and Bax, remained constant over this time period (Figure 3a, and not shown). It should be noted that the Bax antibody used in this experiment recognizes both the inactive and active forms of the protein. Enriched mitochondrial fractions confirmed cyt c release in 24 h CPT-treated MCF-7 cells and showed the accumulation of Puma, Noxa and processed BimEL at the mitochondria (Figure 3b). A proapoptotic function for mitochondrially localized p53, independent of its transcriptional activator function, has been described.14, 15 At the mitochondria, p53 can interact with Bcl-xL as well as Bax; the disruption of a p53/Bcl-xL interaction by BH3-only protein, Puma, allows p53 to interact with Bax and promote cyt c release. Figure 3c shows that Bax is immunoprecipitable only from 24 h CPT-treated mitochondrial fractions with the 6A7 monoclonal antibody that recognizes the active form of the protein (top panel), and that active Bax interacts with the induced p53 protein in this fraction (bottom panel).
Expression of p53 and selected Bcl-2 family members in CPT-treated MCF-7 cell lysates and mitochondria. (a) Western blots of whole-cell lysates (50 μg total protein) harvested at indicated times following CPT exposure. Lysates were resolved on 14% (or 18%) Tris-glycine SDS-polyacrylamide gels and transferred to nitrocellulose. Membranes were sequentially stripped and reprobed (except in the case of the Noxa immunoblot, when an 18% gel was used to resolve the lysate). Actin protein expression (bottom panel) was used as loading control. The actin control is not shown for the Noxa Western. (b) Western blots of mitochondrial fractions from untreated and 24 h CPT-treated MCF-7 cells. Mitochondrially enriched fractions (50 μg total protein) were resolved by SDS-PAGE and transferred to nitrocellulose. Membranes were sequentially stripped and reprobed. Cyt c release (top panel) is an indicator of apoptosis and COX IV expression (second panel) serves both as control for the fractionation procedure as well for loading. (c) Mitochondrially enriched fractions from 24 h CPT-treated and control MCF-7 cells, were lysed in CHAPS lysis buffer. Immunoprecipitation of active Bax from mitochondrial lysate (400 μg) was carried out with anti-Bax 6A7 monoclonal antibody (BD Pharmingen), immunoprecipitates were resolved by SDS-PAGE and immunoblotted with anti-Bax (N-20) polyclonal antibody followed, after stripping, by anti-p53 FL-393 antibody
The induction of p53, transcriptional and translational modulation of selected BH3-only proteins, translocation of p53 to mitochondrial sites, and loss of mitochondrial cyt c were indicative of a prototypical intrinsic response predicted to activate a robust apoptotic caspase cascade initiated by the capase-9 protein. However, data showed instead that cells were dying by a mechanism that was largely caspase independent for at least 48 h (Figure 1). Other groups have noted the lack of an active caspase-9 component in the death response of MCF-7 cells exposed to chemotherapeutic drugs, such as paclitaxel and tamoxifen, and to UV and gamma radiation.10, 11
CPT induces autophagy in MCF-7 cells
The lack of clear apoptotic markers despite substantial evidence for mitochondrial perturbations in MCF-7 cells following CPT exposure was puzzling. Transmission electron microscopy indicated little overt blebbing but revealed the presence of large vacuoles in a number of the CPT-treated cells, suggesting the activation of an autophagic response (Figure 4A). Interesting functional links have been revealed between apoptosis and autophagy. For instance, low levels of mitochondrial membrane permeabilization and depolarization can trigger sequestration and autophagy of damaged mitochondria.16, 17 Recent studies have shown that tamoxifen and nutrient starvation activate autophagy in MCF-7 cells.11, 18, 19 To confirm the presence of autophagosomes, control and CPT-exposed MCF-7 cells were double-labeled with the mitochondria-selective mitotracker green (MTG) dye and with monodansyl cadaverine (MDC), an auto-fluorescent and selective marker for autophagic vacuoles.19, 20 Figure 4B shows aggregated mitochondria labeled with MTG (on left) and red fluorescent vacuolar staining with MDC accumulation (center) in CPT-treated cells. The overlay on the right shows colocalization of a large number of the mitochondria with the autophagic vacuoles. Similarly, GFP-tagged LC321 expressing MCF-7 cells shows that the diffuse GFP fluorescence in control MCF-7-GFP-LC3 cells changes to bright green vacuolar fluorescence often overlapping with using mitotracker red staining following exposure to CPT (Figure 4C). The microtubule-associated protein 1, light chain 3 (MAP-LC3) protein is cytoplasmically localized, but is post-translationally cleaved and lipidated and localized exclusively to autophagosomes during autophagy.21 Quantitative analyses of colocalization by either approach showed that 45–55% of aggregated mitochondria were associated with autophagosomes within 24 h. Figure 4D shows representative fields of 4′-6-diamidino-2-phenylindole (DAPI)-stained GFP-LC3-expressing cells used in quantifying CPT-induced punctate GFP fluorescence (Figure 4E).
