Abstract
Recently, a few genes have been reported to be causative in inflammatory diseases. Still, we are waiting for the vast majority to be discovered. New tools for genotyping and statistical analysis have been developed and emphasis has been put on study design. Coeliac disease (CD) is a disorder, where prolamins in dietary wheat gluten and related proteins from rye or barley are not tolerated. It is one of the most common chronic diseases in humans exceeding a population prevalence of 1%. In this article, we will summarise what is currently known about the genetics influencing CD with the emphasis on the non-HLA genetic component. We will discuss some difficulties when searching for susceptibility genes in disorders with complex inheritance patterns.
Similar content being viewed by others
Introduction
There are basically two genetic approaches to finding disease genes that are inherited to the offspring. The first is the genome-wide search (also referred to as ‘positional cloning’ since the gene is identified using its position in the genome). The second is the ‘functional candidate gene approach’ where genes are selected for investigation based on a known function.
Linkage analysis is used to approximately point out a chromosomal region, which is more often inherited by the affected individuals within families. Association analysis on the other hand depends upon the same allelic variant being present in seemingly unrelated individuals with a common ancestor or ‘founder’. Association analysis has been a method for fine-mapping and for candidate gene studies since the resolution is much higher in comparison to linkage. The number of meiotic events available to ‘cut’ up the chromosomes during recombination are few in linkage (one for each generation and child), while association is the result of a common ancestor many generations ago, and therefore, includes many meiotic events. Unlike linkage analysis, association studies depend upon the analysis of unrelated individuals, where a common sample includes a set of cases and control individuals but the so-called ‘trio’ families (one child and two parents) can also be used. Genetic linkage and association methods are based on DNA analysis and the use of DNA polymorphisms. When it comes to genome-wide studies, microsatellites have been the markers (genetic variants) traditionally used. However, due to the advances in single-nucleotide polymorphism (SNPs) technology, SNPs are now the variants of choice. SNPs can be used for linkage studies but more importantly also for genome-wide association (GWA) studies. Recently, the knowledge about copy number variants (CNVs) has exploded. These variants have been difficult to detect but new technology makes these a target for disease association studies as well. In addition to genetic linkage and association, analysing differential expression of genes and proteins can help by identifying pathways and interesting candidate genes. There is a large amount of mRNA and protein-expression data involving CD including whole-genome mRNA expression profiling studies.1, 2 To limit the scope, we will not discuss this approach further in the present review.
Coeliac disease (CD) is a disease with complex inheritance. After the remarkable success with positional cloning of genes responsible for monogenic traits, there has been comparably few success stories with complex polygenic traits.3 The main problem with linkage and association analysis on phenotypes with complex genetic inheritance compared to monogenic inheritance is that the relationship between a variant/mutation influencing the disease and the disease phenotype is weak and consequently difficult to detect. This is mainly due to the small contribution from each locus to the overall disease susceptibility. A number of problems encountered when dealing with complex inheritance and positional cloning strategies are listed in Box 1.
In light of these problems (described in Box 1), it is important to carefully think through and plan the study design before starting to collect samples on diseases with complex inheritance patterns. For example, very large study populations could be necessary. Whether the whole data set is analysed together or if a stratification approach is adopted, sample size is clearly one of the determinants of success. Thanks to the HapMap project, which was completed and published in October 2005,4 it is now possible to select a limited number of SNPs or CNVs to capture most of the variation in a given segment of the genome. Recent advances in technology have also made it possible to analyse thousands or even millions of genotypes in a few days at a cost that is reasonable. Not only thousands of samples but also thousands of different variants can be analysed.
Information gathered from each subject is of great importance; diagnosis, age of disease onset, disease symptoms, other diseases in the family, family history, ethnic background and birthplace of grandparents. All of these are examples of information, which should be carefully considered and could be very useful when conducting a genetic study. Every individual study and disease is likely to require an optimised study design that takes into account the unique characteristics of both the sample population and the phenotypes studied.5, 6, 7 In some cases, it might be useful to define the phenotype differently compared to the traditional phenotypic characteristics used when making the disease diagnosis. The phenotype–genotype correlation is, in most studies, the weakest link.6, 7
In this paper, we will review what is known so far about the genetics of CD. Several reviews have been written about CD and the human leukocyte antigen (HLA) component8, 9 as well as the epidemiology and diagnosis of CD.10, 11, 12 However, the so-called non-HLA genetic component of CD is currently under intense investigation and this will be the main focus of the present review.
Coeliac disease: the phenotype
CD is characterised by an immunological response to dietary gliadins in wheat and the corresponding prolamins in rye and barley. Individuals with CD characteristically develop a small bowel enteropathy with a villous atrophy, crypt hyperplasia, increased number of intra-epithelial lymphocytes and inflammation of the lamina propria as well as circulating disease-specific IgA-antibodies (anti-endomysium antibodies (EMA) and tissue transglutaminase 2 (TG2) antibodies).
From having been regarded as a gastrointestinal disease, CD is today rather considered a multisystemic complex inflammatory disease.13 Clinically CD is highly variable. Life-threatening states with severe diarrhoea, malnutrition, lethargy, oedema and/or anaemia are seen (even if they are rare) as well as clinically silent, asymptomatic persons without obvious symptoms at all. In dermatitis herpetiformis, a skin manifestation of gluten intolerance, severely itching blisters appear on elbows, knees and buttocks, most often in addition to symptomatic or non-symptomatic enteropathy.
Several diseases are associated with CD. Malignancy, especially in the gastrointestinal tract and particularly intestinal lymphoma, was already in the 1930s found to be more common in CD. A large Swedish registry follow-up study on 12 000 subjects verified this risk in adults.14 The standard incidence ratio (SIR) for all types of cancers was 1.3 (1.2–1.5; 95% confidence interval (CI)), for malignant lymphoma 5.9 (4.3–7.9) and for small intestinal cancer 10 (4.4–20). The risk of cancer declined with increasing length of follow-up after CD diagnosis and was not significantly increased after 10 years. Interestingly, in individuals first hospitalised with the diagnosis of CD before the age of 10 years, no significantly increased risk for malignant lymphoma was found. In contrast, individuals first hospitalised with CD older than 20 years had a considerably higher risk (SIR 7.0, 95% CI: 5.0–9.5). Further, the risk decreased over time for diagnosis of malignant lymphoma during the study period from SIR 12 (95% CI: 3.8–28) in 1970–1979 to 3.4 (95% CI: 1.9–5.7) in 1990–1995.
