Abstract
Glycerol kinase (GK) is at the interface of fat and carbohydrate metabolism and has been implicated in insulin resistance and type 2 diabetes mellitus. To define GK's role in insulin resistance, we examined gene expression in brown adipose tissue in a glycerol kinase knockout (KO) mouse model using microarray analysis. Global gene expression profiles of KO mice were distinct from wild type with 668 differentially expressed genes. These include genes involved in lipid metabolism, carbohydrate metabolism, insulin signaling, and insulin resistance. Real-time polymerase chain reaction analysis confirmed the differential expression of selected genes involved in lipid and carbohydrate metabolism. PathwayAssist analysis confirmed direct and indirect connections between glycerol kinase and genes in lipid metabolism, carbohydrate metabolism, insulin signaling, and insulin resistance. Network component analysis (NCA) showed that the transcription factors (TFs) PPAR-γ, SREBP-1, SREBP-2, STAT3, STAT5, SP1, CEBPα, CREB, GR and PPAR-α have altered activity in the KO mice. NCA also revealed the individual contribution of these TFs on the expression of genes altered in the microarray data. This study elucidates the complex network of glycerol kinase and further confirms a possible role for glycerol kinase deficiency, a simple Mendelian disorder, in insulin resistance, and type 2 diabetes mellitus, a common complex genetic disorder.
Similar content being viewed by others
Main
Glycerol kinase (GK) catalyzes the phosphorylation of glycerol to glycerol 3-phosphate (G3P) which is important in the formation of triacylglycerol (TAG) and fat storage.1 GK is at the interface of fat and carbohydrate metabolism. GK deficiency (GKD) is an X-linked inborn error of metabolism that is characterized biochemically by hyperglycerolemia and glyceroluria and is due to mutations within or deletions of the GK gene on Xp21.1 Isolated GKD can be symptomatic or asymptomatic and we have previously shown that there is no genotype–phenotype correlation in isolated GKD.2, 3 We hypothesize that this lack of genotype–phenotype correlation makes even simple Mendelian disorders complex traits and that this complexity is due to the role of modifier genes, metabolic flux through related pathways, systems dynamics, thresholds of protein functions, networks that the protein functions within as well as the moonlighting (alternative) functions of the enzyme.2, 3, 4, 5
There is an emerging role for GK in type 2 diabetes mellitus (T2DM) as individuals with a GK missense mutation, N288D, have the asymptomatic form of isolated GKD, increased risk for obesity, insulin resistance and T2DM.6 In addition, hepatocyte nuclear factor 4 alpha (HNF4α) is important for GK expression7 and mutations in HNF4α are associated with maturity onset diabetes of youth (MODY).8 HNF4α is an orphan nuclear receptor involved in regulating gluconeogenesis in the liver,9 insulin secretion, and directly activates the insulin gene.10, 11 Recently many studies have focused on the role of HNF4α variants and polymorphisms on T2DM.12, 13, 14, 15, 16, 17 Thiazolidinediones (TZDs) are drugs used to treat T2DM18 and have been shown to induce GK expression in adipocytes, which reduces free fatty acid (FA) secretion and increases insulin sensitivity.19, 20 Insulin resistance and T2DM results from a network of interactions between many genes and environmental factors.21 Identification of these interactions will allow a better understanding of the molecular mechanisms leading to insulin resistance and T2DM.
GK is not normally expressed in white adipocytes but it is induced by TZD's in this tissue.22 However, GK is expressed in brown adipose tissue (BAT) which is metabolically active.23 BAT expresses LPL (lipoprotein lipase) which releases FA from lipoproteins and increases FA uptake. TZDs also affect BAT24, 25 and decreased BAT activity is associated with obesity, insulin resistance, and hyperlipidemia.26 Maintenance of adequate stores of TAG, through esterification of G3P is essential for BAT functioning.27 In addition, insulin deficiency induces BAT glyceroneogenesis to produce more G3P which is important to preserve the normal metabolic activity of BAT. Therefore, we hypothesize that GK has a role in BAT energy homeostasis and insulin resistance.
To relate gene expression data to protein function, we used network component analysis (NCA) which reduces dimensionality of high-dimensional microarray data to a lower dimension.28, 29 This allows identification of hidden dynamics and patterns such as transcription factor activities, (TFAs) which may not be found by microarray analysis.
In this study, we investigated the role of the murine ortholog of GK, Gyk, in metabolism (fat and carbohydrate), and insulin sensitivity using microarray analysis of BAT from wild-type (WT) and Gyk knockout (KO) mice. We determined that Gyk deletion causes alterations in expression of genes involved in carbohydrate and lipid metabolism as well as insulin signaling. NCA determined Gyk's role in adipocyte-specific transcription. Our work confirms a role for GK in metabolism and insulin resistance and helps to understand the complexity of this single gene disorder.
Materials and methods
Animal care
Gyk-deficient mice (courtesy of W J Craigen, Baylor College of Medicine) were generated using 129/SvJ embryonic stem cells, bred to a C57B1/6J mouse, and then backcrossed onto C57B1/6J to make a congenic strain on C57B1/6J.30 WT controls are male littermates of the KO mice and were born to carrier mothers. The mice were on a normal 3.5% fat diet (Harlan Tekland) and experiments were per a UCLA Chancellor's Animal Research Committee approved protocol.