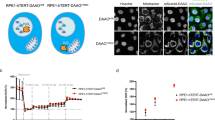
CPT induces autophagy in MCF-7 cells. (A) MCF-7 cells treated with either DMSO or CPT (2 μM) for 30 h were harvested, fixed in 2.5% glutaraldehyde and postfixed in 1% osmium tetroxide. Ultramicrotome sections were poststained and imaged on a JEOL 1200 EX transmission electron microscope at 80 kV. (B) Control (not shown) and 24 h CPT-treated MCF-7 cells were double labeled with 20 nM MTG (left) and 100 μM MDC (center) and observed under a Bio-Rad MRC 1024 Single Photon Confocal microscope at 100 × magnification. Overlay is shown on right. (C) GFP-LC3-expressing MCF-7 cells were treated with DMSO or 2 μM CPT for 24 h, labeled with Mitotracker Red and observed using a confocal microscope. Top panels – DMSO-treated MCF-7-GFP-LC3 cells, bottom panels – 24 h CPT-treated MCF-7-GFP-LC3 cells (a and d, Mitotracker Red; b and e, GFP; c and f, merged). (D and E) MCF-7 cells expressing GFP-LC3 were treated with DMSO or CPT for 24 h, fixed and mounted with DAPI-containing mounting medium and observed under a Zeiss fluorescence microscope (D). The increase in punctate green fluorescence, indicative of autophagy, was quantified using the ‘Analyze particles’ command in the Image J software (E)
Autophagy in mammalian cells as well as yeast is sensitive to inhibition of PI3 kinase activity. Autophagosome precursors are not generated in cells treated with PI3-kinase inhibitors, such as 3-methyl adenine (3-MA) and wortmannin.22 To further determine whether autophagy was contributing to the CPT response, cells were treated with inhibitors of autophagy in the presence of the drug. MCF-7 cells were incubated with either wortmannin, bafilomycin A1 or 3-MA, 4–6 h before harvesting to minimize additional effects on signaling pathways unrelated to the CPT response. All three compounds caused a 5–10% (statistically significant) increase in the dying population of CPT-treated cells (Figure 5a). We also determined the effect of autophagy inhibition on uptake of the mitochondrial membrane potential (ΔpsiM)-sensitive dye and lipophilic cation, tetramethylrhodamine ethyl ester, perchlorate (TMRE), following CPT treatment in the presence and absence of autophagy inhibitors. A few hours of incubation with 3-MA resulted in a 10% increase in the proportion of depolarized mitochondria in CPT-treated cells (Figure 5b). Similar observations were made with wortmannin and bafilomycin A1 (not shown). Caspase-9 activity, as determined using a colorimetric substrate cleavage assay, also increased in 48 h CPT-treated cells that were exposed to 3-MA, although this difference was less discernible at the 24 h time point (Figure 5c).
Inhibitors of autophagy accelerate CPT-induced apoptotic cell death in MCF-7 cells. (a) Cells from exponentially growing and CPT-treated cultures were collected at the times indicated. Inhibitors of autophagy, wortmannin (10 μM), bafilomycin A1 (200 nM) and 3-MA (10 mM) were added to the culture medium 5 h before harvesting. Cell survival was measured by flow cytometric analysis of cells stained with fluorescein-labeled Ann V and PI. Statistical comparisons of mean values were carried out using Student's t-test; P-values ⩽0.05 were considered to be statistically significant. (b) Mitochondrial depolarization following 24 h CPT exposure in the presence or absence of 10 mM 3-MA. MCF-7 cells were incubated with the ΔpsiM-sensitive mitochondrial dye, TMRE (40 nM) for 15 min before harvesting. Profile of unstained, untreated control (black outline) is included with each histogram. (c) Cells were treated with 2 μM CPT in the presence or absence of 3-MA (10 mM) and harvested at the time indicated. Caspase-9 activity levels in cell lysates were determined in triplicate in a colorimetric cleavage assay using LEHD-pNA as substrate
These data suggested that autophagy inhibition was unmasking or accelerating the apoptotic component of the CPT response in the MCF-7 cells. To further validate our observations, we exposed cells knocked down for Beclin 1 expression, to DNA-damaging drugs.
Loss of expression of autophagy proteins unmasks a caspase-dependent apoptotic pathway in DNA-damaged MCF-7 cells
Beclin 1, the mammalian ortholog of Apg6, is a component of the class III PI3-kinase complex involved in autophagic vesicle formation. Beclin 1 overexpression increases the number of autophagic vacuoles and the degradation of long-lived proteins upon amino-acid deprivation in MCF-7 cells.23 This protein is also known to interact with the antiapoptotic protein, Bcl-2.24, 25 It may be noted here that although some groups have referred to MCF-7 as Beclin 1-negative cells, we have consistently detected the expression of endogenous Beclin 1 in MCF-7 Western blots using the sensitive Super Signal West Femto kit (Pierce Biotech). We did not observe increased Beclin 1 expression in CPT-exposed cells, although levels of the protein were reduced in 24 h treated mitochondria suggesting degradation or intracellular translocation (not shown). To determine the effect of Beclin 1 loss on the DNA damage response, its expression in MCF-7 cells was downregulated using small hairpin (sh) RNA constructs. Figure 6a shows a Western blot with three clonal isolates with >80% loss in Beclin 1 expression. Beclin 1low cells expressing GFP-LC3 showed highly reduced punctate green fluorescence compared to controls (Supplementary Figure 1). Next, we determined the response of the Beclin 1low cell lines to DNA damage. Figure 6b shows representative dot plots of PI and Ann V uptake by MCF-7 clone 4 (on the right) and parental MCF-7 cells (on the left) treated with dimethylsulfoxide (DMSO), CPT or etoposide, a DNA-damaging drug that acts by inhibiting topoisomerase II.26 Beclin 1 downregulation substantially increased the number of apoptotic (Ann V positive) cells in response to both CPT and etoposide (similar results were obtained with other Beclin 1low clones). The apoptotic nature of the response further confirmed by its sensitivity to caspase inhibition (Figure 6c) and a 3–4-fold increase in the ability of the Beclin 1low cells to cleave the caspase-9 substrate, LEHD-pNA, within 24 h of CPT exposure (Figure 6d). To confirm that the observed effect of Beclin 1 was not independent of autophagy Atg7, a protein involved in the ubiquitination of MAP-LC3,27, 28 was also downregulated in MCF-7 cells, and shown to activate an apoptotic component following CPT exposure (Figure 7a and b).