Osteopenia/osteoporosis is present at diagnosis of CD in all ages and in patients who fail to adhere to the gluten-free diet.15, 16 Treatment with gluten-free diet reduces or normalises the osteopenia.17 A special form of epilepsy with occipital calcifications is also linked to CD.18
Association between CD and autoimmune disorders has drawn special attention. Type 1 diabetes mellitus (T1D), thyroid diseases, Sjögren's syndrome and other autoimmune diseases are overrepresented in CD and CD is overrepresented when patients with these disorders are serologically screened for CD.19, 20, 21, 22, 23, 24 An Italian multicentre study demonstrated a strong correlation between increasing age at CD diagnosis and occurrence of other autoimmune disease.25 Failure to adhere to the gluten-free diet increases the risk for teenagers of having circulating organ-specific autoantibodies and autoimmune disease26 and first-degree relatives of CD patients have an increased prevalence of autoimmune diseases as well as silent CD.27 These findings point to the possibility of common genetic pathways in CD and autoimmune diseases, and raise the question if the upregulated immunological activity in untreated CD may contribute to the development of autoimmune disease in genetically susceptible individuals.
A higher prevalence of CD is found also among girls with Turner's syndrome28 and children with Down's syndrome.29
Epidemiology, population genetics
Within a few years in the early 1980s, a three to four times increase of childhood CD incidence was described in Sweden, but not in the neighbouring countries.30, 31, 32, 33 After 1996, a sudden decrease was noticed which was followed by a slow increase and a shift to higher ages and more silent phenotypes.33 The changes were parallel to changes in the infant-feeding pattern. Studies showed a higher gluten consumption in Swedish infants compared to countries with a lower incidence and to earlier Swedish studies.34 Also, a higher frequency of terminated breast feeding before the gluten introduction in children with diagnosed CD versus controls was found.35 The question whether these environmental factors are affecting the phenotype (ie, the intensity of the symptoms), and thus, the chance of being diagnosed rather than triggering the onset of the disease has been under debate.36
For genetic issues it is often of interest to compare epidemiological data between different regions, countries and populations. Different Caucasian populations show a prevalence of around 1% with surprisingly low variations when statistical margins of error are taken into consideration.37, 38 A female predominance of around 2:1 is generally found.39 Family studies have shown a sibling relative risk,40, 41 of approximately 10 (λs≈10) using a population prevalence of 1% and a sibling prevalence of 10%.42 In dizygotic twins, the sibling prevalence has been reported to be around 20%.43
Few epidemiological studies have been done in non-Caucasian populations. One exception is a study of Saharawi refugees, who show a surprisingly high prevalence of 5.6%, the highest population prevalence of CD ever found.44 Another study comparing populations from European and Asian origin demonstrated that the Punjabis had a four times higher incidence than Europeans.45
The positional cloning approach in CD
The first evidence of linkage in CD: chromosome 6 and HLA (‘CELIAC 1’)
The most common candidate gene region in inflammatory diseases is the major histocompatibility (MHC) complex on chromosome 6, which encodes for the human leukocyte antigens (HLA). The linkage and the association signals to HLA are typically strong in autoimmune and inflammatory diseases. Linkage and association to HLA were detected several decades ago in many diseases such as type 1 diabetes, rheumatic diseases and CD.46, 47, 48, 49, 50 However, finding the functional genes in HLA influencing these diseases has not been an easy task, in part due to the very strong linkage disequilibrium (LD) in the HLA region. Strong LD can complicate the fine mapping of disease associated genes, but in spite of this and uniquely for CD, the primary disease locus for CD in HLA has been identified.51 The HLA-DQ2 molecule, encoded by specific alleles of HLA-DQA1 and HLA-DQB1, is one of the strongest HLA associations among the HLA-associated diseases. In most populations, over 90% of CD patients carry the DQ2 heterodimer encoded by alleles DQA1*05 and DQB1*02.8 The final genetic evidence that HLA-DQA1 and -DQB1 encoded the functional variant causing CD was the discovery that the DQ2-risk molecule can be encoded from alleles DQA1*05 and DQB1*02 in cis- or in trans-51(Figure 1).
The HLA-DQ2 αβ heterodimer encoded in cis and in trans.8
Nearly all of the CD patients negative for DQ2 carry the DQ8 haplotype containing the risk allelic combination of DQA1*0301-DQB1*0302. A very small proportion of patients are both DQ2 and DQ8 negative, and they have been found to carry one of the two specific alleles coding for DQ2.52, 53 That the risks associated with the different HLA-susceptibility alleles differ between populations have been shown54 as well as a dose dependence of the DQB1*02 allele.55, 56, 57
Non-HLA-linked genes in CD
The alleles HLA-DQ2 or -DQ8 are necessary for the onset of CD but they cannot explain the whole genetic susceptibility to the disease. These alleles are also common in healthy individuals and together the population frequency is about 20–30%.51
The existence of additional susceptibility genes located outside of the HLA region is supported by the difference in concordance between monozygotic twins and dizygotic twins, who are identical in the HLA region and both carrying the susceptibility alleles. The pair wise concordance among monozygotic twins is over 70%, which is the highest reported in a disease with complex inheritance.43 In siblings sharing identical HLA haplotypes, the CD pair-wise concordance is around 20-30%, suggesting importance of genes located outside HLA in disease susceptibility.43, 58, 59 With current estimates of population prevalence (0.5–1%) the relative risk has decreased, and λs is approximately 10–20, λs (HLA) is around 5 and thus with a multiplicative model,60 the non-HLA component λs (non-HLA) is 2–4 and would confer a minor part of the total genetic component. However, these measurements should be taken with caution also considering that HLA is a necessary factor. The estimate λs is a measure of familiarity (a combination of the genetics and other factors which are shared among family members) and there could be an unknown environmental component diluting the effect of the non-HLA genes, if this component also tends to cluster within families.
