Abstract
The chaperonin GroEL has an essential role in mediating protein folding in the cytosol of Escherichia coli. Here we show that GroEL interacts strongly with a well-defined set of approximately 300 newly translated polypeptides, including essential components of the transcription/translation machinery and metabolic enzymes. About one third of these proteins are structurally unstable and repeatedly return to GroEL for conformational maintenance. GroEL substrates consist preferentially of two or more domains with αβ-folds, which contain α-helices and buried β-sheets with extensive hydrophobic surfaces. These proteins are expected to fold slowly and be prone to aggregation. The hydrophobic binding regions of GroEL may be well adapted to interact with the non-native states of αβ-domain proteins.
Similar content being viewed by others
Main
The three-dimensional fold of a protein is determined by the amino-acid sequence of the newly synthesized polypeptide chain. Although proteins can reach their folded states spontaneously, the efficiency of folding is often limited by the side reaction of aggregation. In the cell, misfolding and aggregation of proteins during their biogenesis and under conditions of cellular stress is prevented by molecular chaperones1,2,3.
The chaperonin GroEL, along with its cofactor GroES, is the only chaperone system in E. coli that is essential under all growth conditions4,5. GroEL is a homo-oligomer of 14 subunits, each of relative molecular mass 57,000 (Mr 57K), which are arranged into two heptameric rings, forming a cylindrical structure with two large cavities. Substrate protein, with hydrophobic amino-acid residues exposed, binds in the central cavity of the cylinder, engaging the hydrophobic surfaces exposed by the apical GroEL domains6,7. The ring-shaped cofactor GroES then binds to the apical domains of GroEL in an ATP-dependent reaction, resulting in the displacement of the substrate into an enclosed cavity. Proteins up to Mr ∼ 60K can fold in the GroEL–GroES cage in which aggregation is prevented. After ∼10 s of folding, when the GroEL-bound ATP has been hydrolysed to ADP, ATP binding to the opposite ring of GroEL results in the dissociation of GroES and folded protein from GroEL. Proteins that are strongly dependent on GroEL may require several rounds of interaction with GroEL to reach their native state3,8,9.
GroEL interacts in vitro with almost any non-native model protein7. However, in vivo GroEL is involved in the folding of only ∼10% of newly translated polypeptides10, indicating a preference for a subset of E. coli proteins. We have identified a large number of the endogenous substrates of GroEL, many of which are structurally labile and interact with GroEL not only for the initial folding, but also for conformational maintenance during their lifetime in the cell, both under normal growth conditions and when the cell is exposed to heat stress. Structural analysis revealed that these proteins have a complex domain architecture, preferentially containing two or more domains with αβ-folds.
A defined set of GroEL substrates
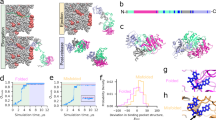
Pulse-chase labelling experiments were performed with live E. coli cells to analyse the set of newly synthesized proteins interacting with GroEL. At different times of chase, cells were lysed on ice in the presence of EDTA, which prevented the ATP-dependent release of protein substrates from GroEL, and GroEL–substrate complexes were isolated by immunoprecipitation with anti-GroEL antibodies10. Total soluble cytoplasmic proteins (Fig. 1a, b) and GroEL-bound proteins (Fig. 1c, d) were separated on two-dimensional polyacrylamide gels (2D gels). Control experiments demonstrated the specificity of the anti-GroEL immunoprecipitations10 (see below).
a–d, E. coli MG1655 cells grown at 30 °C to mid-log phase were pulsed for 15 s and chased for 0–10 min. Equal amounts of total soluble cytoplasmic proteins (a, b) or immunoprecipitated proteins (c, d) were separated on 2D gels. In c, d, circles refer to proteins whose properties of interaction with GroEL are discussed in Figs 2 and 3. e, f, pI (e) and Mr (f) distributions of total soluble cytoplasmic proteins in E. coli (grey bars; mean pI, 5.7; mean Mr, 37.5K) and of GroEL substrates (black bars; mean pI, 5.8; mean Mr, 45.0K) at 0 min of chase (a, c). Similar distributions were obtained at the other time points.