Downregulation of autophagy protein Beclin 1 activates apoptotic pathways in MCF-7 cells following DNA damage. (a) MCF-7 cells were transfected with a Beclin 1 shRNA plasmid using the pGSPU6 expression vector and a lipofectamine transfection protocol. The figure shows Western blots of MCF-7 control cells and three clonal isolates from the transfected populations, probed with antibodies against Beclin 1 (top panel), stripped and reprobed for actin expression (bottom panel). (b) Dot plots of Ann V (FL1 channel) and PI (FL2 channel) uptake of control MCF-7 and Beclin 1low Clone 4 cells exposed to DMSO (top) or DNA-damaging agents, CPT (center) and etoposide (bottom) for 24 h. (c) Cells were incubated with 2 μM CPT in the presence or absence of 25 μM z-VAD-fmk for 24 h. Viability was determined by flow cytometric analysis of Ann V and PI uptake. Results (mean±S.E.) represent the average of three data points. (d) Graph showing caspase-9 activity in MCF-7 transfectants, Beclin 1scr and Beclin 1low (clone 4) following exposure to CPT, as determined by colorimetric substrate (LEHD-pNA) cleavage assays. Results expressed as arbitrary units per microgram protein (mean±S.E.) represent the average of three data points
Downregulation of autophagy protein Atg7 activates apoptosis in CPT-treated MCF-7 cells. (a) MCF-7 cells were transfected with an Atg7 shRNA or control (scrambled) RNA plasmid using the pGSPU6 expression vector and a lipofectamine transfection protocol. Figure shows Western blots of MCF-7 control (untransfected and scrambled RNA-transfected) cells and one pooled and two clonal isolates from the Atg7 shRNA-transfected populations, probed with antibodies against Atg7 (top panel), stripped and reprobed for actin expression (bottom panel). (b) Control and Atg7 shRNA cell lines were incubated with either DMSO or CPT (2 μM) for 24 h. Viability was determined by flow cytometric analysis of Ann V and PI uptake
Autophagy delays apoptosis in noninvasive breast cancers
Taken together, the results suggest that CPT activated an intrinsic pathway and early mitochondrial damage, while simultaneously triggering an autophagic response in the cells that prolonged their survival. We were interested in determining whether autophagy inhibition regulated the DNA damage response in diverse breast epithelial cells and cancer lines. A small selection of commercially available immortalized and tumorigenic breast epithelial cell lines were treated with CPT, in the presence and absence of autophagy inhibitors. The estrogen receptor positive (ERpos), MCF-7 breast cancer cells, although derived from a pleural effusion, are noninvasive cells in mouse models.29 We also determined the response to CPT exposure and autophagy inhibition of the T47D cell line which, like MCF-7, is ERpos, tumorigenic, but noninvasive, primary human mammary epithelial cells (HMECs), immortalized nontumorigenic cell line MCF-10A (not shown), and three metastatic, ERneg cell lines, BT-549, MDA-MB-231 and MDA-MB-435 (representative data are shown in Figure 8). CPT triggered rapid cell death in HMECs and microscopic examination of the dying HMECs revealed blebbing and apoptotic body formation characteristic of apoptotic cells (not shown). The rate of cell death was further enhanced in the presence of autophagy inhibitors, 3-MA (Figure 8) and bafilomycin A1 (not shown), indicating that CPT treatment activated both an apoptotic response as well as an adaptive, autophagic response in normal breast epithelial cells. The T47D cells were more resistant to CPT but, like MCF-7 cells, died an accelerated death when autophagy was inhibited during CPT exposure. The metastatic breast cancer cell lines BT-549, MDA-MB-231 and MDA-MB-435, on the other hand, were resistant to concentrations of CPT that killed the HMEC, MCF-7 and T47D lines, either in the presence or absence of autophagy inhibitors. The inability of invasive breast cancer cell lines to activate apoptosis in response to both DNA damage and autophagy inhibition suggests that the autophagic survival response may either be independent of apoptosis or irrelevant in cells that have acquired resistance to genotoxic stress stimuli. The delayed apoptosis in the noninvasive tumor cells could stem from a defective apoptotic pathway, contributed to by aberrant expression of an intermediate such as caspase-3, further aided by a ‘defective’ autophagic pathway, resulting, for instance, from constitutive expression of low levels of the Beclin 1 protein. In fact, constitutive Beclin 1 expression may be an early factor, though not the only factor, contributing to the maintenance of the malignant phenotype and helping to promote tumor progression (Supplementary Figure 2). Cells treated with either CPT or the TOR inhibitor rapamycin, and immunostained for lysosomal-associated membrane protein 1 (LAMP-1), a marker for lysosomes associated with intermediate and late autophagy,30 suggested that the autophagic response was intact in all three invasive cell lines (Figure 8b and c). These observations suggest that autophagy as an adaptive early response to genotoxic stress enhances survival by retarding the rate of apoptosis in primary or immortalized breast cells and noninvasive breast cancer cells.