Searching the entire genome: linkage studies and association studies
Several genome scans in CD have been performed in search for susceptibility genes outside the HLA region. These studies differ in respect to number of independent families and what types of families that are analysed. Some studies have only a few (multi-affected pedigrees), or even one single large pedigree, while others have adopted an affected sib-pair approach. The differences in outcome can partly be explained by these differences in family structure and partly because sample size is simply not large enough to be able to detect a gene with relatively small effect. Also, if related sib-pairs are analysed and not only independent ones, the linkage statistic can be inflated if it is not corrected for. Another reason for discrepancies is the use of different statistical methods. Some genome-wide scans have used non-parametric linkage (NPL), others have analysed the data using several different parameters and LOD scores or heterogeneity LOD scores (HLOD). Testing of different models and using different types of families (multi-affected pedigrees versus sib-pairs), makes the comparison between studies difficult. The genome-wide linkage scans are summarised in Table 1.
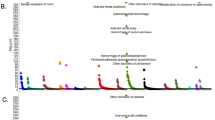
In a European collaboration, raw data from four genome scans including follow-up data were pooled and reanalysed.64 These were the second, third, fourth and fifth studies published.62, 66, 68, 69 The raw data consisted of genotypes from 442 families; 2025 individuals of whom 1056 are affected. The results pointed to chromosome 5q31–33 as being the only significant locus in these families apart from HLA (Figure 2).
Interestingly, the families from the United Kingdom showed negative linkage scores in this region. This could be the result of random variation due to a small number of families and locus heterogeneity. These families also have a somewhat different makeup, being large multigeneration pedigrees with many affected individuals. The three remaining data sets consisted of small nuclear families with typically 2–3 affected individuals. It is possible that many unrelated nuclear families favour the detection of a more common susceptibility gene shared by a large portion of CD patients, while large pedigrees with many affected individuals could be the result of a gene variant, which is relatively rare but with rather strong penetrance. This would illustrate how study design could influence the outcome of the results.
The first GWA analysis in CD has been reported and the results show convincing evidence that the IL2–IL21 region could be involved in CD.78 Several variants were highly significant when analysed in three populations; UK, Dutch and Irish case–control samples (shown in bold in Table 2). Strong LD makes it difficult to isolate one variant as disease causing and both IL2 and IL21 are expressed by T cells and have been implicated in other autoimmune conditions.79, 80 These genes are definitely interesting to follow up on in additional populations. Other SNPs from this GWA analysis, which need to be confirmed, are shown in Table 2. It is noteworthy that one of these variants is located in the 13th exon of LCT, the enzyme necessary for digesting lactose. Could there perhaps be a connection where malfunctioning lactose digestion during the first years of life, increases the risk for CD?
Fine-mapping and positional candidates on chromosomes 5 and 19
After linkage has been established, the next step is to fine map the linked region using association analysis. A simplifying condition for finding a disease gene by association is that most of the individuals with a certain disease are descendants of the same ancestor, therefore carrying a common haplotype with a so-called founder mutation. Association or LD between two alleles is the complex result of several factors, including recombination rate, allele frequencies and mutation rate. Reviews on the subject are written by Lander and Schork81 and Ardlie et al.82 If a disease mutation occurs next to a common allele, a large proportion of the normal population as well as affected individuals will carry this allele in coming generations. A marker that lies a bit further away could by chance have a rare allele on the founder chromosome and the correlation between that marker and the disease variant will be stronger. This illustrates the strong impact allele frequencies have on association.
When it comes to chromosome 5, a few published studies have focused on specific candidate genes in this region but not been able to find association.83, 84, 85, 86, 87 In addition, a large fine-mapping study failed to single out a strong susceptibility candidate.88 There were several genes showing nominal association in this Norwegian/Swedish family sample but no single association could explain the large linkage peak found in the same sample.88 Unless, there are regions inadequately covered due to, for example, copy number variation, these finding suggests that there is too much allelic heterogeneity or too many founder haplotypes to pin a gene down. Fine-mapping of the chromosome 19 region has been more successful than the one of the 5q region. The MYO9B gene was shown to associate in two different CD cohorts.73 It is a gene likely to be involved in Rho-dependent signalling pathways and remodelling of the cytoskeleton and tight junction assembly. This seems to fit very well with the possibility of CD being caused by a more permeable intestinal barrier. Attempts have been made but so far only one group has been able to replicate this finding89 while others have failed.90, 91, 92, 93, 94 Replications of genes with a small effect, or possibly, a population-specific effect can be difficult. However, control allele frequencies, calculated from untransmitted alleles in an Italian population, were significantly different from those of the Dutch control population; while patient allele frequencies were the same in the different populations.93 This questions the Dutch control population and could indicate a false positive finding of the MYO9B gene in the first place.
Other potential candidate genes on chromosome 19 include the intercellular adhesion molecule-1 precursor (ICAM-1) and CD209 both located in chromosomal band 19p13.2. A study of 489 CD patients and 257 families recently suggested that a functional variant of the CD209 promoter is associated with DQ2-negative CD patients.95 The ICAM-1 gene was found to associate to CD when analysing 180 unrelated French CD cases and 212 controls.96 When the authors stratified for age of disease onset, the same variant conferred even stronger predisposition to the adult-onset disease.96 Also from 19p13, cytochrome P450 F3 and F2 genes (CYP4F3 and CYP4F2) showed an effect on familial clustering in a Dutch CD cohort. CYP4F3 and CYP4F2 catalyse the inactivation of leukotriene B4 (LTB4), a potent mediator of inflammation responsible for recruitment and activation of neutrophils.97 Additionally on chromosome 19, a certain combination of expressed killer cell immunoglobulin-like receptor two (KIR2DL5) genotypes has shown association in a Spanish population of 413 cases and 231 controls.98
The candidate gene approach searching for non-HLA genes in CD
Cytotoxic T-lymphocyte-associated protein: CTLA4 (CELIAC 3)
CTLA4 plays an important role in maintaining tolerance to self-antigens, both as a negative regulator of T-cell proliferation through the co-stimulatory signal and as an inducer of clonal anergy. This gene has been under investigation in a number of autoimmune conditions. It is therefore an interesting candidate gene for CD. So far no evidence has come forward showing the involvement of a certain functional genetic variant of this gene in CD. However, there are many reports indicating the involvement of CTLA4 in CD.