At a rate of translation of 10–20 amino acids per s (ref. 11), a 15-s pulse with [35S]-methionine allows for the labelling of an E. coli protein comprising about 150–300 amino acids. Proteins with Mr larger than ∼40K, including GroEL itself, complete synthesis during the chase period with unlabelled methionine (Fig. 1b). In contrast to the expected complexity of newly labelled cytoplasmic proteins resolved on the 2D gels (Fig. 1a, b), the pattern of GroEL-bound proteins (referred to as GroEL substrates) was much simpler and surprisingly well defined (Fig. 1c, d). A core of no more than 250–300 proteins out of a total of ∼2,500 cytoplasmic polypeptides were reproducibly observed in complex with GroEL immediately upon labelling (Fig. 1c), even after prolonged exposure. About half of these proteins were still detectable on GroEL after 10 min of chase (Fig. 1d), albeit in strongly reduced amounts relative to GroEL. Proteins that interact only very transiently with GroEL may have escaped detection in this analysis, but such proteins are not expected to be strongly dependent on the chaperonin for folding.
The pI distributions of total soluble cytoplasmic proteins and of GroEL substrates were found to be very similar (Fig. 1e). However, most GroEL substrates are larger than Mr ∼ 20K (Fig. 1f), and so are likely to have several domains12. The majority of substrates (79%) are smaller than Mr 60K (Fig. 1f).
Maintenance function of GroEL
A systematic analysis of the flux of newly labelled substrates through GroEL revealed three groups of proteins. About two-thirds of the proteins with Mr less than 60K (around 160 proteins) were released completely from GroEL during the chase with time constants between 20 s and 2 min (Fig. 2a), presumably reflecting the requirement of one to several rounds of GroEL binding and release to reach their native state. In contrast, for a core group of around 100 proteins of Mr < 60K, a fraction of the population of a particular protein (5–30% of the initial amount recovered) remained associated with GroEL after the chase (Fig. 2b). In addition, several proteins larger than 60K were also observed to bind GroEL, but these were inefficiently released from the chaperonin (Fig. 2c). These proteins exceed the size limitation of the GroEL–GroES cage and cannot be enclosed in the GroEL cavity by GroES13. However, these proteins contribute only 6% of the total mass of proteins interacting with GroEL, as judged by the analysis of Coomassie-stained 2D gels.
Kinetics of flux through GroEL of some newly translated proteins (indicated by circles in Fig. 1c, d) are shown. For a given kinetic trace, the maximum value obtained was arbitrarily set at 100. Solid lines are traces through the experimental points based on single- or double-exponential fits to the data. a, b, Examples of proteins of Mr < 60K that are either completely (a) or partly (b) released from GroEL, with time constants of decay between 20 s and 2 min. c, Examples of proteins of Mr > 60K that exhibit very slow decay kinetics (time constant >2 min) with inefficient release from GroEL. For clarity, the flux kinetics of only 10 representative proteins of Fig. 1c, d are shown.
The persistence on GroEL of a fraction of newly synthesized polypeptides indicates that there are pre-existing proteins in E. coli (as opposed to newly synthesized proteins) that may undergo repeated cycles of GroEL binding and release. Indeed, when cells were incubated with [35S]-methionine for 5 min or longer and then chased with unlabelled methionine, the amount and pattern of labelled polypeptides bound to GroEL was very similar, from 10 min of chase up to more than 2 hours, provided that the increase in cell mass during the chase time was corrected for (the doubling time of cells is 1 h) (Fig. 3a, left). These proteins were specifically bound to GroEL: they were not recovered with the chaperonin when the cell extract was incubated with SDS10, and they were released from GroEL when ATP and GroES were added (Fig. 3a, right). Rebinding to GroEL was not observed in the dilute cell lysates, confirming the earlier conclusion that the interactions occurred in the intact cells, rather than during lysis10. In contrast to the pre-existing GroEL substrates of Mr < 60K, only a fraction of the GroEL-bound proteins of Mr > 60K were released from the chaperonin when ATP and GroES were added to the lysate (Fig. 3a, right). These proteins may represent dead-end species.