RNA isolation
Day three of life WT and Gyk KO mice in the fed state were killed and interscapular BAT was extracted, homogenized in Trizol reagent (GibcoBRL Life Technologies, Rockville, MD, USA), frozen in liquid nitrogen, and stored at −80°C. Three to five tissues were pooled, then RNA was isolated (Trizol reagent, Gibco BRL Life Technologies), purified (RNeasy minielute, Qiagen, Valencia, CA, USA), and DNase treated (Turbo DNA-free, Ambion Inc., Austin, TX, USA).
cDNA synthesis and hybridization
cDNA synthesis was performed on pooled RNA and hybridized to Affymetrix Mouse Genome 430 2.0. Gene chips (three WT and four KO) as described.31
Microarray analysis
Microarray data was analyzed using DNA-Chip (dChip) analyzer software.32 Two unsupervised learning methods (multidimensional scaling and hierarchical clustering with an Euclidean distance measure) were used on genes with a coefficient of variance between 0.3 and 10, and a percent present call of 20% to define the ‘most varying probesets’ (3776 probesets). Differentially expressed genes were filtered out using the criteria: fold change >1.5 between baseline (WT) and experimental (KO), absolute difference in the expression level between WT and KO >100, Student's t-test P-value <0.05, and percent present call of ⩾20%.
Gene annotation
Genes that met the above criteria were used to identify enriched biological themes by Expression Analysis Systemic Explorer (EASE)33 using the categorical overrepresentation function and the one-tailed Fisher exact probability for overrepresentation.
Real-time polymerase chain reaction
RNA was extracted as described above. KO and WT samples included some of the samples from microarray analysis as well as single animal samples. Gene expression assays (Applied Biosystems Inc. (ABI), Foster City, CA, USA) were used for all genes. Predeveloped Taqman assay reagents for 18 s rRNA (ABI) were the endogenous controls. cDNA synthesis was performed using Superscript III (Invitrogen Corp., Carlsbad, CA, USA) per the manufacturer's instructions with random primers. The reaction was carried out using Taqman master mix (ABI) and Real-time polymerase chain reaction (RT-PCR) products detected using ABI prism 7700/7500 Sequence detection system. Fold differences for each of the genes were calculated using the 2−ΔΔCT method.34
PathwayAssist analysis
Altered genes were analyzed using PathwayAssist (version 3.0,. Stratagene, La Jolla, CA, USA) by searching for connections of the genes listed in Tables 1 and 2 by looking for common regulators, and finding the shortest paths between nodes.
NCA
PUBMED was used to construct the connectivity matrices between transcription factors (TFs) important in adipocytes and genes differentially expressed in the microarray analysis. NCA was performed with criteria of 1.2-fold change, absolute difference in the expression level between the WT and KO larger than 100, Student's t-test P-value <0.05, and percent present call ⩾20%. Ten TF were used to construct the final connectivity matrix. The data matrices were decomposed and the control strengths (CS), and transcription factor activity (TFA) matrices and contribution plots were obtained using the NCA toolbox (http://www.seas.ucla.edu/~liaoj/downloads/htm).
Results
Three-day-old mice were chosen for this study because this is the first day that the mice have statistically significant clinical symptoms including hypoglycemia, lower pH, lower bicarbonate, and lower base excess, which mimics the human disease.35 It was felt that on day of life two the important changes from glycerol kinase deficiency would not be present without the clinical changes seen in mice (and humans). We believe that the majority of the changes seen on day 3 will be owing to the effects of GKD, however one cannot rule out the possibility that some will be due to the perimortum state of the mice as they die on DOL4. Measurements of the brown fat pads in subsequent WT and KO mice have shown no statistical difference in weight (in g) (data not shown).
Unsupervised hierarchical clustering and multidimensional scaling using the 3776 most varying probesets showed that the gene expression in BAT in a Gyk KO mouse model grouped together according to KO and WT status (Figures 1a and b). In both the hierarchical clustering tree (Figure 1a) and the multidimensional scaling (Figure 1b) the bfko2 sample is linked to the WT samples however, it is still distinct from them. This unsupervised learning analysis demonstrates that Gyk KO mice have a distinct global gene expression profile compared to WT. Differential gene expression analysis (gene filtering) revealed 888 probesets (668 genes) significantly differentially expressed between KO and WT mice with a median false discovery rate of 5%. Of the 668 genes, 388 genes were downregulated and 280 were upregulated.
Unsupervised learning analysis of KO and WT mice samples using the 3777 most varying genes. (a) Average linkage hierarchical clustering tree of KO samples and WT samples. (b) Multidimensional scaling plot of the same samples. Black circles are KO, and open circles are WT. bfko represents brown fat knockout and bfmwt represent brown fat male wild type.