The relative risk of CTLA4 in diabetes is estimated to be as low as 1.299 and if the same is true for CD, almost all studies in CD have too little power to be able to detect association. Still, some linkage to the region has been reported in a few studies68, 100, 74 and most published studies in CD do show signs of association as well.100, 74, 101, 102, 103, 104, 105, 106, 107, 108, 109, 110, 111 However, they are not completely congruent. Different studies show association to different haplotypes107, 112 and some studies fail to show association.113, 114, 115 Other studies with negative results could remain unpublished. Nevertheless, a negative result is not surprising if the relative risk is low and study samples are small. Different haplotypes showing association could be a result of allelic heterogeneity between and within populations but also of that CTLA4 can have different impacts on disease sub-phenotypes such as disease onset and severity. If one study includes many silent cases and another includes classical cases, this could be a reason for the different associations shown. Genotypes at HLA have also been suggested to influence the outcome of susceptibility to alleles in CTLA4.68, 110
Other non-HLA candidate genes potentially involved in CD
We searched PubMed (www.pubmed.gov) for ‘genetic AND (celiac OR coeliac) AND (associated OR association)’ and the database returned 405 hits. The genes presented in Table 3 are implicated from candidate gene studies, where a reported significance was below a P-value of 0.05. Furthermore, only those studies based on more than 100 unrelated CD patients are included below. This is indeed a very small sample size in a complex disease like CD, where odds ratios in the order of two or even less is expected. Some studies include many more individuals and thus these results could be more reliable. After excluding the HLA, chromosomes 5, 19 and CTLA4 regions, only seven studies in total fulfilled these criteria. In some cases, there have already been attempts made to replicate these findings. However, in most cases no convincing evidence can be presented (Table 3).
It is important that other research groups try and replicate suggestive findings since it is difficult to interpret the significance of genetic associations from a single report. Without a doubt, many of these studies will prove to be false-positive findings.
Evolutionary perspective
When searching for genes causing a complex disorder, it is interesting to try and understand the evolutionary and historical implications and how the disease is moulded by time.
One would think that individuals with variants conferring risk to a disease would be reduced in the population over time. It could be argued that if intolerance to gluten was an original state for human beings, we would never have adapted to this food source. However, in a disease like CD, it is possible that the disease state is the wild type or ‘natural state’ much like lactose intolerance is considered the wild type for humans. There are several factors that can be used to argue for how ‘intolerance’ variants could have been widespread, which in turn, would explain how humans adapted to the new food source all the same.
When looking at the HLA molecule known to cause CD – DQ2 – this molecule is frequent in the population and is considered ‘normal’ (ie, no damaging mutation hindering a protein to function properly). How is this possible, from an evolutionary perspective?
First, the complexity of the disorder makes individuals respond very different. Many affected individuals live to a high age and reproduce in spite of the disease.
Second, variants conferring susceptibility to disease could give an advantage in other situations or genetic backgrounds. HLA genes are examples of such genes; many variants of HLA are beneficial at the same time as they confer susceptibility to disease. If an individual is lacking the non-HLA-susceptibility genes it could be an advantage to have the DQ2 molecule conferring susceptibility to CD and vice versa.
Third, when it comes to CD, the percentage of gluten in today’s crops is increased by human intervention. When the first humans started to eat diploid wild-wheat the amount of gluten ingested was much lower than in today’s tetraploid breed of wheat. Although very small amounts of gluten induces relapse of the flat mucosa in CD patients, we do not know so much about the amount of gluten needed to trigger the disease. Maybe, the environmental trigger (gluten) was not strong enough to cause the disease in many individuals even if they had the genetic susceptibility variants.
Factors that contribute to keeping the CD-susceptibility genes in HLA fairly common in the population can equally keep non-HLA genes common.
Taking these factors in consideration it is possible that non-identified CD-susceptibility alleles might have a similar allele frequency as the HLA-susceptibility alleles, and could then also be considered the wild-type allele. If a CD-susceptibility locus was polymorphic 10 000 years ago, then this can mean that the genetic variant influencing CD could be common and ancient. For the search of this kind of variation, the haplotype-mapping approach and GWA analysis are very suitable.127, 128, 129
On the other hand, before humans started to eat wheat, CD-susceptibility variants could have had any frequency. If the allele influencing CD was the wild-type and non-polymorphic at the time when gluten was introduced in the population, it is possible that protective and relatively rare variants have developed over the years. These variants would have escalated in the population because of an increased selective pressure in a changing environment/gluten consumption. More recent mutations would suggest increased allelic heterogeneity. From this follows that analysing patients jointly from different parts of the world or even different parts of the same country, should be done with caution. However, if this would be the case, then we would expect a high variability of disease prevalence in different populations and the fact that CD seems equally prevalent in many populations speaks against this.
Future challenges
Current technologies have made it possible to also use association analysis for whole-genome scans. Although it is under debate if this is a good strategy to use, several such studies have already reported disease associations and more will follow. The IL2–IL21 region in CD is one of these findings and even if this is not the only non-HLA region influencing disease, there is evidence that one of these genes plays a part in CD susceptibility.
Two linked regions have reached a level of significant genome-wide linkage according to Lander and Kruglyak. These regions are 5q31—33, where linkage has been verified in several different independent populations64, 65 and 19p13.1 which show linkage in the Dutch population.73 One would think that these regions should be the first place to look for susceptibility genes. However, so far no convincing evidence has pinpointed a certain gene in any of these regions. New strategies using innovative phenotyping and using gene–gene interaction models might be necessary to locate disease genes. It is very possible that a combination of several different genes in the linked regions alone may be influencing the outcome of the disease and a strategy that takes this and other gene–gene interactions into account might be the way to move forward.
The puzzle is slowly being pieced together. Undoubtedly, several genes and genetic variants outside of the HLA complex add to the disease risk. Whether this risk increase is mainly due to rare variants, common ancestral variants or both remain to be seen.
References
Juuti-Uusitalo K, Maki M, Kaukinen K et al: cDNA microarray analysis of gene expression in coeliac disease jejunal biopsy samples. J Autoimmun 2004; 22: 249–265.