a, Cells were treated as in Fig. 1a–d except that the chase was added after 5 min of labelling. Left, the pattern of GroEL-bound proteins separated on 16% SDS–PAGE isolated at the indicated long chase times. The same amount of radioactive GroEL was loaded in each lane. Right, anti-GroEL immunoprecipitations (IP) from cells incubated before the addition of antibodies, with either 1% SDS or with ATP regenerating system and GroES. b, GroEL–substrate complexes were isolated as in a, left, and separated on a 2D gel. c, d, Cells, at 30 °C, were labelled for 5 min, chased for a further 5 min, and converted into spheroplasts10. Spheroplasts were incubated in the presence of unlabelled methionine for 5 min at 30 °C (c) or 43 °C (d) and then lysed immediately by dilution into hypo-osmotic buffer. GroEL–substrate complexes were isolated by immunoprecipitation and separated on 2D gels. The 2D gels are shown at exposures with similar intensities for the GroEL spot. Circles refer to proteins numbered as in Fig. 1c, d.
The 2D gel pattern of pre-existing GroEL substrates (Fig. 3b) was very similar to that of newly translated proteins recovered in complex with GroEL after long chase times (Fig. 1d), indicating that the pre-existing proteins may also be using the chaperonin for initial folding. From the quantification of GroEL immunoprecipitates, about 1% of the cellular content of these proteins is associated with GroEL at any one time, assuming average abundance. Given a GroEL folding reaction of 10 s, the entire population of a pre-existing GroEL substrate will thus cycle on and off GroEL about four times per doubling time of the cells (1 h). The normal concentration of the GroEL oligomer in the cytosol is ∼3 µM14, but each ring of GroEL may be active in polypeptide binding15. At a total protein concentration of 200 g l-1 in the cytosol16 and an average Mr of cytosolic proteins of 33K, the set of pre-existing GroEL substrates would occupy about 30% of the total polypeptide-binding capacity of GroEL.
It seemed plausible that the pre-existing proteins interacting with GroEL are relatively unstable, populating partly folded states to a significant extent in vivo. This idea could be confirmed by analysing the interaction of pre-existing proteins with GroEL under conditions of heat stress (Fig. 3c, d). For this experiment, spheroplasts were used rather than cells, as they can be lysed rapidly in a hypo-osmotic buffer. It was found that, under heat stress at 43 °C, predominantly the same set of pre-existing proteins interacted more extensively with GroEL (compare Fig. 3c and d), although GroEL was about threefold more abundant at 43 °C than at 30 °C. The total amount of pre-existing substrates on GroEL increased roughly 5-fold, varying between 3- and 12-fold for individual polypeptides (data not shown). Thus, under heat stress, the fraction of GroEL devoted to conformational maintenance of proteins approximately doubles.
GroEL substrates
To identify the proteins that interact with GroEL for initial folding and conformational maintenance, large-scale immunoprecipitations of GroEL–substrate complexes were performed from E. coli cells under the conditions described above. Isolated complexes were separated on 2D gels (Fig. 4) and protein spots were analysed by mass spectroscopy. This procedure identified unequivocally 52 of the most abundant GroEL-bound proteins (Table 1), including several components of the transcription/translation machinery, such as subunits of the RNA polymerase, elongation factor Tu, and several aminoacyl transfer RNA synthetases, as well as a variety of important metabolic enzymes. We verified that these proteins transit GroEL on synthesis by 2D-gel analysis of a large-scale immunoprecipitation of GroEL complexes that had been mixed with an immunoprecipitate from pulse-labelled cells prepared as in Fig. 1c (data not shown). The extent to which these proteins are dependent on GroEL and GroES for their folding remains to be determined. A preliminary analysis of the flux kinetics through GroEL indicates that the proteins NUSA (Mr 55K), S-adenosylmethionine synthase (42K), elongation factor Tu (43K), RNA polymerase α-chain (37K) and 50S ribosomal protein L7/L12 (12K) interact with GroEL for initial folding and return to GroEL for conformational maintenance. They are released from GroEL in vitro when GroES and ATP are added (data not shown).
The positions of 52 identified GroEL substrates on the Coomassie-stained 2D gel are indicated (see Table 1). In some cases, several spots correspond to the same protein, perhaps as a result of carbamylation of proteins after prolonged exposure to urea. Bottom right, insets show the region around the GroEL spot from Coomassie-stained 2D gels of total soluble cytoplasmic proteins obtained before and after immunoprecipitation.