CPT-treated primary and noninvasive, but not metastatic, breast cell lines show enhanced apoptosis in response to autophagy inhibition. (a) The breast cell lines HMEC, MCF-7, T47D, BT-549, MDA-MB-231 and MDA-MB-435, treated with DMSO or CPT for 24 h, were incubated in the presence or absence of 3-MA for 5 h before harvesting. Cell viability was determined by FACS analysis as described earlier. Results (mean±S.E.) represent the average of at least three data points. (b) Breast cancer cells mount autophagic responses to both CPT and rapamycin. Cells were grown to 30–50% confluence on chamber slides and treated with DMSO or 2 μM CPT for 24 h, or 20 nM rapamycin for 36 h, fixed, permeabilized and immunolabeled with polyclonal anti-LAMP-1 primary antibody, followed by Alexa 488-tagged secondary antibody. Fixed and labeled cells were mounted in Vectastain containing DAPI and visualized using a Bio-Rad MRC 1024 Multi-Photon Confocal microscope. Data were quantified (c) using the ‘Analyze particles’ command in Image J software. 231 – MDA-MB-231, 435 – MDA-MB-435
Discussion
We have investigated the mechanisms underlying the delayed death response of the MCF-7 breast cancer line to the drug, CPT. Consistent with published studies, induction of a number of BH3-only transcripts and proteins, including Puma and Noxa, was preceded by an increase in the expression of p53, and the induced BH3 proteins localized to mitochondria within 24 h of drug exposure. The BimEL isoform was converted to a shorter protein indicative of dephosphorylation, a modification previously shown to reduce its ability to bind Bax and prepare it for proteasomal degradation.31, 32 The presence of mitochondrially localized active Bax associated with p53, and finally cyt c release, all predicted an impending apoptotic caspase cascade. Despite early events indicative of a classical intrinsic apoptotic pathway, however, the proportion of Ann V-positive cells was low, death was largely caspase-independent and the characteristic DNA fragmentation observed in apoptotic cells was absent (not shown). Transmission electron microscopy and confocal microscopy revealed the presence of large vacuoles often colocalizing with mitochondria, suggesting the activation of autophagic mechanisms. Furthermore, exposure to autophagy inhibitors such as bafilomycin A1 and 3-MA caused a small, but statistically significant, enhancement of both apoptosis and caspase-9 activity in the CPT-treated cell population.
Although sustained autophagy contributes to cell death, this pathway may be an early attempt to establish homeostasis. Mitochondrial membrane permeabilization and cyt c release below the threshold required for apoptosis can promote sequestration and turnover of the damaged mitochondria in degradative vacuoles or autophagosomes. Such a ‘checkpoint’ may serve as an effective survival strategy for the cells, providing an immediate source of metabolic energy in the form of ATP from the damaged mitochondria. The overt PCD II response in MCF-7 cells may be due to lack of caspase-3 activity and possible defects in apoptosome formation/caspase-9 activation, although studies have shown that restoration of caspase-3 expression only partially restores apoptosis in these cells.11 Furthermore, breast cancers expressing normal caspase-3, such as T47D, also mount an autophagic response to a variety of therapeutics, including DNA-damaging agents (Figure 8 and Mooney et al.33).
We provide further evidence that inhibiting autophagy stimulates apoptotic pathways in DNA-damaged MCF-7 cells by eliminating the expression of Beclin 1 and Atg7 proteins. Atg7 is an E1-like enzyme involved in the ubiquitination of MAP-LC3 early in the pathway,21 whereas Beclin 1 controls autophagosome formation by entering into a complex with class III PI3 kinase.23, 34, 35 Pattingre et al.36 have demonstrated that the release of Beclin 1 from a complex with ER localized Bcl-2 underlies the nutrient starvation response in MCF-7 cells. Our studies show that Beclin 1 expression is also necessary for suppression of the apoptotic response to DNA damage in MCF-7 cells. Beclin 1 is monoallelically deleted in 40–75% of sporadic human breast cancers, and protein expression is consistently reduced in breast tumors that have lost one allele.34 Our studies show that further downregulation or total loss of Beclin 1 expression essentially abrogates the autophagic ‘survival’ response to DNA damage, allowing cells to die by apoptosis. Similarly, knockdown of Atg7 increased apoptosis in response to CPT, although the response was weaker than that observed in Beclin 1low cells.
A number of treatments used against breast and ovarian cancers, including tamoxifen, rapamycin and radiation, have been shown to activate autophagy at low doses.18, 37, 38 This study is the first demonstration that DNA-damaging chemotherapeutic drugs, CPT and etoposide, activate an autophagic stress response in the MCF-7 breast cancer cells. This work also shows that knock down of proteins, such as Beclin 1, critical for autophagosome formation, can expose a latent apoptotic pathway in these cells. Studies showing that stable overexpression of Beclin 1 reduces the malignant phenotype and ability for anchorage-independent growth in MCF-7 cells,23 have led to the suggestion that autophagy is a tumor-suppressing event and Beclin 1, a tumor suppressor. Although a fully activated autophagic response may be tumor suppressing, we would argue that autophagy resulting from low, but constitutive, Beclin 1 protein expression serves a survival function by blocking the onset of an apoptotic cascade during periods of stress. Implications of monoallelic deletion of the beclin 1 gene notwithstanding, it is highly significant that the protein shows constitutive steady-state expression in all the breast cancer cell lines examined, but not in primary HMECs (Supplementary Figure 2) or in immortalized MCF-10A cells (not shown). Basal expression of Beclin 1 may allow for a faster autophagic response to stress stimuli, further contributing to the suppression of apoptosis. This may be particularly evident if cells harbor defective apoptotic machinery that results in a muted apoptotic response, as in the case of the MCF-7 cells.
A number of provocative questions are raised by this study. For instance, is there a common critical initiating event in the DNA damage response for both PCD I and II in breast cancer cells, or are the two pathways mutually exclusive? What is the trigger for autophagy initiation in invasive breast cancer cells, particularly those with high basal levels of autophagy, such as the MDA-MB-231 cells (Figure 8b and c)? Our observations indicate that autophagy inhibition does not activate latent apoptotic pathways in the three invasive cell lines examined, and it is tempting to suggest that (DNA damaging) stimuli, which activate autophagy as an adaptive response in breast cancers cells, eventually select for cells with a survival advantage thus promoting tumor progression. The selection process could result from the accumulation of compensatory DNA mutations by the exposed cell as it survives by ‘self-eating’ in the face of damaged or poorly repaired DNA. Studies aimed at determining whether controlled exposure of breast epithelial and noninvasive breast cancer cells to DNA damage promotes tumor progression and drug-resistance via activation or maintenance of an increasingly overt autophagic phase are currently underway.