Diosdado B, Wapenaar MC, Franke L et al: A microarray screen for novel candidate genes in coeliac disease pathogenesis. Gut 2004; 53: 944–951.
Botstein D, Risch N : Discovering genotypes underlying human phenotypes: past successes for Mendelian disease, future approaches for complex disease. Nat Genet 2003; 33 (Suppl): 228–237.
Altshuler D, Brooks LD, Chakravarti A, Collins FS, Daly MJ, Donnelly P : A haplotype map of the human genome. Nature 2005; 437: 1299–1320.
Weiss KM, Buchanan AV : Evolution by phenotype: a biomedical perspective. Perspect Biol Med 2003; 46: 159–182.
Terwilliger JD, Goring HH : Gene mapping in the 20th and 21st centuries: statistical methods, data analysis, and experimental design. Hum Biol 2000; 72: 63–132.
Terwilliger JD, Haghighi F, Hiekkalinna TS, Goring HH : A bias-ed assessment of the use of SNPs in human complex traits. Curr Opin Genet Dev 2002; 12: 726–734.
Sollid LM : Molecular basis of celiac disease. Annu Rev Immunol 2000; 18: 53–81.
Louka AS, Sollid LM : HLA in coeliac disease: unravelling the complex genetics of a complex disorder. Tissue Antigens 2003; 61: 105–117.
Lee SK, Green PH : Celiac sprue (the great modern-day imposter). Curr Opin Rheumatol 2006; 18: 101–107.
Alaedini A, Green PH : Narrative review: celiac disease: understanding a complex autoimmune disorder. Ann Intern Med 2005; 142: 289–298.
Green PH, Rostami K, Marsh MN : Diagnosis of coeliac disease. Best Pract Res Clin Gastroenterol 2005; 19: 389–400.
Sollid LM : Coeliac disease: dissecting a complex inflammatory disorder. Nat Rev Immunol 2002; 2: 647–655.
Askling J, Linet M, Gridley G, Halstensen TS, Ekstrom K, Ekbom A : Cancer incidence in a population-based cohort of individuals hospitalized with celiac disease or dermatitis herpetiformis. Gastroenterology 2002; 123: 1428–1435.
Valdimarsson T, Lofman O, Toss G, Strom M : Reversal of osteopenia with diet in adult coeliac disease. Gut 1996; 38: 322–327.
Kalayci AG, Kansu A, Girgin N, Kucuk O, Aras G : Bone mineral density and importance of a gluten-free diet in patients with celiac disease in childhood. Pediatrics 2001; 108: E89.
Molteni N, Caraceni MP, Bardella MT, Ortolani S, Gandolini GG, Bianchi P : Bone mineral density in adult celiac patients and the effect of gluten-free diet from childhood. Am J Gastroenterol 1990; 85: 51–53.
Gobbi G : Coeliac disease, epilepsy and cerebral calcifications. Brain Dev 2005; 27: 189–200.
Scott BB, Losowsky MS : Coeliac disease: a cause of various associated diseases? Lancet 1975; 2: 956–957.
Collin P, Reunala T, Pukkala E, Laippala P, Keyrilainen O, Pasternack A : Coeliac disease – associated disorders and survival. Gut 1994; 35: 1215–1218.
Sategna-Guidetti C, Volta U, Ciacci C et al: Prevalence of thyroid disorders in untreated adult celiac disease patients and effect of gluten withdrawal: an Italian multicenter study. Am J Gastroenterol 2001; 96: 751–757.
Ansaldi N, Palmas T, Corrias A et al: Autoimmune thyroid disease and celiac disease in children. J Pediatr Gastroenterol Nutr 2003; 37: 63–66.
Ch’ng CL, Biswas M, Benton A, Jones MK, Kingham JG : Prospective screening for coeliac disease in patients with Graves’ hyperthyroidism using anti-gliadin and tissue transglutaminase antibodies. Clin Endocrinol (Oxf) 2005; 62: 303–306.
Reunala T, Collin P : Diseases associated with dermatitis herpetiformis. Br J Dermatol 1997; 136: 315–318.
Ventura A, Magazzu G, Greco L : Duration of exposure to gluten and risk for autoimmune disorders in patients with celiac disease. SIGEP study group for autoimmune disorders in celiac disease. Gastroenterology 1999; 117: 297–303.
Toscano V, Conti FG, Anastasi E et al: Importance of gluten in the induction of endocrine autoantibodies and organ dysfunction in adolescent celiac patients. Am J Gastroenterol 2000; 95: 1742–1748.
Petaros P, Martelossi S, Tommasini A, Torre G, Caradonna M, Ventura A : Prevalence of autoimmune disorders in relatives of patients with celiac disease. Dig Dis Sci 2002; 47: 1427–1431.
Bonamico M, Pasquino AM, Mariani P et al: Prevalence and clinical picture of celiac disease in Turner syndrome. J Clin Endocrinol Metab 2002; 87: 5495–5498.
Bonamico M, Mariani P, Danesi HM et al: Prevalence and clinical picture of celiac disease in Italian Down syndrome patients: a multicenter study. J Pediatr Gastroenterol Nutr 2001; 33: 139–143.
Ascher H, Krantz I, Kristiansson B : Increasing incidence of coeliac disease in Sweden. Arch Dis Child 1991; 66: 608–611.
Carlsson AK, Axelsson IE, Borulf SK, Bredberg AC, Ivarsson SA : Serological screening for celiac disease in healthy 2.5-year-old children in Sweden. Pediatrics 2001; 107: 42–45.
Cavell B, Stenhammar L, Ascher H et al: Increasing incidence of childhood coeliac disease in Sweden. Results of a national study. Acta Paediatr 1992; 81: 589–592.
Ivarsson A, Persson LA, Nystrom L et al: Epidemic of coeliac disease in Swedish children. Acta Paediatr 2000; 89: 165–171.
Ascher H, Holm K, Kristiansson B, Maki M : Different features of coeliac disease in two neighbouring countries. Arch Dis Child 1993; 69: 375–380.
Ivarsson A, Hernell O, Stenlund H, Persson LA : Breast-feeding protects against celiac disease. Am J Clin Nutr 2002; 75: 914–921.