We observed that the α and β subunits of the RNA polymerase core complex and the ω subunit of the holoenzyme17 are substrates of GroEL (Table 1), in accord with previous genetic and biochemical evidence18,19. The α subunit of the RNA polymerase represents 0.6% of the total cellular protein20. Under normal growth conditions, ∼2% of the subunit was found to be associated with the chaperonin at any one time (Fig. 4), in agreement with GroEL being involved in its maintenance.
A preferred structural motif
Sequence analysis of the 52 identified GroEL substrates failed to reveal statistically significant consensus sequences or clusters of consensus sequences. Given the preference of GroEL for a subset of E. coli proteins in vivo, we considered the possibility that many of these proteins contain common structural motifs or folds that form the basis of their interaction with the chaperonin.
To investigate this possibility, we focused the analysis on the GroEL substrates with known 3D structures or with homologues of known structures, using the protein domain-classification databases SCOP21 and CATH22. For every protein, domains were classified independently through sequence homology to domains in these databases. Proteins containing a stretch of more than 100 contiguous amino acids not covered by sequence homology to domains in SCOP (or CATH) were excluded from the analysis. Of the 52 identified GroEL substrates, 24 were amenable to structural analysis (18 proteins using homologies with >17% identities, or 24 with >13% identities; see Methods and Table 1). The 24 proteins were representative GroEL substrates, based on their Mr distributions and predictions of secondary structure. Both SCOP and CATH gave similar results. We found that, with high statistical significance, GroEL substrates preferentially contain several αβ domains compared with E. coli proteins (Fig. 5a). Of the multidomain GroEL substrates, 13 of 17 have at least two αβ domains (Table 1). In contrast, no significant preference for all α, all β, discontinuous domains, or oligomeric state was found for GroEL substrates, compared with E. coli proteins, although these types of proteins are not excluded from the set of substrates.
a, SCOP or CATH domain-classification databases were used to determine the number of proteins in E. coli (grey bars, ∼400 proteins) or in the set of identified GroEL substrates (black bars, 18 proteins) with at least one, two or three αβ domains. (A very similar result was obtained when including the additional six GroEL substrates which have a lower homology score.) The differences observed (asterisk) between the E. coli set and the GroEL substrates were found to be statistically significant at the 95% confidence limit by using equations developed for sampling statistics (see ref. 47). b, Structures of representative GroEL substrates. Top, the whole protein subunit chain, colour coded by domains. Domains that are classified as three-layer αβα or two-layer αβ sandwiches according to CATH, except for RNA polymerase α chain (RPOA) whose structural classification is based on ref. 48 and was not included in the statistical analysis, are depicted individually with helices in blue, strands in red and other secondary structure elements in grey. The hydrophobic face of the buried sheet for each of the domains is shown as molecular surface colour-coded according to hydrophobicity (yellow is hydrophilic and blue is hydrophobic) based on a scale defined in GRASP49. The names of GroEL substrates are abbreviated according to Table 1. Protein Database files used are METK, 1mxb; G3P1, 1gadO; PYRB, 1raiA; RECA, 2reb; RPOA, 1bdfA + 1coo.
The preference of GroEL for multiple αβ domains in the GroEL substrates, compared with soluble E. coli proteins, is not due to a difference in Mr distribution between the two sets. If the structural analysis is restricted to proteins of Mr > 20K, to match the size bias of GroEL-bound proteins (Fig. 1f), the same significant preference for multiple αβ domains is obtained for the GroEL substrates. Furthermore, based on their Mr distribution and predicted secondary structure, the set of E. coli proteins from SCOP or CATH faithfully represent the set of total soluble proteins in terms of their fold classification (data not shown).
The most common domain architectures in GroEL substrates were those of the three-layer αβα sandwich and, with higher preference, the two-layer αβ sandwich (data not shown). As a common occurrence in such structural motifs23, the β-sheets expose a hydrophobic surface packed against the hydrophobic surfaces of α-helices (Fig. 5b). These β-sheets and the corresponding hydrophobic faces of the α-helices would provide ideal hydrophobic surfaces to mediate high-affinity interactions with the apical domains of GroEL. Several stringent model substrates of GroEL that are typically used for in vitro studies, including ornithine transcarbamylase24, malate dehydrogenase25, rhodanese10 and RUBISCO26, belong to this category of αβ proteins.