Materials and Methods
Cell lines, chemicals and antibodies
MCF-7 breast cancer cells were grown in minimal essential medium (MEM) with 10% fetal calf serum (FCS), 2 mM L-glutamine, non-essential amino acids, 1 mM sodium pyruvate, 100 units/ml penicillin and 100 μg/ml streptomycin, BT549 cells were grown in RPMI 1640 with 10% FCS, 2 mM L-glutamine, 0.15 μg/ml insulin, 100 units/ml penicillin and 100 μg/ml streptomycin. T47D cells were cultured in MEM with 5% FCS, non-essential amino acids, 2 mM L-glutamine, 0.15 μg/ml human insulin, 100 units/ml penicillin and 100 μg/ml streptomycin. MDA-MB-231 and MA-MB-435 lines were maintained in DMEM with 10% FCS, 11.25 nM human insulin, 100 units/ml penicillin and 100 μg/ml streptomycin. Primary HMECs were cultured in mammary epithelial basal medium(Clonetics), supplemented with growth factors, cytokines and supplements according to the manufacturer's instructions. The cells were maintained in a humidified atmosphere of 5% CO2 at 37°C.
Bafilomycin A1, 3-methyl adenine and rapamycin were purchased from Sigma (St. Louis, MO, USA). MitoTracker Red, MitoTracker Green, TMRE and MDC were obtained from Molecular Probes, Inc. (Eugene, OR, USA). CPT was from BioVision Inc. (Palo Alto, CA, USA) and wortmannin was from EMD Biosciences (La Jolla, CA, USA). Antibodies were obtained or purchased from the following sources/suppliers: mouse monoclonal anti-actin antibody from Oncogene (San Diego, CA, USA); rabbit polyclonal anti-human Bak, NT from Upstate Biotechnology (Lake Placid, NY, USA); rabbit polyclonal anti-Bax (N-20), rabbit polyclonal p53 (Pab 1801) and mouse monoclonal anti-p53 (DO-1) antibodies from Santa Cruz Biotechnology (Santa Cruz, CA, USA); mouse monoclonal anti-Bax, Bcl-2 and rabbit polyclonal anti-Beclin 1 from BD Pharmingen (San Diego, CA, USA); rabbit polyclonal anti-human Bid from Cell Signaling (Beverly, MA, USA); rabbit polyclonal anti-human Bim EL/L from Stressgen (British Columbia, Canada); cytochrome oxidase subunit IV (COX IV) mouse monoclonal antibody from Molecular Probes (Eugene, OR, USA); rabbit polyclonal anti-human PUMA from Abcam (Cambridge, MA, USA); mouse monoclonal anti human-Noxa from IMGENEX (San Diego, CA, USA); rabbit polyclonal Atg7 antibody from ProSci Inc. (Poway, CA, USA) and anti-LAMP-1 antibody from Affinity BioReagents (Golden, CO, USA). The monoclonal anti-cyt c antibody was a generous gift from Ron Jemmerson (University of Minnesota) and rabbit polyclonal anti-Bcl-xL (13.6) has been described previously.39 The GFP-LC3 plasmid was a kind gift from Drs. T Yoshimori (National Institute of Genetics, Shizuoka-ken, Japan) and N Mizushima (The Tokyo Metropolitan Institute of Medical Science, Japan).
Induction of apoptosis and apoptosis assays
MCF-7 cells were allowed to reach ⩽80% confluency in fresh medium before treatment. Exponentially growing cells were exposed to 2 μM CPT for specific periods of time. For the apoptosis assays, which were carried out in triplicate, 0.5–1 × 106 cells from exponentially growing and CPT-treated cultures were collected at the times indicated. Inhibitors of autophagy, such as wortmannin (10 μM), bafilomycin A1 (200 nM) and 3-methyladenine (10 mM) were added to the culture medium 4–6 h before harvesting. Adherent MCF-7 cells were detached from culture plates with Accutase (Innovative Cell Technologies, San Diego, CA, USA) or trypsin-ethylene diaminetetra acetic acid (EDTA) and pelleted with predetached floaters. All cell cultures were washed in cold phosphate-buffered saline (PBS) and resuspended in Ann V binding buffer. Cell survival was measured by flow cytometric analysis of cells stained with fluorescein isothiocyanate-labeled Ann V and PI and analyzed using CellQuest Pro Software (BD Biosciences, San Jose, CA, USA). Ann V single positive, PI single positive and double-positive populations were collectively counted as ‘dead’, whereas double-negative cells were considered viable. To label mitochondria, MCF-7 cells were incubated with 40 nM TMRE or 75 nM MitoTracker Red at 37°C and 5% CO2 in culture medium for 15 min before harvesting for flow cytometry, or fixing in 3.7% paraformaldehyde for confocal microscopy, respectively.
