Abstract
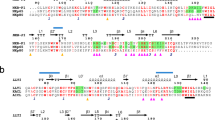
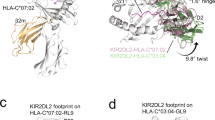
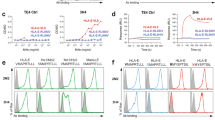
Natural killer (NK) cell function is regulated by NK receptors that interact with MHC class I (MHC-I) molecules on target cells. The murine NK receptor Ly49A inhibits NK cell activity by interacting with H-2Dd through its C-type-lectin-like NK receptor domain. Here we report the crystal structure of the complex between the Ly49A NK receptor domain and unglycosylated H-2Dd. The Ly49A dimer interacts extensively with two H-2Dd molecules at distinct sites. At one interface, a single Ly49A subunit contacts one side of the MHC-I peptide-binding platform, presenting an open cavity towards the conserved glycosylation site on the H-2Dd α2 domain. At a second, larger interface, the Ly49A dimer binds in a region overlapping the CD8-binding site. The smaller interface probably represents the interaction between Ly49A on the NK cell and MHC-I on the target cell, whereas the larger one suggests an interaction between Ly49A and MHC-I on the NK cell itself. Both Ly49A binding sites on MHC-I are spatially distinct from that of the T-cell receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lanier,L. L. NK cell receptors. Annu. Rev. Immunol. 16, 359–393 (1998).
Yokoyama,W. M. in Fundamental Immunology (ed. Paul, W. E.) 575–603 (Lippincott-Raven, Philadelphia, 1999).
Long,E. O. Regulation of immune responses through inhibitory receptors. Annu. Rev. Immunol. 17, 875–904 (1999).
Kärre,K., Ljunggren,H. G., Piontek,G. & Kiessling,R. Selective rejection of H-2-deficient lymphoma variants suggests alternative immune defense strategy. Nature 319, 675–678 (1986).
Ljunggren,H. G. & Kärre,K. In search of the “missing self”: MHC molecules and NK cell recognition. Immunol. Today 11, 237–244 (1990).
Fan,Q. R. et al. Structure of the inhibitory receptor for human natural killer cells resembles haematopoietic receptors. Nature 389, 96–100 (1997).
Snyder,G. A., Brooks,A. G. & Sun,P. D. Crystal structure of the HLA-Cw3 allotype-specific killer cell inhibitory receptor KIR2DL2. Proc. Natl Acad. Sci. USA 96, 3864–3869 (1999).
Maenaka,K., Juji,T., Stuart,D. I. & Jones,E. Y. Crystal structure of the human p58 killer cell inhibitory receptor (KIR2DL3) specific for HLA-Cw3-related MHC class I. Structure 7, 391–398 (1999).
Boyington,J. C. et al. Structure of CD94 reveals a novel C-type lectin fold: implications for the NK cell-associated CD94/NKG2 receptors. Immunity 10, 75–82 (1999).
Garcia,K. C., Teyton,L. & Wilson,I. A. Structural basis of T cell recognition. Annu. Rev. Immunol. 17, 369–397 (1999).
Natarajan,K. et al. Interaction of the NK cell inhibitory receptor Ly49A with H-2Dd: identification of a site distinct from the TCR site. Immunity 11, 591–601 (1999).
Li,H., Natarajan,K., Malchiodi,E. L., Margulies,D. H. & Mariuzza,R. A. Three-dimensional structure of H-2Dd complexed with an immunodominant peptide from human immunodeficiency virus envelope glycoprotein 120. J. Mol. Biol. 283, 179–191 (1998).
Weis,W. I., Taylor,M. E. & Drickamer,K. The C-type lectin superfamily in the immune system. Immunol. Rev. 163, 19–34 (1998).
Weis,W. I., Kahn,R., Fourme,R., Drickamer,K. & Hendrickson,W. A. Structure of the calcium-dependent lectin domain from a rat mannose-binding protein determined by MAD phasing. Science 254, 1608–1615 (1991).