Substrate interaction with GroEL
The finding of a preferred domain topology in GroEL substrates provides insight into why and how these proteins interact with the chaperonin. For αβ domains, the formation of the β-sheet is expected to be the most difficult step in the folding process because, unlike the formation of an α-helix, assembly of the β-sheet requires the formation of a large number of specific long-range contacts in the proper orientation. In general, these domains are expected to exhibit relatively slow folding rates27, and misfolding or kinetic trapping may occur either through the improper packing of helices and sheets within one domain or between domains, or owing to the packing of helices in one molecule against a sheet in another molecule. As GroEL substrates consist preferentially of two or more αβ domains, these proteins may be particularly prone to aggregation as a consequence of 3D domain swapping28.
The estimated number of proteins with multiple αβ domains in the E. coli cytoplasm is between 200 and 600. GroEL may assist in the initial folding and conformational maintenance of a subset of these proteins in several ways. The hydrophobic residues exposed on two flexible helices and a loop region in the apical domain of GroEL, which mediate polypeptide binding6, could provide an adjustable scaffold to stabilize the β-sheet of the substrate protein, essentially by acting as a substitute for the helices in the native protein. Subsequently, folding of a single protein molecule would proceed upon GroES-mediated displacement into the enclosed GroEL–GroES cavity. If helices and sheet(s) in a substrate protein have been packed improperly, multiple apical domains of GroEL could interfere by binding the helices29, the β-sheet or both, thereby dissociating their improper interaction. Similarly, for structurally labile αβ proteins, partial unfolding would lead to the exposure of the hydrophobic surfaces of the helices and the buried β-sheets which can be bound by GroEL. These proteins would therefore continually require GroEL for conformational maintenance and refolding. For several model proteins, the native state has been shown to be in equilibrium with unfolding intermediates that bind to GroEL30,31,32.
The typical GroEL substrate
The role of GroEL in assisting the folding or maintenance of a particular protein in vivo is expected to depend on the presence of appropriate binding sites for GroEL in the folding or unfolding intermediates of that protein, and on the rate at which these sites are buried during folding or refolding, respectively. Our findings indicate that a typical GroEL substrate has an Mr of between 20K and 60K, and consists preferentially of several αβ domains with buried hydrophobic β-sheets. The non-native states of these topologically complex proteins are likely to expose extensive hydrophobic surfaces that could be recognized by GroEL, either during folding or on misfolding. Co-expression of chaperonin may provide a rational strategy to improve the folding efficiency of foreign proteins expressed in E. coli when these proteins contain multiple αβ domains.
Methods
Cell labelling, immunoprecipitation and 2D-gel analysis
Pulse-chase experiments on E. coli MG1655 cells (F-λ-) followed by immunoprecipitation of GroEL–substrate complexes were carried out as described10. Radioactive proteins on SDS–PAGE or on 2D gels were visualized by using a FUJIFILM FLA-2000 phosphorimager. Control immunoprecipitations were performed from cell lysates that were heated to 95 °C for ∼1 min with 1% SDS and then diluted 10 times into lysis buffer containing 0.5% Triton X-100 (Fig. 3a, right, first lane). For controls of ATP with GroES (Fig. 3a, right, second lane), cells were lysed hypo-osmotically without EDTA, and then 500 µl of the cytoplasmic fraction was incubated for 30 min at room temperature with 10 mM ATP, 200 µg ml-1 creatine kinase and 10 mM creatine phosphate with 0.2 µM purified GroES33. Samples for 2D-gel analysis were focused using a 13-cm Immobiline DryStrip pH 4–7 L and the Multiphor II system (Pharmacia), followed by SDS–PAGE34,35. Kinetics of flux through GroEL of newly translated proteins (Fig. 2) were determined by measuring the volume of the radioactive spots corresponding to each protein from the 2D gel of the anti-GroEL immunoprecipitation at different chase times using ImageMaster 2D Elite (Pharmacia). The volume obtained at each time point was normalized to the GroEL volume at that time point and scaled according to the increase in the intensity of the GroEL band as a function of chase time visualized on SDS–PAGE.