Ascher H, Kristiansson B : The highest incidence of celiac disease in Europe: the Swedish experience. J Pediatr Gastroenterol Nutr 1997; 24: S3–S6.
Dube C, Rostom A, Sy R et al: The prevalence of celiac disease in average-risk and at-risk Western European populations: a systematic review. Gastroenterology 2005; 128: S57–S67.
Ascher H : Paediatric aspects of coeliac disease: old challenges and new ones. Dig Liver Dis 2002; 34: 216–224.
Ivarsson A, Persson LA, Nystrom L, Hernell O : The Swedish coeliac disease epidemic with a prevailing twofold higher risk in girls compared to boys may reflect gender specific risk factors. Eur J Epidemiol 2003; 18: 677–684.
Risch N : Assessing the role of HLA-linked and unlinked determinants of disease. Am J Hum Genet 1987; 40: 1–14.
Risch N : Linkage strategies for genetically complex traits. II. The power of affected relative pairs. Am J Hum Genet 1990; 46: 229–241.
Gudjonsdottir AH, Nilsson S, Ek J, Kristiansson B, Ascher H : The risk of celiac disease in 107 families with at least two affected siblings. J Pediatr Gastroenterol Nutr 2004; 38: 338–342.
Greco L, Romino R, Coto I et al: The first large population based twin study of coeliac disease. Gut 2002; 50: 624–628.
Catassi C, Ratsch IM, Gandolfi L et al: Why is coeliac disease endemic in the people of the Sahara? Lancet 1999; 354: 647–648.
Sher KS, Fraser RC, Wicks AC, Mayberry JF : High risk of coeliac disease in Punjabis. Epidemiological study in the south Asian and European populations of Leicestershire. Digestion 1993; 54: 178–182.
Falchuk ZM, Strober W : HL-A antigens and adult coeliac disease. Lancet 1972; 2: 1310.
Ludwig H, Polymenidis Z, Granditsch G, Wick G : [Association of HL-A1 and HL-A8 with childhood celiac disease]. Z Immunitatsforsch Exp Klin Immunol 1973; 146: 158–167.
Dausset J, Degos L, Hors J : The association of the HL-A antigens with diseases. Clin Immunol Immunopathol 1974; 3: 127–149.
Bluestone R : HL-A antigens in clinical medicine. Dis Mon 1976; 23: 1–27.
Stenszky V, Kozma L, Ambro I, Karmazsin L : Calculation of disease susceptibility gene frequency in insulin-dependent diabetes mellitus. Haematologia (Budap) 1978; 12: 141–148.
Sollid LM, Markussen G, Ek J, Gjerde H, Vartdal F, Thorsby E : Evidence for a primary association of celiac disease to a particular HLA-DQ alpha/beta heterodimer. J Exp Med 1989; 169: 345–350.
Karell K, Louka AS, Moodie SJ et al: HLA types in celiac disease patients not carrying the DQA1*05-DQB1*02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease. Hum Immunol 2003; 64: 469–477.
Spurkland A, Sollid LM, Polanco I, Vartdal F, Thorsby E : HLA-DR and -DQ genotypes of celiac disease patients serologically typed to be non-DR3 or non-DR5/7. Hum Immunol 1992; 35: 188–192.
Margaritte-Jeannin P, Babron MC, Bourgey M et al: HLA-DQ relative risks for coeliac disease in European populations: a study of the European Genetics Cluster on Coeliac Disease. Tissue Antigens 2004; 63: 562–567.
Ploski R, Ek J, Thorsby E, Sollid LM : On the HLA-DQ (alpha 1*0501, beta 1*0201)-associated susceptibility in celiac disease: a possible gene dosage effect of DQB1*0201. Tissue Antigens 1993; 41: 173–177.
Vader W, Stepniak D, Kooy Y et al: The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc Natl Acad Sci USA 2003; 100: 12390–12395.
Louka AS, Moodie SJ, Karell K et al: A collaborative European search for non-DQA1*05-DQB1*02 celiac disease loci on HLA-DR3 haplotypes: analysis of transmission from homozygous parents. Hum Immunol 2003; 64: 350–358.
Mearin ML, Biemond I, Pena AS et al: HLA-DR phenotypes in Spanish coeliac children: their contribution to the understanding of the genetics of the disease. Gut 1983; 24: 532–537.
Petronzelli F, Bonamico M, Ferrante P et al: Genetic contribution of the HLA region to the familial clustering of coeliac disease. Ann Hum Genet 1997; 61: 307–317.
Risch N : Linkage strategies for genetically complex traits. I. Multilocus models. Am J Hum Genet 1990; 46: 222–228.
Zhong F, McCombs CC, Olson JM et al: An autosomal screen for genes that predispose to celiac disease in the western counties of Ireland. Nat Genet 1996; 14: 329–333.
Greco L, Corazza G, Babron MC et al: Genome search in celiac disease. Am J Hum Genet 1998; 62: 669–675.
Greco L, Babron MC, Corazza GR et al: Existence of a genetic risk factor on chromosome 5q in Italian coeliac disease families. Ann Hum Genet 2001; 65: 35–41.
Babron MC, Nilsson S, Adamovic S et al: Meta and pooled analysis of European coeliac disease data. Eur J Hum Genet 2003; 11: 828–834.
Percopo S, Babron MC, Whalen M et al: Saturation of the 5q31–q33 candidate region for coeliac disease. Ann Hum Genet 2003; 67: 265–268.
King AL, Yiannakou JY, Brett PM et al: A genome-wide family-based linkage study of coeliac disease. Ann Hum Genet 2000; 64: 479–490.
King AL, Fraser JS, Moodie SJ et al: Coeliac disease: follow-up linkage study provides further support for existence of a susceptibility locus on chromosome 11p11. Ann Hum Genet 2001; 65: 377–386.
Naluai AT, Nilsson S, Gudjonsdottir AH et al: Genome-wide linkage analysis of Scandinavian affected sib-pairs supports presence of susceptibility loci for celiac disease on chromosomes 5 and 11. Eur J Hum Genet 2001; 9: 938–944.