To uncover enriched biological themes among the differentially expressed genes we used the gene ontology analysis EASE software33 (Table 1). The analysis identified the most significant enrichment of differentially expressed genes to be in cell proliferation (P<0.001). Other suggestive (P-value ⩽0.02) biological and molecular themes included cell cycle, regulation of cellular process, regulation of cell cycle, regulation of cell proliferation, carbohydrate transport, lipid metabolism, sugar porter activity, and sugar transporter activity. EASE analysis of genes involved in lipid and carbohydrate metabolism revealed 40 genes (excluding Gyk) differentially expressed between the KO and WT mice (Tables 2A and B). Of these, 28 were in lipid metabolism and 12 in carbohydrate metabolism. Fifteen genes involved in lipid metabolism were downregulated in the KO mouse including lysosomal acid lipase 1 (Lip1), fatty acid synthase (Fasn), and leptin (Lep), (Table 2A), and 13 genes involved in lipid metabolism were upregulated, which included Lipin 1 (Lpin1), very low density lipoprotein receptor (Vldlr), and adiponectin receptor 1 (Adipor1) (Table 2A). Twelve genes involved in carbohydrate metabolism were differentially expressed (excluding Gyk); eight of which were downregulated including glycogen synthase 2 (Gys2), alpha-N-acetylglucosaminidase (Naglu), and pyruvate dehydrogenase beta (Table 2B). Four genes were upregulated including sorbitol dehydrogenase 1 (Table 2B).
To assess GK's role in insulin resistance and T2DM, we examined the differential gene expression of genes involved in insulin sensitivity including genes encoding insulin-receptor-associated proteins, components of and downstream effectors of the phosphatidylinositol 3-kinase (PI3K) and MAP kinase pathways, primary target genes for insulin resistance, effectors of insulin signaling, and target genes for peroxisome proliferator-activated receptor gamma (PPAR-γ). We found 25 probesets (19 genes) that were differentially expressed between KO and WT mice (Table 3) that relate to insulin signaling or insulin resistance. Ten probesets (7 genes) were downregulated including insulin-like growth factor 1 (Igf1), eukaryotic translation initiation factor 4E (Eif4e), sterol regulatory element binding factor 2 (Srebf2), leptin (Lep), jun oncogene (Jun), mitogen-activated protein kinase 6, and Fasn. Fifteen probesets (12 genes) were upregulated in the Gyk KO mouse including adrenergic receptor beta 1 and 3 (Adrb1 and Adrb3), CCAAAT/enhancer binding protein delta (Cebpd), and insulin-like growth factor binding protein 3 (Igfbp3).
Supervised hierarchical clustering based on the 28 and 12 differentially expressed genes (29 and 13 probesets) belonging to lipid and carbohydrate metabolism, respectively showed a clustering distinction between WT and KO mice (Figure 2a). The 19 genes (25 probesets) differentially expressed involved in insulin signaling and insulin resistance also showed distinct clustering of the WT and KO mice (Figure 2b).
Supervised hierarchical clustering of KO and WT samples; (a) dChip clustering tree of top 46 most significant differentially expressed probesets involved in lipid and carbohydrate metabolism. (b) dChip clustering tree of top 25 most significant differentially expressed probesets involved in insulin signaling and insulin resistance. The top genes for both clusters were filtered out using fold change larger than 1.5, absolute difference in the expression level larger than 100, Student's t-test P-value smaller than 0.05, and percent present call of at least 20%. bfko represents brown fat knockout and bfmwt represent brown fat male wild type.
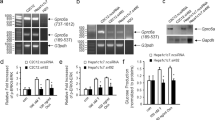
RT-PCR was performed on 30 genes found to be differentially expressed by microarray analysis. RT-PCR confirmed 20 of these 30 genes including the downregulation of carbohydrate and lipid metabolism genes including Gyk, Gys2, Naglu, and galactose-4-epimerase (Gale) (Figure 3 and Table 4). Sphingosine phosphate (Sgpl1), Lpin1, sulfotransferase family 1A, phenol-preferring, member 1 (Sult1a1), and flavin containing monooxygenase 1 (Fmo1) were upregulated in the KO mice by both RT-PCR and microarray expression data (Figure 3 and Table 4).