Identification of GroEL substrates
GroEL–substrate complexes were isolated by large-scale immunoprecipitation from 2-l cultures of MG1655 cells grown in minimal media to mid-log phase. Cells were lysed hypo-osmotically on ice and about 16 mg of affinity-purified goat anti-GroEL antibodies crosslinked to protein-G-Sepharose were used in the immunoprecipitations. Proteins were eluted from the beads using 8 M urea/4% CHAPS. Portions of the cytoplasmic fraction taken before and after immunoprecipitation of GroEL complexes were exchanged into 8 M urea/4% CHAPS. Samples were concentrated to about 300 µl for 2D-gel analysis (Amicon concentrators with Mr 3K cutoff) and loaded into 18-cm Immobiline DryStrip pH 3–10 NL for isoelectric focusing, followed by 9–16% SDS–PAGE. The pattern of spots on the 2D gel from the large-scale immunoprecipitation was reproducibly obtained in three separate repeats. Approximately 110 protein spots in the 2D gel were treated with trypsin followed by peptide-mass fingerprint analysis using MALDI-TOF mass spectrometry36. All proteins identified were found to be E. coli proteins, which further supported their correct identification.
Sequence and structure analysis
Pairwise and multiple sequence alignments were carried out using PSI-BLAST37 (BLOSUM62 matrix, filtering using SEG38, default gap penalty, and an e-value of 0.01). An all-against-all sequence comparison of the identified GroEL substrates gave only three pairs of proteins with significant sequence similarity: GATD_ECOLI and TDH_ECOLI; AROG_ECOLI and AROH_ECOLI, and PSTB_ECOLI and YCFV_ECOLI. Because up to one third of the E. coli proteins are estimated to have resulted from gene duplication events39, GroEL displays no preference for a particular gene family.
For secondary structure predictions, membrane proteins, defined as having two or more predicted transmembrane regions, were first identified using ALOM40. Structural predictions of the remaining globular proteins were then calculated using PREDATOR41. Proteins were attributed to αβ, allα, allβ, or irregular category classes42.
For homology-based fold assignments, iterative similarity searches using PSI-BLAST were carried out with each query protein sequence against SCOP21 or CATH22. Duplicate hits from different parts of discontinuous domains were removed. For E. coli, 492 proteins were obtained for the structural analysis using SCOP and 401 using CATH; for the group of 52 identified GroEL substrates, this number was 18 using either database. Homologies obtained with this procedure had identities of 17–100%. By using PSI-BLAST searches against a combination of the full-protein sequence database and SCOP or CATH domains43, six additional proteins were structurally assigned from the set of GroEL substrates through homologies with identities between 13 and 17%.
In CATH, protein domains are divided into four classes of αβ, mainly α, mainly β, and few secondary structures. Each class is then subdivided into different architectures. The most common architectures of the αβ class are those of the three-layer αβα sandwich, consisting of a relatively flat β-sheet sandwiched between two layers of α helices, and the two-layer αβ sandwich, consisting of a relatively flat β-sheet packed against a layer of α helices. In SCOP, as in CATH, a similar but more specific division of protein domains into different classes is found. The main domain classes found in SCOP are: α and β (typically consisting of a central β-sheet with repeated units of β strand–α helix–β strand), α plus β (containing segregated α and β regions), all α, all β, and multidomain classes. When using SCOP, we grouped the (α and β) and (α plus β) classes into one αβ class to be consistent with the CATH classification. Results of the structural and functional characterization of the 52 identified GroEL substrates will be available on the PEDANT server44 (http://pedant.mips.biochem.mpg.de).
References
Georgopoulos,C. & Welch,W. J. Role of the major heat shock proteins as molecular chaperones. Annu. Rev. Cell Biol. 9, 601–634 (1993).
Ellis,R. J. Roles of molecular chaperones in protein folding. Curr. Opin. Struct. Biol. 4, 117–122 (1994).
Hartl,F. U. Molecular chaperones in cellular protein folding. Nature 381, 571–580 (1996).
Fayet,O., Ziegelhoffer,T. & Georgopoulos,C. The groES and groEL heat shock gene products of Escherichia coli are essential for bacterial growth at all temperatures. J. Bacteriol. 171, 1379–1385 (1989).
Horwich,A. L., Low,K. B., Fenton,W. A., Hirshfield,I. N. & Furtak,K. Folding in vivo of bacterial cytoplasmic proteins: role of GroEL. Cell 74, 909–917 (1993).