In vitro transcription and multiprobe ribonuclease protection assays
Total RNA was extracted from untreated cells or cells treated with CPT at the indicated periods of time using the RNeasy Midi kit (Qiagen, Valencia, CA, USA). Two BH3-only multiple RPA template sets generated in our laboratory were used in the protection assays.12 Probe set 1 included Puma, Bim, BNip3, Bak, Bik, Mcl-1, Bid and housekeeping gene GAPDH as a loading control, and probe set 2 included Bax, Bmf, MAP-1, Rad9, Bcl-x, Bad, Bcl-G, Hrk and GAPDH. Multiple RPA was performed with the HybSpeed™ RPA kit according to the protocol previously outlined12 using [32P]-UTP labeled antisense RNA probes and 10 μg total cell RNA. Protected fragments in autoradiographs of dried exposed gels were quantified by densitometry using BioRad GS-363 Molecular Imager and Molecular Analyst software. Optical density values of BH3-only RNA bands were normalized relative to GAPDH bands and RNA levels were expressed as arbitrary units (mean and standard deviations, n=3). Statistical comparisons of mean values were carried out using Student's t-test; P-values ⩽0.05 were considered to be statistically significant.
Preparation of whole-cell lysates and mitochondrial fractions
After treatment with 2 μM CPT for 0, 3, 6, 9, 12 and 24 h, MCF-7 cells were pelleted by centrifugation and washed once with PBS. Cells were then resuspended in RIPA buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1% v/v Nonidet P-40, 0.5% v/v sodium deoxycholate and 0.1% SDS) supplemented with protease inhibitor cocktail (Calbiochem) and lysed on ice for 5 min with intermittent vortexing. The lysates were centrifuged for 5 min at 10 600 × g and supernatant was collected and used for further experiments. Mitochondrial fractions were enriched from untreated exponentially growing MCF-7 cells or cells treated with CPT for 24 h. Cells were dislodged with Accutase, washed in cold PBS and resuspended in 1.0–1.5 ml of ice-cold mitochondria isolation buffer (250 mM sucrose, 10 mM Tris-HCl, pH 7.4 and 0.1 mM ethylene glycol-bis (b-aminoethyl ether)) containing a protease inhibitor cocktail. Mitochondria were isolated either using the Qproteome™ Mitochondria Isolation kit (Qiagen) or by a nitrogen cavitation technique. For the latter, cells were lysed by nitrogen cavitation (650 psi, 10 min), lysates were collected and centrifuged at 2500 r.p.m. for 3 min. Supernatants were recentrifuged at 10 000 × g for 10 min and pellets, representing the mitochondrial fractions, were resuspended in isolation buffer. Mitochondrial integrity was confirmed by Western blotting using COX IV antibody.
Western blotting and immunoprecipitation
For Western blotting, 1 × 107 cells were lysed in 150 μl of RIPA buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.5% v/v sodium deoxycholate, 1% v/v Nonidet P-40, 0.1% SDS) supplemented with protease inhibitor cocktail (Calbiochem). Lysates (50–60 μg total protein) were electrophoretically separated by 14 or 18% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to nitrocellulose membrane. Membrane blocking, washing, primary and secondary antibody incubations and chemiluminescence reactions were carried out according to the Amersham ECL protocol. Antibody dilutions were carried out as per the data sheet provided by the manufacturer or as suggested by the provider's laboratory. Blots were stripped for reuse by washing for 30 min to 2 h in TBS-T buffer (pH 2.5–3.0) at room temperature. For immunoprecipitation, cell extracts were prepared in lysis buffer (100 mM NaCl, 1 mM EDTA, 20 mM Tris (pH 8.0), 0.1% IGEPAL) supplemented with a protease inhibitor cocktail. Cell lysate (100 μg) was incubated with 1–5 μg/ml of antibody and immune complexes were captured with protein G Sepharose. Proteins were resolved by SDS-PAGE, transferred to nitrocellulose membranes and antigen–antibody complexes were detected using ECL (Amersham Biosciences, Piscataway, NJ, USA). The endogenous Beclin 1 protein was detected using the Super Signal West Femto kit (Pierce Biotech) as per the manufacturer's instructions.
Active Bax was detected by immunoprecipitation in CHAPS lysis buffer, with an anti-Bax monoclonal antibody, 6A7 (BD Pharmingen), that recognizes Bax only in the active conformation.40 Mitochondrially enriched fractions were prepared from 24 h CPT-treated and control MCF-7 cells lysed in CHAPS lysis buffer (2% CHAPS, 150 mM NaCl, 1 mM N-2-hydroxyl piperazine-N′-2-ehane sulfonic acid, pH 7.5) containing protease inhibitors. For immunoprecipitation, 400 μg of mitochondrial lysate was incubated with anti-Bax 6A7 monoclonal antibody, complexes were immobilized with protein-G agarose analyzed by SDS-PAGE/immunoblot with anti-Bax polyclonal antibody (Santa Cruz Biotech).
Caspase activity assays
Cells were collected following treatment with 2 μM CPT for the time indicated. Cells were centrifuged to pellet and washed once in 1 × PBS. The cell pellet was then flash frozen in a dry ice/ethanol bath. After collection of all time points, cell pellets were lysed on ice for 10 min with intermittent vortexing in 160 μl lysis buffer (R&D Systems, Minneapolis, MN, USA). Lysates were centrifuged for 1 min at 10 600 × g and a small volume of supernatant was set aside for total protein estimation. The remainder was tested for caspase activity in triplicate using a colorimetric assay kit from R&D Systems. Caspase-9 activity was determined using LEHD-pNA as substrate. Reactions were allowed to develop for 2 h at 37°C and absorbance measured at 405 nm using a microplate reader.
Transmission electron microscopy
Cells were fixed in 2.5% glutaraldehyde in 0.1 M sodium cacodylate buffer (pH 7.4) for 1 h at room temperature, then washed 3 times in cacodylate buffer. Cells were postfixed in 1% osmium tetroxide in 0.1 M cacodylate buffer for 1 h at room temperature. Following this, 70 nm sections were cut on a ‘Reichert Ultracut S’ ultramicrotome. The sections were subsequently post-stained with 4% uranyl acetate for 10 min and Reynald's lead citrate for 1.5 min. Sections were imaged at 80 kV on a JEOL 1200 EX transmission electron microscope.