PathwayAssist analysis was used to represent the networks affected by Gyk deletion (Figure 4). Genes involved in lipid and carbohydrate metabolism, insulin signaling, and insulin resistance that were found to be altered in the microarray were used to create the pathway. For example, Gyk deletions lead to disruption of genes such as leptin that is implicated in insulin resistance. Central regulators in the pathway include PPAR-γ, PPAR-α, jun oncogene (JUN), tumor necrosis factor (TNF), glucose, and glucocorticoid. JUN is the most highly connected node, which likely reflects its global role as a transcription factor and it's involvement in a wide variety of biological processes.36, 37, 38, 39
PathwayAssist analysis involving differentially expressed genes from carbohydrate and lipid metabolism groups and from the insulin signaling and insulin resistance groups. Black nodes represent proteins coded by genes upregulated in the microarray analysis, gray nodes represent proteins coded by genes that were downregulated, and white nodes represents no change in the micorarray. Ovals represent genes involved in lipid and carbohydrate metabolism, hexagons represent genes involved in insulin and insulin resistance, rectangles represent genes that belong to both groups, diamonds represent common regulators of the differentially expressed genes, and octagonal nodes represent small molecules. Gyk is denoted by a triangle. ABCD2, ATP-binding cassette subfamily D member 2; ACACB, acetyl-Coenzyme A carboxylase beta; ADIPOR1, ADRB1, adrenergic receptor beta 1; ADRB3, adrenergic receptor beta 3; CEBPA, CCAAT/enhancer binding protein, alpha; CEBPD, CCAAT/enhancer binding protein delta; CEBPG, CCAAT/enhancer binding protein gamma; CITED2, Cbp/p300-interacting transactivator 2; CSF2, colony-stimulating factor 2; PRKCM, DGAT2, diacylglycerol O-acyltransferase 2; EIF4E, FDFT1, farnesyl diphosphate farnesyl transferase 1; GALE, galactose-4-epimerase, UDP; GH1, growth hormone 1; Glut1, glucose transporter 1; GC, glucocorticoid; GYS2, glycogen synthase 2; HPGD, hydroxyprostaglandin dehydrogenase 15; IFNG, interferon gamma; IGF1, insulin-like growth factor 1; IGFBP3, insulin-like growth factor binding protein 3; IL1B, interleukin 1 beta; IL1F8, interleukin 1 family, member 8; IL4, interleukin 4; IL5, interleukin 5; IL9, interleukin 9; INS, insulin; JUN, jun Oncogene; Lep, leptin; LIPA, lipaseA; LRPAP1, lipoprotein receptor-associated protein; LTC4S, leukotriene C4 synthase; NEU1, sialidase 1; NF κB1, nuclear factor kappa-B, subunit 1; PLAA, phospholipase A2-activating protein; PLCD3, phospholipase C, delta 3; PLD2, phospholipase d2; PPARA, peroxisome proliferator-activated receptor alpha; PPARG, peroxisome proliferator-activated receptor gamma; PRKCM, protein kinase C, mu; PTPN11, protein tyrosine phosphatase non-receptor type 11; SLC35A2, solute carrier family 35, member 2; SORD, sorbital dehydrogenase; SP1, sp1 transcription factor; SREBF2; SULT1a1, sulfotransferase family 1A; TCF1, transcription factor 1; TNF, VLDLR.
We elucidated transcription factor activities of TFs important in adipose tissue in the absence of Gyk using NCA of our microarray data as described in methods. Sixty-seven genes and 10 TFs resulted from the analysis. PPAR-γ, trans-acting transcription factor 1 (SP1), CCAAT/enhancer binding protein alpha, and glucocorticoid receptor (GR) activities were increased in the KO compared to the WT (Figure 5) whereas SREBP-2, SREBP-1, signal transducer and activator of transcription 3 (STAT3), STAT5, cAMP responsive element binding protein 1 (CREB), and PPAR-α activities were decreased in the KO (Figure 5). The connectivity, CS, expression matrices, and references used to deduce TFA and control strengths (CS) are provided as supplemental material. In order to rule out false discovery, we performed 100 permutations by shuffling the gene expression data and tested for significance. All activities of the 10 TFs showed perturbations when compared to the 100 permutated (shuffled gene expression) data (P-value <0.05) (data not shown). Contribution plots for phosphogluconate dehydrogenase (Pgd), angiopoietin-like 4 (Angptl4), fatty acid binding protein 4, adipocyte (Fabp4), Cebpd, Fasn, solute carrier family 2 (facilitated glucose transporter) member 3 (Slc2a3), insulin-like growth factor 1 receptor (Igf1r), and Igf1 are depicted in Figure 6. The graph shows the contribution to the total gene expression of the specific gene by each TF (hatched and white bars each represent a TF) and the actual gene expression from the microarray data is represented by black bars.
TFA of selected TFs important in adipose tissue. The transcriptional network containing 10 TF and 67 genes resulting from microarray analysis of 1.2-fold change. Peroxisome proliferator-activated receptor gamma (PPAR-γ), sterol regulatory element binding factor 2 (SREBP-2), sterol regulatory element binding factor 1 (SREBP-1), signal transducer and activator of transcription 3 (STAT3), signal transducer and activator of transcription 5 (STAT5), trans-acting transcription factor 1 (SP1), CCAAT/enhancer binding protein (C/EBP), alpha (CEBP-α), cAMP responsive element binding protein 1 (CREB), glucocorticoid receptor (GR), and peroxisome proliferator-activated receptor alpha (PPAR-α).
Scaling-independent contribution plots of a subset of genes. TF gene expression contribution for phosphogluconate dehydrogenase (Pgd), angiopoietin-like 4 (Angptl4), fatty acid binding protein 4, adipocyte (Fabp4), CCAAT/enhancer binding protein (C/EBP), delta (Cebp), Fasn, solute carrier family 2 (facilitated glucose transporter), member 3 (Slc2a3), insulin-like growth factor I receptor (Igf1r), and Igf1. Each gene is represented by two bars, white/hatched, and black. The white and hatched bar represents two TFs as indicated contributing to the gene expression of that gene; the sum of the white and hatched areas represents the fitted gene expression computed by the NCA software. The black bar represents the actual gene expression data from the microarray analysis.
