Abstract
TNF receptor-associated periodic syndrome (TRAPS) is an autosomal dominant disorder characterized by recurrent attacks of fever and serositis. To date, more than 30 mutations have been reported in TNFRSF1A, the responsible gene. In Caucasian populations, the P46L (c.224C>T) TNFRSF1A sequence variation is considered as a low-penetrance mutation because its allele frequency is similar in patients and controls (∼1%). Whereas the spectrum of TNFRSF1A gene mutations has been well established in Caucasian and several Mediterranean populations, it remains unknown in sub-Saharan African populations. In this study, we found an unexpected high P46L allele frequency (∼10%) in two groups from West Africa – a group of 145 patients with sickle cell anaemia and a group of 349 healthy controls. These data suggest that the P46L variant is rather a polymorphism than a TRAPS causative mutation. We propose that the P46L high frequency in West African populations could be explained by some biological advantage conferred to carriers.
Similar content being viewed by others
Introduction
TNF receptor-associated periodic syndrome (TRAPS, MIM#142680) is the most frequent autosomal dominant disorder characterized by recurrent attacks of fever, abdominal pain, acute synovial and cutaneous inflammation. More than 30 mutations have been so far reported in TNFRSF1A, the causative gene. Most of these mutations are clustered in the region spanning exons 2–4 of the genomic sequence, encoding cysteine-rich extracellular domains of the protein. Upon cellular activation, impaired cleavage of the TNFRSF1A ectodomain with correlative decrease in the plasma concentration of the soluble receptor has been proposed as one of the mechanisms underlying the hyper-inflammatory response in TRAPS.1 The P46L (c.224C>T) mutation was initially reported in one patient with TRAPS, but was also found in 1.23% of Caucasian and African American control chromosomes.2 As the authors also demonstrated in vitro that P46L reduced TNFRSF1A shedding in monocytes, they concluded that P46L is a low-penetrance mutation. A broader study including patients and controls of Caucasian, Arab, Armenian and Sephardic Jewish extractions confirmed previously reported results by demonstrating P46L similar allele frequencies in patients (1.6%) and controls (1.3%).3 Whereas the spectrum of TNFRSF1A gene mutations has been well established in several Caucasian and Mediterranean populations, it remains unknown in sub-Saharan African populations. In this study, we investigated P46L allele frequency in 494 unrelated African individuals not known to have any symptoms compatible with TRAPS.
Materials and methods
Population analysis
The studied population consisted of two groups. The first group was composed of 145 patients (71 males and 74 females, average age 33.9 years) with typical sickle cell anaemia caused by homozygous SS beta-globin gene mutation; all originated from different African sub-Saharan countries (Benin, Cameroon, Congo, Ivory Coast, Togo and Zaire) and living in France. The second group was composed of 349 healthy women living in Togo and Ivory Coast (average age 26.1 years). Informed consent was obtained from the subjects according to the French Ethics Committee recommendations. The expected genotype distributions were calculated assuming a Hardy–Weinberg equilibrium. The observed and expected genotype distributions and allele frequencies were compared using χ2 test.
Mutational analysis
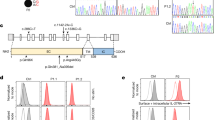
The P46L mutation creates a StuI restriction site. After genomic DNA extraction, exon 3 of TNFRSF1A gene was amplified by PCR using the following oligonucleotides: forward primer 5′-GGGCTCCTTCCTTGTGTTCT-3′ and reverse primer 5′-CTGACTCTCCTGCCTGTGC-3′, and digested for 4 h at 37°C with Eco147I (StuI) (Fermentas, MBI, USA). PCR products were visualized on 2.5% agarose gels stained with ethidium bromide.
Results
In the group of sickle cell anaemia patients of different African origins (see Materials and methods for details) and without any suspicion of TRAPS (n=145), 13 P46L heterozygous and one P46L homozygous individual were found among men and nine heterozygotes were found among women (total allele frequency 8.3%, Table 1). We found 48 heterozygotes and two homozygotes for P46L among 264 symptom-free women living in Togo (allele frequency 9.8%) and 17 heterozygotes and two homozygotes among 85 healthy women from Ivory Coast (allele frequency 12.4%). The P46L frequency was comparable in studied groups (8.3 vs 9.8%, P=0.45; 8.3 vs 12.4%, P=0.16; 9.8 vs 12.4%, P=0.35). We did not find any significant difference between observed and expected genotype distributions according to the Hardy–Weinberg equilibrium in the studied population (not shown). In addition, we did not find any individual carrying the P46L allele among 328 healthy control subjects of Caucasian (n=88), Sephardic Jewish (n=176) and Arab origins (n=64).