Fenton,W. A., Kashi,Y., Furtak,K. & Horwich,A. L. Residues in chaperonin GroEL required for polypeptide binding and release. Nature 371, 614–619 (1994).
Coyle,J. E., Jaeger,J., Gross,M., Robinson,C. V. & Radford,S. E. Structural and mechanistic consequences of polypeptide binding by GroEL. Fold. Design 2, R93–R104 (1997).
Sigler,P. B. et al. Structure and function in GroEL-mediated protein folding. Annu., Rev. Biochem. 67, 581–608 (1998).
Ranson,N. A., White,H. E. & Saibil,H. R. Chaperonins. Biochem. J. 333, 233–242 (1998).
Ewalt,K. L., Hendrick,J. P., Houry,W. A. & Hartl,F. U. In vivo observation of polypeptide flux through the bacterial chaperonin system. Cell 90, 491–500 (1997).
Pedersen,S. Escherichia coli ribosomes translate in vivo with variable rate. EMBO J. 3, 2895–2898 (1984).
Xu,D. & Nussinov,R. Favorable domain size in proteins. Fold. Design 3, 11–17 (1998).
Xu,Z. H., Horwich,A. L. & Sigler,P. B. The crystal structure of the asymmetric GroEL–GroES–(ADP)7 chaperonin complex. Nature 388, 741–750 (1997).
Ellis,R. J. & Hartl,F. U. Protein folding in the cell: Competing models of chaperonin function. FASEB J. 10, 20–26 (1996).
Rye,H. S. et al. GroEL–GroES cycling: ATP and nonnative polypeptide direct alternation of folding-active rings. Cell 97, 325–338 (1999).
Ellis,R. J. Molecular chaperones: avoiding the crowd. Curr. Biol. 7, R531–R533 (1997).
Gentry,D. R. & Burgess,R. R. The cloning and sequence of the gene encoding the omega subunit of Escherichia coli RNA polymerase. Gene 48, 33–40 (1986).
Wada,M., Fujita,H. & Itikawa,H. Genetic suppression of a temperature-sensitive groES mutation by an altered subunit of RNA polymerase of Escherichia coli K-12. J. Bacteriol. 169, 1102–1106 (1987).
Ziemienowicz,A. et al. Both the Escherichia coli chaperone systems, GroEL/GroES and DnaK/DnaJ/GrpE, can reactivate heat-treated RNA polymerase. Different mechanisms for the same activity. J. Biol. Chem. 268, 25425–25431 (1993).
VanBogelen,R. A., Sankar,P., Clark,R. L., Bogan,J. A. & Neidhardt,F. C. The gene–protein database of Escherichia coli: edition 5. Electrophoresis 13, 1014–1054 (1992).
Hubbard,T. J. P., Ailey,B., Brenner,S. E., Murzin,A. G. & Chothia,C. SCOP: a structural classification of proteins database. Nucleic Acids Res. 27, 254–256 (1999).
Orengo,C. A. et al. The CATH database provides insights into protein structure/function relationships. Nucleic Acids Res. 27, 275–279 (1999).
Eisenberg,D., Wilcox,W. & McLachlan,A. D. Hydrophobicity and amphiphilicity in protein structure. J. Cell. Biochem. 31, 11–17 (1986).
Cheng,M. Y. et al. Mitochondrial heat-shock protien hsp60 is essential for assembly of proteins imported into yeast mitochondria. Nature 337, 620–625 (1989).
Rye,H. S. et al. Distinct actions of cis and trans ATP within the double ring of the chaperonin GroEL. Nature 388, 792–798 (1997).
Viitanen,P. V. et al. Chaperonin-facilitated refolding of the ribulsebisphosphate carboxylase and ATP hydrolysis by chaperonin 60 (GroEL) are K+ dependent. Biochemistry 29, 5665–5671 (1990).
Plaxco,K. W., Simons,K. T. & Baker,D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 277, 985–994 (1998).
Schlunegger,M. P., Bennett,M. J. & Eisenberg,D. Oligomer formation by 3D domain swapping: a model for protein assembly and misassembly. Adv. Protein Chem. 50, 61–122 (1997).
Landry,S. J., Jordan,R., McMacken,R. & Gierasch,L. M. Different conformations for the same polypeptide bound to chaperones DnaK and GroEL. Nature 355, 455–457 (1992).