Discussion
We report gene expression profiles in the Gyk KO mouse to provide insights into the role of GK in brown fat, insulin resistance, and T2DM. The Gyk KO mouse model mimics human GKD in that the mice have hyperglycerolemia, metabolic acidosis, and hypoglycemia.30, 35 However, the Gyk KO mouse has a more severe phenotype as the mice have growth retardation and die in the neonatal period.30
Of the biological themes identified using EASE analysis, cell proliferation was the most highly enriched (P-value <0.0001). Two related biological themes, regulation of cell proliferation (P-value <0.001) and negative regulation of cell proliferation (P-value=0.01), were also significantly enriched in the KO mouse. This may relate to the emerging role of GK in apoptosis40 and energy metabolism at the outer mitochondrial membrane.41 GK binds to the voltage dependent anion channel (VDAC or porin) of the outer mitochondrial membrane,41, 42 a component of the mitochondrial permeability transition pore complex (PTPC) involved in cytochrome c release and apoptosis.41, 43
A large number of enriched biological themes were identified in the Gyk KO mouse (Table 1). This attests to the complex biological network, within which GK is functioning and may be due in part to its role in phosphorylating glycerol. Of particular interest is carbohydrate and lipid metabolism, which we focused on for further analysis.
Within the category of carbohydrate metabolism, the levels of Gys2 were downregulated in the KO mice. Glycogen synthase (GYS) is expressed in the liver, catalyzes the incorporation of a glycosyl residue into glycogen, and is regulated by insulin, glucose, and glucose-6- phosphate.44, 45 In the absence of GK, the cell may become more dependent upon glucose metabolism to drive cellular processes. Therefore, glucose will not be converted into glycogen and a reduction of glycogen synthase would be expected.
In the lipid metabolism group, Lpin1, involved in fat adipocyte differentiation, is upregulated in the Gyk KO, whereas Lip1, involved FA release, is downregulated. LIP1 catalyzes hydrolysis of cholesteryl esters and triglycerides, and is suppressed by insulin.46 These findings suggest that GK deletion possibly stimulates adipocyte differentiation and decreases fat hydrolysis. The stimulation of adipocyte differentiation is intriguing given the role of PPAR-γ in adipocyte differentiation and stimulation of Gyk expression.19, 20 The levels of PPAR-γ mRNA were not changed in the Gyk KO mice compared to the WT, although PPAR-γ is activated as determined by NCA thus demonstrating the importance of NCA. These studies suggest that GK has a role in adipocyte differentiation and may relate to its role in T2DM.
Differentially expressed genes involved in insulin signaling, insulin resistance, and T2DM identify a link between GK and T2DM. Phospholipase d2 (Pld2) and protein kinase C (Pkc) mRNA levels were upregulated in Gyk KO mice 2.3- and 1.5- fold respectively. PLD2 releases diacylglycerol (DAG) during the breakdown of phospholipids. DAG is involved in many different signal transduction pathways including the activation of PKC,47 which ultimately inhibits insulin action in muscle.48, 49, 50 The increase of PLD2 and PKC in the absence of GK suggests that the presence of GK is important for maintaining normal insulin action, which is consistent with insulin resistance in individuals with GKD.6
The pathways affected by GK deletion were examined (Figure 4). Pathway Assist analysis showed that JUN was the node with the highest connectivity. This is possibly due to the increase of leptin in the KO mice because leptin increases JUN transcriptional activity.51 JUN's central role is likely due to its extensive role as a transcription factor in numerous mechanisms.36, 37, 38, 39 SP1 was implicated to regulate five of the genes that were altered by Gyk deletions: Lep, Adrb1, solute carrier family 2 (facilitated glucose transporter) (Slc2a1), member 1, Jun, and leukotriene C4 synthase. This confirms SP1's role as a key mediator of ‘cross-talk’ between signaling pathways such as insulin sensitivity, and gene transcription.52 Although SP1 regulates genes that are affected by GK deletions, the direct role of GK on SP1 remains unknown. Insulin was also implicated in the regulation of five of the genes differentially expressed in the microarray: Adrb3, Pld2, Dgat2, Gys2, and Lip1. Other key regulators in the pathway include PPAR-γ, PPAR-α, glucose, glucocorticoid, and TNF. PathwayAssist analysis provides a better understanding of the complexity of the network that GK functions in and suggests that GK is a critical component in multiple pathways.
NCA determined the TFA of TFs important in BAT when GK is deleted including PPAR-γ, SREBP-1, SREBP-2, STAT3, STAT5, SP1, CEBPα, CREB, GR, and PPAR-α. The gene expression level of Srebf2, which codes for SREBP-2 was significantly decreased in the KO mice vs the WT mice (1.86-fold). All other TFs gene expression levels were not significantly altered. This variance of gene expression level and TFA shows the importance of NCA to determine TF protein function from gene expression data.
A total of 12 PPAR-γ target genes were altered in our microarray data including Angptl4 and Adipor1, which are both upregulated. Angptl4 has been shown to be upregulated during fasting.53 GKD is similar to fasting in that glycerol and FA are released. Fasn, uncoupling protein 3 (Ucp3), and Dgat2 are downregulated in the KO mice. Fasn is involved in FA synthesis and is probably decreased in the KO as a feedback response by the cells to compensate for the absence of GK, which blocks the esterification of FA. Dgat2 is responsible for the synthesis of triglycerides and needs G3P.54 We would predict TFs such as PPAR-γ involved in adipocyte differentiation to be altered in the KO mice as the cells attempt to compensate for the lack of triglyceride synthesis by activating adipocyte differentiation.