Discussion
The P46L variant was first described as a TRAPS-causative mutation because it was found in one patient suffering from periodic fevers and because it reduced TNFRSF1A shedding in monocytes.2 The defective TNFRSF1A shedding, proposed as the mechanism underlying the hyper-inflammatory response in TRAPS,1 has been demonstrated in monocytes from patients with P46L.2, 4 However, the defective shedding is neither correlated with the presence of TNFRSF1A gene mutations nor necessary for the development of TRAPS. Indeed, an impaired TNFRSF1A shedding was not present in TRAPS patients with C30S, T37I, R42Q, del42, C52R and N65I TNFRSF1A mutations,2, 5, 6 but it was observed in affected members of three TNFRSF1A mutation-negative families.6 Supplementary investigations must be performed to understand the exact molecular mechanisms of TRAPS and the significance of an impaired TNFRSF1A shedding.
Until now, P46L was considered as a low-penetrance mutation. It was found in 0.6% of Caucasian control chromosomes and in 1.9% of African American control chromosomes.2 A recent study3 showed similar P46L allele frequencies in patients (1.6%) and controls (1.3%) of Caucasian, Arab, Armenian and Sephardic Jewish extractions. The present study highlighted an unexpected high P46L allele frequency (9.8%) in sub-Saharan African populations. Surprisingly, none of the P46L carriers identified in our study displayed symptoms compatible with TRAPS. Thus, this observation strongly suggests that the P46L allele may not be a deleterious mutation since it is not sufficient for TRAPS development.
The P46L frequency discrepancy observed between African and non-African populations may have no biological significance. Indeed, a founder effect and a genetic drift could not be excluded, thus a larger study implicating other sub-Saharan African populations and a haplotype study of the P46L carrier chromosomes must be undertaken. Nevertheless, we propose that P46L is a functional variant that may have a broader influence on susceptibility to inflammation. Indeed, this allele was found only in 1.9% (3/156) of African American control chromosomes,2 whereas in our study it was found in 9.8% (97/988, P<0.005) of West Africa native's chromosomes. This difference prompts us to speculate that the P46L allele may provide some biological advantage to carriers living in West Africa. A few infectious diseases actually cause selective genetic pressure on host allele frequencies7 and the implication of TNFα and its receptors in infectious diseases has already been suggested.8, 9, 10 Further investigations must be performed to test this hypothesis.
References
McDermott MF, Aksentijevich I, Galon J et al: Germline mutations in the extracellular domains of the 55 kDa TNF receptor, TNFR1, define a family of dominantly inherited autoinflammatory syndromes. Cell 1999; 97: 133–144.
Aksentijevich I, Galon J, Soares M et al: The tumor-necrosis-factor receptor-associated periodic syndrome: new mutations in TNFRSF1A, ancestral origins, genotype–phenotype studies, and evidence for further genetic heterogeneity of periodic fevers. Am J Hum Genet 200; 69: 301–314.
Dode C, Andre M, Bienvenu T et al: The enlarging clinical, genetic, and population spectrum of tumor necrosis factor receptor-associated periodic syndrome. Arthritis Rheum 2002; 46: 2181–2188.
Arkwright PD, McDermott MF, Houten SM et al: Hyper IgD syndrome (HIDS) associated with in vitro evidence of defective monocyte TNFRSF1A shedding and partial response to TNF receptor blockade with etanercept. Clin Exp Immunol 2002; 130: 484–488.
Dode C, Papo T, Fieschi C et al: A novel missense mutation (C30S) in the gene encoding tumor necrosis factor receptor 1 linked to autosomal-dominant recurrent fever with localized myositis in a French family. Arthritis Rheum 2000; 43: 1535–1542.
Aganna E, Hammond L, Hawkins PN et al: Heterogeneity among patients with tumor necrosis factor receptor-associated periodic syndrome phenotypes. Arthritis Rheum 2003; 48: 2632–2644.
Segal S, Hill AV : Genetic susceptibility to infectious disease. Trends Microbiol 2003; 11: 445–448.
Rothe J, Lesslauer W, Lotscher H et al: Mice lacking the tumour necrosis factor receptor 1 are resistant to TNF-mediated toxicity but highly susceptible to infection by Listeria monocytogenes. Nature 1999; 364: 798–802.
Keane J, Gershon S, Wise RP et al: Tuberculosis associated with infliximab, a tumor necrosis factor alpha-neutralizing agent. N Engl J Med 2001; 345: 1098–1104.
Mombo LE, Ntoumi F, Bisseye C et al: Human genetic polymorphisms and asymptomatic Plasmodium falciparum malaria in Gabonese schoolchildren. Am J Trop Med Hyg 2003; 68: 186–190.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tchernitchko, D., Chiminqgi, M., Galactéros, F. et al. Unexpected high frequency of P46L TNFRSF1A allele in sub-Saharan West African populations. Eur J Hum Genet 13, 513–515 (2005). https://doi.org/10.1038/sj.ejhg.5201344
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201344