Confocal and fluorescence microscopy
Cells were plated at 50% confluence on glass chamber slides and treated with 2 μM CPT or DMSO for 24 h and observed using a confocal microscope. Cells were incubated with 100 μM MDC and 20 nM MitoTracker Green for 15–30 min, washed thoroughly in PBS, fixed for 30 min at 37°C in 3.7% paraformaldehyde in PBS, rinsed, dried and mounted under coverslips using Vectashield mounting medium containing DAPI (Vector Laboratories, Inc.). Cells were observed at 60 × magnification using a Bio-Rad MRC 1024 Single or Multi-Photon Confocal microscope. Alternatively, GFP-LC3-expressing MCF-7 cells treated with CPT for 18 h were labeled with Mitotracker Red and observed with the Single Photon Confocal microscope. Images were captured using the Bio-Rad Laser Sharp 3.0 and processed with Image J or Photoshop CS software. GFP-LC3 translocation was quantified using the Image J software command ‘Analyze Particles’, which counts and measures objects in binary or thresholded images.
Beclin 1 and Atg7 sh RNA constructs, transfection and expression
The human Beclin 1 shRNA sequence (CAGTTTGGCACAATCAATA) or human Atg7 sequence (GGAGTCACAGCTCTTCCTT) was cloned into BamH1 and EcoR1 sites of the pGSU6 vector (Gene Therapy Systems). Control scrambled and shRNA plasmids (1.5 μg) were transfected into MCF-7 cells plated at 50% confluence in six-well plates using lipofectamine (Invitrogen, Carlsbad, CA, USA). Transfectants were selected in culture medium containing 1.42 mg/ml Geneticin (Cellgro, Herndon, VA, USA) and single-cell clones were isolated using cloning cylinders (Fisher Biotech).
Abbreviations
- CPT:
-
camptothecin
- cyt c:
-
cytochrome c
- Ann V:
-
Annexin V
- PI:
-
propidium iodide
- TMRE:
-
tetramethylrhodamine ethyl ester, perchlorate
- DAPI:
-
4′-6-diamidino-2-phenylindole
- MAP-LC3:
-
microtubule-associated protein 1, light chain 3
- MDC:
-
monodansyl cadaverine
- 3-MA:
-
3-methyl adenine
- LAMP-1:
-
lysosomal-associated membrane protein 1
References
Kaufmann SH, Hengartner MO . Programmed cell death: alive and well in the new millennium. Trends Cell Biol 2001; 11: 526–534.
Budihardjo I, Oliver H, Lutter M, Luo X, Wang X . Biochemical pathways of caspase activation during apoptosis. Annu Rev Cell Dev Biol 1999; 15: 269–290.
Gozuacik D, Kimchi A . Autophagy as a cell death and tumor suppressor mechanism. Oncogene 2004; 23: 2891–2906.
Wang CW, Klionsky DJ . The molecular mechanism of autophagy. Mol Med 2003; 9: 65–76.
Adams JM, Cory S . The Bcl-2 protein family: arbiters of cell survival. Science 1998; 281: 1322–1326.
Huang DC, Strasser A . BH3-Only proteins-essential initiators of apoptotic cell death. Cell 2000; 103: 839–842.
Kelekar A, Thompson CB . Bcl-2-family proteins: the role of the BH3 domain in apoptosis. Trends Cell Biol 1998; 8: 324–330.
Chipuk JE, Green DR . Cytoplasmic p53: bax and forward. Cell Cycle 2004; 3: 429–431.
Schuler M, Green DR . Transcription, apoptosis and p53: catch-22. Trends Genet 2005; 21: 182–187.
Ferguson HA, Marietta PM, Van Den Berg CL . UV-induced apoptosis is mediated independent of caspase-9 in MCF-7 cells: a model for cytochrome c resistance. J Biol Chem 2003; 278: 45793–45800.
Kottke TJ, Blajeski AL, Meng XW, Svingen PA, Ruchaud S, Mesner Jr PW et al. Lack of correlation between caspase activation and caspase activity assays in paclitaxel-treated MCF-7 breast cancer cells. J Biol Chem 2002; 277: 804–815.
Wang D, McDonnell MA, Kelekar A . Multi-probe RPA template sets to study RNA modulation and transcriptional control of BH3-only members of the Bcl-2 family. Cancer Detect Prev 2005; 29: 189–200.
Ley R, Balmanno K, Hadfield K, Weston C, Cook SJ . Activation of the ERK1/2 signaling pathway promotes phosphorylation and proteasome-dependent degradation of the BH3-only protein, Bim. J Biol Chem 2003; 278: 18811–18816.
Chipuk JE, Kuwana T, Bouchier-Hayes L, Droin NM, Newmeyer DD, Schuler M et al. Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science 2004; 303: 1010–1014.
Mihara M, Erster S, Zaika A, Petrenko O, Chittenden T, Pancoska P et al. p53 has a direct apoptogenic role at the mitochondria. Mol Cell 2003; 11: 577–590.
Elmore SP, Qian T, Grissom SF, Lemasters JJ . The mitochondrial permeability transition initiates autophagy in rat hepatocytes. FASEB J 2001; 15: 2286–2287.
Elmore SP, Nishimura Y, Qian T, Herman B, Lemasters JJ . Discrimination of depolarized from polarized mitochondria by confocal fluorescence resonance energy transfer. Arch Biochem Biophys 2004; 422: 145–152.
Paglin S, Hollister T, Delohery T, Hackett N, McMahill M, Sphicas E et al. A novel response of cancer cells to radiation involves autophagy and formation of acidic vesicles. Cancer Res 2001; 61: 439–444.