Laminet,A. A., Ziegelhoffer,T., Georgopoulos,C. & Pluckthun,A. The Escherichia coli heat shock proteins GroEL and GroES modulate the folding of the beta-lactamase precursor. EMBO J. 9, 2315–2319 (1990).
Vitanen,P. V., Donaldson,G. K., Lorimer,G. H., Lubben,T. H. & Gatenby,A. A. Complex interactions between the chaperonin 60 molecular chaperone and dihydrofolate reductase. Biochemistry 30, 9716–9723 (1991).
Smith,K. E., Voziyan,P. A. & Fisher,M. T. Partitioning of rhodanese onto GroEL–chaperonin binds a reversibly oxidized form derived from the native protein. J. Biol. Chem. 273, 28677–28681 (1998).
Hayer-Hartl,M. K., Weber,F. & Hartl,F. U. Mechanism of chaperonin action: GroES binding and release can drive GroEL-mediated protein folding in the absence of ATP hydrolysis. EMBO J. 15, 6111–6121 (1996).
Bjellqvist,B., Pasquali,C., Ravier,F., Sanchez,J. C. & Hochstrasser,D. A nonlinear wide-range immobilized pH gradient for two-dimensional electrophoresis and its definition in a relevant pH scale. Electrophoresis 14, 1357–1365 (1993).
Gorg,A., Postel,W. & Gunther,S. The current state of two-dimensional electrophoresis with immobilized pH gradients. Electrophoresis 9, 531–546 (1988).
Fountoulakis,M. & Langen,H. Identification of proteins by matrix-assisted laser desorption ionization-mass spectrometry following in-gel digestion in low-salt, nonvolatile buffer and simplified peptide recovery. Anal. Biochem. 250, 153–156 (1997).
Altschul,S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Wootton,J. & Federhen,S. Statistics of local complexity in amino acid sequences and sequence databases. Comput. Chem. 17, 149–163 (1993).
Labedan,B. & Riley,M. Gene products of Escherichia coli: sequence comparisons and common ancestries. Mol. Biol. Evol. 12, 980–987 (1995).
Klein,P., Kanehisa,M. & DeLisi,C. The detection and classification of membrane-spanning proteins. Biochim. Biophys. Acta 815, 468–476 (1985).
Frishman,D. & Argos,P. Seventy-five percent accuracy in protein secondary structure prediction. Proteins 27, 329–335 (1997).
Nakashima,H., Nishikawa,K. & Ooi,T. The folding type of a protein is relevant to the amino acid composition. J. Biochem. 99, 153–162 (1986).
Huynen,M. et al. Homology-based fold predictions for Mycoplasma genitalium proteins. J. Mol. Biol. 280, 323–326 (1998).
Frishman,D. & Mewes,H. W. Pedantic genome analysis. Trends Genet. 13, 415–416 (1997).
Bairoch,A. & Apweiler,R. The SWISS-PROT protein sequence data bank and its supplement TrEMBL in 1998. Nucleic Acids Res. 26, 38–42 (1998).
Karp,P. D., Riley,M., Paley,S. M., Pellegrini-Toole,A. & Krummenacker,M. EcoCyc: Encyclopedia of Escherichia coli genes and metabolism. Nucleic Acids Res. 26, 50–53 (1998).
Meyer,S. L. Data Analysis for Scientists and Engineers. (Wiley, New York, 1975).
Zhang,G. & Darst,S. A. Structure of the Escherichia coli RNA polymerase alpha subunit amino terminal domain. Science 281, 262–266 (1998).
Nicholls,A., Sharp,K. A. & Honig,B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank K. Andersson for technical assistance and M. C. Bewley for help with Fig. 5b.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Houry, W., Frishman, D., Eckerskorn, C. et al. Identification of in vivo substrates of the chaperonin GroEL. Nature 402, 147–154 (1999). https://doi.org/10.1038/45977
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/45977
This article is cited by
-
An inventory of interactors of the human HSP60/HSP10 chaperonin in the mitochondrial matrix space
Cell Stress and Chaperones (2020)
-
Improved 1, 2, 4-butanetriol production from an engineered Escherichia coli by co-expression of different chaperone proteins
World Journal of Microbiology and Biotechnology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.