Liu J, Juo SH, Holopainen P et al: Genomewide linkage analysis of celiac disease in Finnish families. Am J Hum Genet 2002; 70: 51–59.
Woolley N, Holopainen P, Ollikainen V et al: A new locus for coeliac disease mapped to chromosome 15 in a population isolate. Hum Genet 2002; 111: 40–45.
Popat S, Bevan S, Braegger CP et al: Genome screening of coeliac disease. J Med Genet 2002; 39: 328–331.
Neuhausen SL, Feolo M, Camp NJ, Farnham J, Book L, Zone JJ : Genome-wide linkage analysis for celiac disease in North American families. Am J Med Genet 2002; 111: 1–9.
Van Belzen MJ, Meijer JW, Sandkuijl LA et al: A major non-HLA locus in celiac disease maps to chromosome 19. Gastroenterology 2003; 125: 1032–1041.
Rioux JD, Karinen H, Kocher K et al: Genomewide search and association studies in a Finnish celiac disease population: identification of a novel locus and replication of the HLA and CTLA4 loci. Am J Med Genet A 2004; 130: 345–350.
van Belzen MJ, Vrolijk MM, Meijer JW et al: A genomewide screen in a four-generation Dutch family with celiac disease: evidence for linkage to chromosomes 6 and 9. Am J Gastroenterol 2004; 99: 466–471.
Garner CP, Ding YC, Steele L et al: Genome-wide linkage analysis of 160 North American families with celiac disease. Genes Immun 2007; 8: 108–114.
Lander E, Kruglyak L : Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
van Heel DA, Franke L, Hunt KA et al: A genome-wide association study for celiac disease identifies risk variants in the region harboring IL2 and IL21. Nat Genet 2007; 39: 827–829.
Sadlack B, Merz H, Schorle H, Schimpl A, Feller AC, Horak I : Ulcerative colitis-like disease in mice with a disrupted interleukin-2 gene. Cell 1993; 75: 253–261.
Monteleone G, Monteleone I, Fina D et al: Interleukin-21 enhances T-helper cell type I signaling and interferon-gamma production in Crohn’s disease. Gastroenterology 2005; 128: 687–694.
Lander ES, Schork NJ : Genetic dissection of complex traits. Science 1994; 265: 2037–2048.
Ardlie KG, Kruglyak L, Seielstad M : Patterns of linkage disequilibrium in the human genome. Nat Rev Genet 2002; 3: 299–309.
Louka AS, Torinsson Naluai A, D’Alfonso S et al: The IL12B gene does not confer susceptibility to coeliac disease. Tissue Antigens 2002; 59: 70–72.
Seegers D, Borm ME, van Belzen MJ et al: IL12B and IRF1 gene polymorphisms and susceptibility to celiac disease. Eur J Immunogenet 2003; 30: 421–425.
Ryan AW, Thornton JM, Brophy K et al: Haplotype variation at the IBD5/SLC22A4 locus (5q31) in coeliac disease in the Irish population. Tissue Antigens 2004; 64: 195–198.
Ryan AW, Thornton JM, Brophy K et al: Chromosome 5q candidate genes in coeliac disease: genetic variation at IL4, IL5, IL9, IL13, IL17B and NR3C1. Tissue Antigens 2005; 65: 150–155.
Wapenaar MC, Monsuur AJ, Poell J et al: The SPINK gene family and celiac disease susceptibility. Immunogenetics 2007; 59: 349–357.
Amundsen SS, Adamovic S, Hellqvist A et al: A comprehensive screen for SNP associations on chromosome region 5q31–33 in Swedish/Norwegian celiac disease families. Eur J Hum Genet 2007.
Sanchez E, Alizadeh BZ, Valdigem G et al: MYO9B gene polymorphisms are associated with autoimmune diseases in Spanish population. Hum Immunol 2007; 68: 610–615.
Hunt KA, Monsuur AJ, McArdle WL et al: Lack of association of MYO9B genetic variants with coeliac disease in a British cohort. Gut 2006; 55: 969–972.
Amundsen SS, Monsuur AJ, Wapenaar MC et al: Association analysis of MYO9B gene polymorphisms with celiac disease in a Swedish/Norwegian cohort. Hum Immunol 2006; 67: 341–345.
Nunez C, Marquez A, Varade J et al: No evidence of association of the MYO9B polymorphisms with celiac disease in the Spanish population. Tissue Antigens 2006; 68: 489–492.
Giordano M, Marano C, Mellai M et al: A family-based study does not confirm the association of MYO9B with celiac disease in the Italian population. Genes Immun 2006; 7: 606–608.
Cirillo G, Di Domenico MR, Corsi I et al: Do MYO9B genetic variants predispose to coeliac disease? An association study in a cohort of South Italian children. Dig Liver Dis 2007; 39: 228–231.
Nunez C, Rueda B, Martinez A et al: A functional variant in the CD209 promoter is associated with DQ2-negative celiac disease in the Spanish population. World J Gastroenterol 2006; 12: 4397–4400.
Abel M, Cellier C, Kumar N, Cerf-Bensussan N, Schmitz J, Caillat-Zucman S : Adulthood-onset celiac disease is associated with intercellular adhesion molecule-1 (ICAM-1) gene polymorphism. Hum Immunol 2006; 67: 612–617.
Curley CR, Monsuur AJ, Wapenaar MC, Rioux JD, Wijmenga C : A functional candidate screen for coeliac disease genes. Eur J Hum Genet 2006; 14: 1215–1222.
Santin I, Castellanos-Rubio A, Perez de Nanclares G, Vitoria JC, Castano L, Bilbao JR : Association of KIR2DL5B gene with celiac disease supports the susceptibility locus on 19q13.4. Genes Immun 2007; 8: 171–176.
Ueda H, Howson JM, Esposito L et al: Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 2003; 423: 506–511.
Holopainen P, Arvas M, Sistonen P et al: CD28/CTLA4 gene region on chromosome 2q33 confers genetic susceptibility to celiac disease. A linkage and family-based association study. Tissue Antigens 1999; 53: 470–475.
Djilali-Saiah I, Schmitz J, Harfouch-Hammoud E, Mougenot JF, Bach JF, Caillat-Zucman S : CTLA-4 gene polymorphism is associated with predisposition to coeliac disease. Gut 1998; 43: 187–189.