Aquaporin 7 (Aqp7), a PPAR-γ target gene, functions to transport glycerol into the blood stream when needed.55 Aqp7 is decreased in the KO mice, which is intriguing, as previous studies have shown that Aqp7 disruption leads to increased GK levels in adipocytes.56 The studies show that Aqp7 deficiency leads to obesity by increasing GK levels. The decrease in Aqp7 in the Gyk KO mice is most likely in response to the excess glycerol in the bloodstream (hyperglycerolemia). Therefore, glycerol does not need to be transported into the bloodstream and Aqp7 is not expressed.
We assessed the relative contributions of various TFs on specific gene expression. The expression of Slc2a3, a glucose transporter gene is increased in the Gyk KO mice. Slc2a3 gene expression is controlled by SP1 (inhibitor) and CREB (activator). The actual levels of Slc2a3 expression are higher in the KO because the activation of Slc2a3 by CREB is greater than the inhibition by SP1 (Figure 6). Fabp4 gene expression is increased in the Gyk KO mouse owing to the activation by both PPAR-γ and CEBP-α (Figure 6). Fasn encodes Fasn, which catalyzes the synthesis of palmitate from acetyl-CoA and malonyl-CoA into long-chain saturated FA. PPAR-γ antagonizes SREBP-1 to contribute to an overall decrease of Fasn gene expression (Figure 6). For all of the TFA, the fitted expression represented by the contributions of the TFs (hatched and white bars) correlated with the actual expression from the microarray data (represented by the black bars). This study confirms and expands our previous work indicating that NCA can be used in mammalian systems31 and allows better definition of the transcriptional network within which GK functions.
This study confirmed that GK has an integral part in the overall metabolic network including insulin signaling and begins to elucidate the relationship between GKD, insulin resistance, and T2DM. Specifically, our study showed that GK deletion causes alterations in gene expression levels of genes involved in lipid and carbohydrate metabolism, as well as other metabolic and transcriptional networks including genes involved in insulin resistance. Further investigations into the mechanisms of these alterations may provide greater insight into the links between glycerol and glucose metabolism, and the role of GK in insulin resistance and T2DM. Such investigations will help us understand the complexity of GKD and serve as a model for other complex genetic diseases.
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
McCabe ER : Disorders of glycerol metabolism; In: Scriver CR B, AL, Sly, WS, Valle, D, Childs, B, Vogelstein, B (eds): The metabolic and molecular basis of inherited disease. New York: McGraw-Hill, 2001, pp 2217–2237.
Dipple KM, McCabe ER : Phenotypes of patients with ‘simple’ Mendelian disorders are complex traits: thresholds, modifiers, and systems dynamics. Am J Hum Genet 2000; 66: 1729–1735.
Dipple KM, Zhang YH, Huang BL et al: Glycerol kinase deficiency: evidence for complexity in a single gene disorder. Hum Genet 2001; 109: 55–62.
Sriram G, Martinez JA, McCabe ER, Liao JC, Dipple KM : Single-gene disorders: what role could moonlighting enzymes play? Am J Hum Genet 2005; 76: 911–924.
Dipple KM, Phelan JK, McCabe ER : Consequences of complexity within biological networks: robustness and health, or vulnerability and disease. Mol Genet Metab 2001; 74: 45–50.
Gaudet D, Arsenault S, Perusse L et al: Glycerol as a correlate of impaired glucose tolerance: dissection of a complex system by use of a simple genetic trait. Am J Hum Genet 2000; 66: 1558–1568.
Stepanian SV, Huyn ST, McCabe ER, Dipple KM : Characterization of the human glycerol kinase promoter: identification of a functional HNF-4alpha binding site and evidence for transcriptional activation. Mol Genet Metab 2003; 80: 412–418.
Yamagata K FH, Oda N, Kaisaki PJ, Menzel S, Cox NJ, Fajans SS et al: Mutations in the hepatocyte nuclear factor-4alpha gene in maturity-onset diabetes of the young (MODY1). Nature 1996; 384: 458–460.
Rhee J, Inoue Y, Yoon JC et al: Regulation of hepatic fasting response by PPARgamma coactivator-1alpha (PGC-1): requirement for hepatocyte nuclear factor 4alpha in gluconeogenesis. Proc Natl Acad Sci USA 2003; 100: 4012–4017.
Bartoov-Shifman R, Hertz R, Wang H, Wollheim CB, Bar-Tana J, Walker MD : Activation of the insulin gene promoter through a direct effect of hepatocyte nuclear factor 4 alpha. J Biol Chem 2002; 277: 25914–25919.
Wang H, Maechler P, Antinozzi PA, Hagenfeldt KA, Wollheim CB : Hepatocyte nuclear factor 4alpha regulates the expression of pancreatic beta -cell genes implicated in glucose metabolism and nutrient-induced insulin secretion. J Biol Chem 2000; 275: 35953–35959.