Poget,S. F. et al. The structure of a tunicate C-type lectin from Polyandrocarpa misakiensis complexed with D-galactose. J. Mol. Biol. 290, 867–879 (1999).
Ng,K. K.-S., Park-Snyder,S. & Weis,W. I. Ca2+-dependent structural changes in C-type mannose-binding proteins. Biochemistry 37, 17965–17976 (1998).
Weis,W. I. & Drickamer,K. Trimeric structure of a C-type mannose-binding protein. Structure 2, 1227–1240 (1994).
Sheriff,S., Chan,C. Y. & Ezekowitz,R. A. B. Human mannose-binding protein carbohydrate recognition domain trimerizes through a triple α-helical coiled-coil. Nature Struct. Biol. 1, 789–794 (1994).
Graves,B. J. et al. Insight into E-selectin/ligand interaction from the crystal structure and mutagenesis of the lec/EGF domains. Nature 367, 532–538 (1994).
Lo Conte,L., Chothia,C. & Janin,J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
Wang,J.-H. et al. Structure of a heterophilic adhesion complex between the human CD2 and CD58 (LFA-3) counterreceptors. Cell 97, 791–803 (1999).
Lawrence,M. C. & Colman,P. M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993).
Karlhofer,F. M., Ribaudo,R. K. & Yokoyama,W. M. MHC class I alloantigen specificity of Ly-49+ IL-2 activated natural killer cells. Nature 358, 66–70 (1992).
Gao,G. F. et al. Crystal structure of the complex between human CD8αα and HLA-A2. Nature 387, 630–634 (1997).
Kern,P. S. et al. Structural basis of CD8 coreceptor function revealed by crystallographic analysis of a murine CD8αα ectodomain fragment in complex with H-2Kb. Immunity 9, 519–530 (1998).
Achour,A. et al. The crystal structure of H-2Dd MHC class I complexed with the HIV-1-derived peptide P18-I10 at 2.4 Å resolution: implications for T cell and NK cell recognition. Immunity 9, 199–208 (1998).
Correa,I. & Raulet,D. H. Binding of diverse peptides to MHC class I molecules inhibits target cell lysis by activated natural killer cells. Immunity 2, 61–71 (1995).
Orihuela,M., Margulies,D. H. & Yokoyam,W. M. The natural killer cell receptor Ly49A recognizes a peptide-induced conformational determinant on its major histocompatibility complex class I ligand. Proc. Natl Acad. Sci. USA 93, 11792–11797 (1996).
Matsumoto,N., Ribaudo,R. K., Abastado,J. P., Margulies,D. H. & Yokoyama,W. M. The lectin-like NK cell receptor Ly-49A recognizes a carbohydrate-independent epitope on its MHC class I ligand. Immunity 8, 245–254 (1998).
Sundbäck,J. et al. The α2 domain of H-2Dd restricts the allelic specificity of the murine NK cell inhibitory receptor Ly-49A. J. Immunol. 160, 5971–5978 (1998).
Karlhofer,F. M., Ribaudo,R. K. & Yokoyama,W. M. MHC class I alloantigen specificity of Ly-49+ IL-2 activated natural killer cells. Nature 358, 66–70 (1992).
Brennan,J., Mahon,G., Mager,D. L., Jefferies,W. A. & Takei,F. Recognition of class I major histocompatibility complex molecules by Ly-49: specificities and domain interactions. J. Exp. Med. 183, 1553–1559 (1996).
Daniels,B. F., Nakamura,M. C., Rosen,S. D., Yokoyama,W. M. & Seaman,W. E. Ly49A, a receptor for H-2Dd, has a functional carbohydrate recognition domain. Immunity 1, 785–792 (1994).
Brennan,J., Takei,F., Wong,S. & Mager,D. L. Carbohydrate recognition by a natural killer cell receptor, Ly-49C. J. Biol. Chem. 270, 9691–9694 (1995).
Chang,C. S. & Kane,K. P. Evidence for sulphate modification of H-2Dd on N-linked carbohydrate(s): possible involvement in Ly-49A interaction. J. Immunol. 160, 4367–4374 (1998).