Naluai AT, Nilsson S, Samuelsson L et al: The CTLA4/CD28 gene region on chromosome 2q33 confers susceptibility to celiac disease in a way possibly distinct from that of type 1 diabetes and other chronic inflammatory disorders. Tissue Antigens 2000; 56: 350–355.
Popat S, Hearle N, Hogberg L et al: Variation in the CTLA4/CD28 gene region confers an increased risk of coeliac disease. Ann Hum Genet 2002; 66: 125–137.
Popat S, Hearle N, Wixey J et al: Analysis of the CTLA4 gene in Swedish coeliac disease patients. Scand J Gastroenterol 2002; 37: 28–31.
King AL, Moodie SJ, Fraser JS et al: Coeliac disease: investigation of proposed causal variants in the CTLA4 gene region. Eur J Immunogenet 2003; 30: 427–432.
Mora B, Bonamico M, Indovina P et al: CTLA-4 +49 A/G dimorphism in Italian patients with celiac disease. Hum Immunol 2003; 64: 297–301.
Amundsen SS, Naluai AT, Ascher H et al: Genetic analysis of the CD28/CTLA4/ICOS (CELIAC3) region in coeliac disease. Tissue Antigens 2004; 64: 593–599.
Haimila K, Smedberg T, Mustalahti K, Maki M, Partanen J, Holopainen P : Genetic association of coeliac disease susceptibility to polymorphisms in the ICOS gene on chromosome 2q33. Genes Immun 2004; 5: 85–92.
van Belzen MJ, Mulder CJ, Zhernakova A, Pearson PL, Houwen RH, Wijmenga C : CTLA4 +49 A/G and CT60 polymorphisms in Dutch coeliac disease patients. Eur J Hum Genet 2004; 12: 782–785.
Zhernakova A, Eerligh P, Barrera P et al: CTLA4 is differentially associated with autoimmune diseases in the Dutch population. Hum Genet 2005; 118: 58–66.
Brophy K, Ryan AW, Thornton JM et al: Haplotypes in the CTLA4 region are associated with coeliac disease in the Irish population. Genes Immun 2006; 7: 19–26.
Hunt KA, McGovern DP, Kumar PJ et al: A common CTLA4 haplotype associated with coeliac disease. Eur J Hum Genet 2005; 13: 440–444.
Clot F, Fulchignoni-Lataud MC, Renoux C et al: Linkage and association study of the CTLA-4 region in coeliac disease for Italian and Tunisian populations. Tissue Antigens 1999; 54: 527–530.
Martin-Pagola A, Perez de Nanclares G, Vitoria JC et al: No association of CTLA4 gene with celiac disease in the Basque population. J Pediatr Gastroenterol Nutr 2003; 37: 142–145.
Rueda B, Zhernakova A, Lopez-Nevot MA et al: CTLA4/CT60 polymorphism is not relevant in susceptibility to autoimmune inflammatory intestinal disorders. Hum Immunol 2005; 66: 321–325.
Nunez C, Rueda B, Martinez A et al: Involvement of macrophage migration inhibitory factor gene in celiac disease susceptibility. Genes Immun 2007; 8: 168–170.
Rueda B, Zhernakova A, Lopez-Nevot MA, Martin J, Koeleman BP : Association study of functional genetic variants of innate immunity related genes in celiac disease. BMC Med Genet 2005; 6: 29.
Zhernakova A, Alizadeh BZ, Eerligh P et al: Genetic variants of RANTES are associated with serum RANTES level and protection for type 1 diabetes. Genes Immun 2006; 7: 544–549.
Rueda B, Koeleman BP, Lopez-Nevot MA et al: Poly (ADP-ribose) polymerase-1 haplotypes are associated with coeliac disease. Int J Immunogenet 2005; 32: 245–248.
Boniotto M, Braida L, Baldas V et al: Evidence of a correlation between mannose binding lectin and celiac disease: a model for other autoimmune diseases. J Mol Med 2005; 83: 308–315.
Wu J, Alizadeh BZ, Veen TV, Meijer JW, Mulder CJ, Pena AS : Association of FAS (TNFRSF6)-670 gene polymorphism with villous atrophy in coeliac disease. World J Gastroenterol 2004; 10: 717–720.
Rueda B, Martinez A, Lopez-Nevot MA et al: A functional variant of IFNgamma gene is associated with coeliac disease. Genes Immun 2004; 5: 517–519.
Lio D, Scola L, Forte GI et al: TNFalpha, IFNgamma and IL-10 gene polymorphisms in a sample of Sicilian patients with coeliac disease. Dig Liver Dis 2005; 37: 756–760.
Wapenaar MC, van Belzen MJ, Fransen JH et al: The interferon gamma gene in celiac disease: augmented expression correlates with tissue damage but no evidence for genetic susceptibility. J Autoimmun 2004; 23: 183–190.
Mora B, Bonamico M, Ferri M et al: Association of the matrix metalloproteinase-3 (MMP-3) promoter polymorphism with celiac disease in male subjects. Hum Immunol 2005; 66: 716–720.
Alizadeh BZ, Valdigem G, Coenen MJ et al: Association analysis of functional variants of the FcgRIIa and FcgRIIIa genes with type 1 diabetes, celiac disease and rheumatoid arthritis. Hum Mol Genet 2007; e-pub ahead of print 25 July 2007.
Thompson D, Stram D, Goldgar D, Witte JS : Haplotype tagging single nucleotide polymorphisms and association studies. Hum Hered 2003; 56: 48–55.
Reich DE, Cargill M, Bolk S et al: Linkage disequilibrium in the human genome. Nature 2001; 411: 199–204.
Reich DE, Lander ES : On the allelic spectrum of human disease. Trends Genet 2001; 17: 502–510.
Kong A, Cox NJ : Allele-sharing models: LOD scores and accurate linkage tests. Am J Hum Genet 1997; 61: 1179–1188.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Naluai, Å., Ascher, H., Nilsson, S. et al. Searching for genes influencing a complex disease: the case of coeliac disease. Eur J Hum Genet 16, 542–553 (2008). https://doi.org/10.1038/sj.ejhg.5201918
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201918