Scarlatti F, Bauvy C, Ventruti A, Sala G, Cluzeaud F, Vandewalle A et al. Ceramide-mediated macroautophagy involves inhibition of protein kinase B and up-regulation of beclin 1. J Biol Chem 2004; 279: 18384–18391.
Biederbick A, Kern HF, Elsasser HP . Monodansylcadaverine (MDC) is a specific in vivo marker for autophagic vacuoles. Eur J Cell Biol 1995; 66: 3–14.
Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J 2000; 19: 5720–5728.
Eskelinen EL, Prescott AR, Cooper J, Brachmann SM, Wang L, Tang X et al. Inhibition of autophagy in mitotic animal cells. Traffic 2002; 3: 878–893.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Liang XH, Kleeman LK, Jiang HH, Gordon G, Goldman JE, Berry G et al. Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J Virol 1998; 72: 8586–8596.
Liang XH, Yu J, Brown K, Levine B . Beclin 1 contains a leucine-rich nuclear export signal that is required for its autophagy and tumor suppressor function. Cancer Res 2001; 61: 3443–3449.
Hande KR . Etoposide: four decades of development of a topoisomerase II inhibitor. Eur J Cancer 1998; 34: 1514–1521.
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol 2005; 169: 425–434.
Tanida I, Mizushima N, Kiyooka M, Ohsumi M, Ueno T, Ohsumi Y et al. Apg7p/Cvt2p: A novel protein-activating enzyme essential for autophagy. Mol Biol Cell 1999; 10: 1367–1379.
Rizk NN, El-Rakhawy MT . Tissue culture and scanning electron microscopy of breast carcinoma ‘cell line MCF-7’ from pleural effusion. Acta Anat (Basel) 1981; 109: 70–74.
Mizushima N . Methods for monitoring autophagy. Int J Biochem Cell Biol 2004; 36: 2491–2502.
Harada H, Quearry B, Ruiz-Vela A, Korsmeyer SJ . Survival factor-induced extracellular signal-regulated kinase phosphorylates BIM, inhibiting its association with BAX and proapoptotic activity. Proc Natl Acad Sci USA 2004; 101: 15313–15317.
Seward RJ, von Haller PD, Aebersold R, Huber BT . Phosphorylation of the pro-apoptotic protein Bim in lymphocytes is associated with protection from apoptosis. Mol Immunol 2003; 39: 983–993.
Mooney LM, Al-Sakkaf KA, Brown BL, Dobson PR . Apoptotic mechanisms in T47D and MCF-7 human breast cancer cells. Br J Cancer 2002; 87: 909–917.
Aita VM, Liang XH, Murty VV, Pincus DL, Yu W, Cayanis E et al. Cloning and genomic organization of beclin 1, a candidate tumor suppressor gene on chromosome 17q21. Genomics 1999; 59: 59–65.
Kihara A, Kabeya Y, Ohsumi Y, Yoshimori T . Beclin-phosphatidylinositol 3-kinase complex functions at the trans-Golgi network. EMBO Rep 2001; 2: 330–335.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Bursch W, Ellinger A, Kienzl H, Torok L, Pandey S, Sikorska M et al. Active cell death induced by the anti-estrogens tamoxifen and ICI 164 384 in human mammary carcinoma cells (MCF-7) in culture: the role of autophagy. Carcinogenesis 1996; 17: 1595–1607.
Ogier-Denis E, Codogno P . Autophagy: a barrier or an adaptive response to cancer. Biochim Biophys Acta 2003; 1603: 113–128.
Boise LH, Gonzalez-Garcia M, Postema CE, Ding L, Lindsten T, Turka LA et al. bcl-x, a bcl-2-related gene that functions as a dominant regulator of apoptotic cell death. Cell 1993; 74: 597–608.
Nechushtan A, Smith CL, Hsu YT, Youle RJ . Conformation of the Bax C-terminus regulates subcellular location and cell death. EMBO J 1999; 18: 2330–2341.
Acknowledgements
We thank Drs. T Yoshimori and N Mizushima for sharing the GFP-LC3 plasmid, Drs. Doug Yee, Carol Lange and Vitaly Polunovsky for sharing breast cell lines, Dr. Robert Hafner for help with the transmission electron microscopy, and Jerry Sedgewick and John Oja for help with the confocal microscopy. We also thank Dr. Thomas Neufeld for helpful discussions. This work was supported by research grants from the Minnesota Medical Foundation and from The University of Minnesota Cancer Center, a Comprehensive Cancer Center designated by the National Cancer Institute and the Regis Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by E Baehrecke
Supplementary Information accompanies the paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Supplementary information
Rights and permissions
About this article
Cite this article
Abedin, M., Wang, D., McDonnell, M. et al. Autophagy delays apoptotic death in breast cancer cells following DNA damage. Cell Death Differ 14, 500–510 (2007). https://doi.org/10.1038/sj.cdd.4402039
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4402039
Keywords
This article is cited by
-
Gaillardin inhibits autophagy and induces apoptosis in MCF-7 breast cancer cells by regulating JAK/STAT pathway
Molecular Biology Reports (2024)
-
Apoptosis and autophagy markers predict survival in neoadjuvant treated oesophageal adenocarcinoma patients
BMC Cancer (2022)
-
Endonuclease G promotes autophagy by suppressing mTOR signaling and activating the DNA damage response
Nature Communications (2021)
-
Intracellular leucine-rich alpha-2-glycoprotein-1 competes with Apaf-1 for binding cytochrome c in protecting MCF-7 breast cancer cells from apoptosis
Apoptosis (2021)
-
Inhibition of autophagy by chloroquine enhances the antitumor activity of gemcitabine for gallbladder cancer
Cancer Chemotherapy and Pharmacology (2020)