Damcott CM, Hoppman N, Ott SH et al: Polymorphisms in both promoters of hepatocyte nuclear factor 4-{alpha} are associated with type 2 diabetes in the Amish. Diabetes 2004; 53: 3337–3341.
Ek J, Rose CS, Jensen DP et al: The functional Thr130Ile and Val255Met polymorphisms of the hepatocyte nuclear factor-4{alpha} (HNF4A): gene associations with type 2 diabetes or altered {beta}-cell function among Danes. J Clin Endocrinol Metab 2005; 90: 3054–3059.
Hansen SK, Rose CS, Glumer C et al: Variation near the hepatocyte nuclear factor (HNF)-4alpha gene associates with type 2 diabetes in the Danish population. Diabetologia 2005; 48: 452–458.
Bagwell AM, Bento JL, Mychaleckyj JC, Freedman BI, Langefeld CD, Bowden DW : Genetic analysis of HNF4A polymorphisms in Caucasian-American type 2 diabetes. Diabetes 2005; 54: 1185–1190.
Mohlke KL, Boehnke M : The role of HNF4A variants in the risk of type 2 diabetes. Curr Diab Rep 2005; 5: 149–156.
Silander K, Mohlke KL, Scott LJ et al: Genetic variation near the hepatocyte nuclear factor-4 alpha gene predicts susceptibility to type 2 diabetes. Diabetes 2004; 53: 1141–1149.
Lehmann JM, Moore LB, Smith-Oliver TA, Wilkison WO, Willson TM, Kliewer SA : An antidiabetic thiazolidinedione is a high affinity ligand for peroxisome proliferator-activated receptor gamma (PPAR gamma). J Biol Chem 1995; 270: 12953–12956.
Guan HP, Li Y, Jensen MV, Newgard CB, Steppan CM, Lazar MA : A futile metabolic cycle activated in adipocytes by antidiabetic agents. Nat Med 2002; 8: 1122–1128.
Guan HP, Ishizuka T, Chui PC, Lehrke M, Lazar MA : Corepressors selectively control the transcriptional activity of PPAR{gamma} in adipocytes. Genes Dev 2005; 19: 453–461.
Ashcroft F, Rorsman P : Type 2 diabetes mellitus: not quite exciting enough? Hum Mol Genet 2004; 13 (Spec no.): 1R21–1R31.
Salway J : Metabolism at a Glance. Nashville: Blackwell publisher, 2004.
Chakrabarty K, Chaudhuri B, Jeffay H : Glycerokinase activity in human brown adipose tissue. J Lipid Res 1983; 24: 381–390.
Valverde AM, Benito M : The brown adipose cell: a unique model for understanding the molecular mechanism of insulin resistance. Mini Rev Med Chem 2005; 5: 269–278.
Valverde AM, Benito M, Lorenzo M : The brown adipose cell: a model for understanding the molecular mechanisms of insulin resistance. Acta Physiol Scand 2005; 183: 59–73.
Sell H, Deshaies Y, Richard D : The brown adipocyte: update on its metabolic role. Int J Biochem Cell Biol 2004; 36: 2098.
Festuccia WTL, Kawashita NH, Garofalo MAR et al: Control of glyceroneogenic activity in rat brown adipose tissue. Am J Physiol Regul Integr Comp Physiol 2003; 285: R177–182.
Liao JC, Boscolo R, Yang YL, Tran LM, Sabatti C, Roychowdhury VP : Network component analysis: reconstruction of regulatory signals in biological systems. Proc Natl Acad Sci USA 2003; 100: 15522–15527.
Kao KC, Yang YL, Boscolo R, Sabatti C, Roychowdhury V, Liao JC : Transcriptome-based determination of multiple transcription regulator activities in Escherichia coli by using network component analysis. Proc Natl Acad Sci USA 2004; 101: 641–646.
Huq AH, Lovell RS, Ou CN, Beaudet AL, Craigen WJ : X-linked glycerol kinase deficiency in the mouse leads to growth retardation, altered fat metabolism, autonomous glucocorticoid secretion and neonatal death. Hum Mol Genet 1997; 6: 1803–1809.
MacLennan NK, Rahib L, Shin C et al: Targeted disruption of glycerol kinase gene in mice: expression analysis in liver shows alterations in network partners related to glycerol kinase activity. Hum Mol Genet 2006; 15: 405–415.
Li CW : WH: DNA-Chip Analyzer (dChip); In: Parmigiani G GE, Irizarry R, Zeger, SL (eds): The analysis of gene expression analysis and software. Springer: New York, 2003, pp 120–141.
Hosack D, Dennis G, Sherman B, Lane H, Lempicki R : Identifying biological themes within lists of genes with EASE. Genome Biology 2003; 4: R70.
Livak KJ, Schmittgen TD : Analysis of relative gene expression data sing real-time quantitative PCR and the 2-[Delta][Delta]CT method. Methods 2001; 25: 402–408.
Kuwada N, Nagano K, MacLennan N et al: Gene therapy for murine glycerol kinase deficiency: Importance of murine ortholog. Biochem Biophys Rese Commun 2005; 335: 247.
Angel P, Allegretto EA, Okino ST et al: Oncogene jun encodes a sequence-specific trans- activator similar to AP-1. Nature 1988; 332: 166.