Lian,R. H., Freeman,J. D., Mager,D. L. & Takei,F. Role of conserved glycosylation site unique to murine class I MHC in recognition by Ly-49 NK cell receptor. J. Immunol. 161, 2031–2036 (1998).
Hanke,T. et al. Direct assessment of MHC class I binding by seven Ly49 inhibitory NK cell receptors. Immunity 11, 67–77 (1999).
Madden,D. R. The three-dimensional structure of peptide-MHC complexes. Annu. Rev. Immunol. 13, 587–622 (1995).
Kåse,A., Johansson,M. H., Olsson-Alheim,M. Y., Kärre,K. & Höglund,P. External and internal calibration of the MHC class I-specific receptor Ly49A on murine natural killer cells. J. Immunol. 161, 6133–6138 (1998).
Andersson,M. et al. MHC class I mosaic mice reveal insights into control of Ly49C inhibitory receptor expression in NK cells. J. Immunol. 161, 6475–6479 (1998).
Franksson,L. et al. Peptide dependency and selectivity of the NK cell inhibitory receptor Ly-49C. Eur. J. Immunol. 29, 2748–2758 (1999).
Valés-Gómez,M., Reyburn,H. T., Erskine,R. A., López-Botet,M. & Strominger,J. L. Kinetics and peptide dependency on the binding of the inhibitory NK receptor CD94/NKG2-A and the activating receptor CD94/NKG2-C to HLA-E. EMBO J. 18, 4250–4260 (1999).
Chung,D. H. et al. NK and CTL recognition of a single chain H-2Dd molecule: distinct sites of H-2Dd interact with NK and T cell receptors. J. Immunol. 163, 3699–3708 (1999).
Bendelac,A., Rivera,M. N., Park,S.-H. & Roark,J. H. Mouse CD1-specific NK1 T cells: development, specificity, and function. Annu. Rev. Immunol. 15, 535–562 (1997).
Otwinowski,Z. & Minor,W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–327 (1997).
Navaza,J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Brünger,A. T. et al. Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones,T. A., Zou,J. Y., Cowan,S. W. & Kjeldgaard,M. Improved methods for building models in electron density maps and the location of errors in those models. Acta Crystallogr. A 47, 110–119 (1991).
Esnouf,R. M. An extensively modified version of Molscript that includes greatly enhanced colouring capabilities. J. Mol. Graph. 15, 133–138 (1997).
Merrit,E. A. & Bacon,D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Acknowledgements
We thank W. Yang and B. A. Fields for their help with data collection, L. Boyd and R. Carey for assistance with protein expression and characterization, M. Garfield for protein sequencing, and J. C. Boyington and P. D. Sun for providing coordinates of the CD94 structure before release. We thank W. Yokoyama for encouragement and for critical comments on the manuscript. We also thank the staff at the Advanced Photon Source, Argonne National Laboratory, which is operated by the Department of Energy, Office of Basic Energy Sciences. This work was supported, in part, by grants from the National Institutes of Health and the National Multiple Sclerosis Society (R.A.M.) and the Spanish “Comision Interministerial de Ciencia y Tecnologia” (J.T.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Tormo, J., Natarajan, K., Margulies, D. et al. Crystal structure of a lectin-like natural killer cell receptor bound to its MHC class I ligand. Nature 402, 623–631 (1999). https://doi.org/10.1038/45170
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/45170
This article is cited by
-
Structure of MHC class I-like MILL2 reveals heparan-sulfate binding and interdomain flexibility
Nature Communications (2018)
-
Recognition of host Clr-b by the inhibitory NKR-P1B receptor provides a basis for missing-self recognition
Nature Communications (2018)
-
The Ly49 natural killer cell receptors: a versatile tool for viral self‐discrimination
Immunology & Cell Biology (2014)
-
The activating Ly49W and inhibitory Ly49G NK cell receptors display similar affinities for identical MHC class I ligands
Immunogenetics (2014)
-
Targeting of a natural killer cell receptor family by a viral immunoevasin
Nature Immunology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.