Davis RJ : Signal transduction by the JNK group of MAP kinases. Cell 2000; 103: 239.
Angel P, Karin M : The role of Jun, Fos and the AP-1 complex in cell-proliferation and transformation. Biochim et Biophys Acta (BBA) - Rev Cancer 1991; 1072: 129.
Ip YT, Davis RJ : Signal transduction by the c-Jun N-terminal kinase (JNK) -- from inflammation to development. Curr Opin Cell Biol 1998; 10: 205.
Martinez Agosto JA, McCabe ERB : Conserved family of glycerol kinase loci in Drosophila melanogaster. Mol Genet Metab 2006; 88: 334.
McCabe ER : Microcompartmentation of energy metabolism at the outer mitochondrial membrane: role in diabetes mellitus and other diseases. J Bioenerg Biomembr 1994; 26: 317–325.
Seltzer WK, McCabe ER : Subcellular distribution and kinetic properties of soluble and particulate-associated bovine adrenal glycerol kinase. Mol Cell Biochem 1984; 64: 51–61.
Cesura AM, Pinard E, Schubenel R et al: The voltage-dependent anion channel is the target for a new class of inhibitors of the mitochondrial permeability transition pore. J Biol Chem 2003; 278: 49812–49818.
Ortmeyer HK : In vivo insulin regulation of skeletal muscle glycogen synthase in calorie-restricted and in ad libitum-fed rhesus monkeys. J Nutr 2001; 131: 907S–9912.
Villar-Palasi C, Guinovart J : The role of glucose 6-phosphate in the control of glycogen synthase. FASEB J 1997; 11: 544–558.
Abe H, Bandai A, Makuuchi M et al: Hyperinsulinaemia accelerates accumulation of cholesterol ester in aorta of rats with transplanted pancreas. Diabetologia 1996; 39: 1276–1283.
Huang KP : The mechanism of protein kinase C activation. Trends Neurosci 1989; 12: 425–432.
Griffin M, Marcucci M, Cline G et al: Free fatty acid-induced insulin resistance is associated with activation of protein kinase C theta and alterations in the insulin signaling cascade. Diabetes 1999; 48: 1270–1274.
Schmitz-Peiffer C, Browne C, Oakes N et al: Alterations in the expression and cellular localization of protein kinase C isozymes epsilon and theta are associated with insulin resistance in skeletal muscle of the high-fat-fed rat. Diabetes 1997; 46: 169–178.
Chen J-S, Exton JH : Regulation of phospholipase D2 activity by protein kinase C{alpha}. J Biol Chem 2004; 279: 22076–22083.
Olorunseun OO, Ian LPB : The anti-apoptotic and growth stimulatory actions of leptin in human colon cancer cells involves activation of JNK mitogen activated protein kinase, JAK2 and PI3 kinase/Akt. Int J Colorectal Dis 2007; 22: 401–409.
Samson S, Wong N : Role of Sp1 in insulin regulation of gene expression. J Mol Endocrinol 2002; 29: 265–279.
Koster A, Chao YB, Mosior M et al: Transgenic angiopoietin-like (Angptl)4 overexpression and targeted disruption of angptl4 and angptl3: regulation of triglyceride metabolism. Endocrinology 2005; 146: 4943–4950.
Cases S, Stone SJ, Zhou P et al: Cloning of DGAT2, a second mammalian diacylglycerol acyltransferase, and related family members. J Biol Chem 2001; 276: 38870–38876.
Maeda N, Funahashi T, Hibuse T et al: Adaptation to fasting by glycerol transport through aquaporin 7 in adipose tissue. Proc Natl Acad Sci USA 2004; 101: 17801–17806.
Hibuse T, Maeda N, Funahashi T et al: From the cover: aquaporin 7 deficiency is associated with development of obesity through activation of adipose glycerol kinase. Proc Natl Acad Sci USA 2005; 102: 10993–10998.
Acknowledgements
We thank Dr. Julian Martinez for helpful discussions and Dr WJ Craigen for the Gyk KO mice used to establish our colony. This work was supported by NIH Grants [DK60055 and GM067929].
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Rights and permissions
About this article
Cite this article
Rahib, L., MacLennan, N., Horvath, S. et al. Glycerol kinase deficiency alters expression of genes involved in lipid metabolism, carbohydrate metabolism, and insulin signaling. Eur J Hum Genet 15, 646–657 (2007). https://doi.org/10.1038/sj.ejhg.5201801
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201801
Keywords
This article is cited by
-
Identification of candidate genes that specifically regulate subcutaneous and intramuscular fat deposition using transcriptomic and proteomic profiles in Dingyuan pigs
Scientific Reports (2022)
-
Glycerol kinase 5 confers gefitinib resistance through SREBP1/SCD1 signaling pathway
Journal of Experimental & Clinical Cancer Research (2019)
-
Multiple-trait QTL mapping and genomic prediction for wool traits in sheep
Genetics Selection Evolution (2017)
-
Signaling network prediction by the Ontology Fingerprint enhanced Bayesian network
BMC Systems Biology (2012)
-
A dynamic network of transcription in LPS-treated human subjects
BMC Systems Biology (2009)