Abstract
Pinus chiapensis (Pinaceae) is a large conifer, endemic to central and southern Mexico and north-western Guatemala. In order to assess the extent of genetic variation within and between populations of this species, samples were obtained from throughout the natural range and analysed using random amplified polymorphic DNA (RAPD) and mtDNA RFLPs markers. Probes for the CoxI mitochondrial gene enabled two mitotypes to be observed. Populations from the eastern and western limit of the range of the species were fixed for one mitotype (‘A’), whereas two populations distributed near the centre of the range were fixed for another (‘B’). When the samples were screened with eight 10-mer RAPD primers, a total of 12 polymorphic bands were detected. The proportion of polymorphic bands was unusually low (24.5%) compared with other tree species. AMOVA analysis indicated that a significant proportion of the variation (P < 0.002) was distributed between populations; the extent of population differentiation detected (Φst = 0.226; GST = 0.194) was exceptionally high for a pine species. Pair-wise comparison of Φst values derived from AMOVA indicated that populations were significantly (P < 0.05) different from each other in virtually every case. These results are interpreted in the context of the evolutionary history of the species, and the implications for its in- and ex situ conservation are discussed.
Similar content being viewed by others
Introduction
Pines (Pinus spp., Pinaceae) are considered to be the most important genus of trees, both from ecological and economic perspectives (Richardson and Rundel, 1998). Many species of pine are of high economic value as sources of timber, pulp, nuts, resin and a variety of other products (Richardson and Rundel, 1998). Ecologically, pines are an often dominant component of the forests of the northern hemisphere, providing an essential source of food and habitat for a wide variety of animal and bird species, and playing a crucial role in biogeochemical processes (Richardson and Rundel, 1998). However, a third of all pine species are now considered to be threatened with extinction, as a result of logging, browsing and clearance for agriculture, among other factors (Farjon and Page, 1999).
Genetically, pines are among the most variable of organisms, and are characterised by high variation both within and between populations, as indicated by assessments of both quantitative (Cornelius, 1994) and isozyme variation (Hamrick and Godt, 1990; Ledig, 1998). The high diversity generally recorded within populations of pines is attributable to reproductive characteristics such as wind pollination, high reproductive capacity, effective mechanisms of seed dispersal, and flexible mating systems which permit inbreeding and selfing in isolated trees (Ledig, 1998; Richardson and Rundel, 1998). Pines are monoecious, and in most species of the genus, mechanisms of self-incompatibility appear to be lacking (Ledig, 1998). However, rates of outcrossing are generally very high, which appear to be maintained by partial self-sterility (Ledig, 1998).
Although patterns of genetic variation have been studied in many species of pine, most previous investigations have focused on northern temperate species (Hamrick and Godt, 1990; Ledig, 1998). The pine species of Mexico and Central America have been relatively little studied, despite the fact that this area has the greatest pine diversity of any area of comparable size, with some 47 native species, 74% of which are endemic to the region (Perry et al, 1998).
Pinus chiapensis (Mart.) Andresen is a white pine restricted to southern Mexico and neighbouring areas of Guatemala, occurring from latitudes 15°N and 19°N at altitudes of 150–2300 m (Dvorak et al, 1996). The tree can reach an impressively large stature, with a height of over 50 m with a diameter at breast height (d.b.h.) of over 1 m, and is found both on fertile and infertile soils in tropical and subtropical environments, generally on humid sites with an annual rainfall of 1500–2000 mm (Dvorak et al, 1996). Pinus chiapensis is morphologically similar to the well-known P. strobus L., which grows in southward from eastern Canada along the Appalachians as far as northern Georgia. Because of this similarity, P. chiapensis has attracted much taxonomic and phytogeographic interest (Wright et al, 1996). The species was originally classified as a variety of P. strobus, but subsequent taxonomic study has indicated that the taxon deserves specific rank (Price et al, 1998), a position which has been supported by recent phylogenetic analyses (Liston et al, 1999).
As with many other pine species in Mexico (Perry et al, 1998), most populations of P. chiapensis have been heavily exploited for timber. The tree is harvested by at least 11 native ethnic groups in southern Mexico, who use the wood for construction, furniture manufacture and firewood; resin from the tree is also used for medicinal purposes (del Castillo and Acosta, 2002). Many remaining stands are under immediate threat of extinction (Zamora-Serrano and Velasco-Fiscal, 1977; Dvorak and Brouard, 1987). The species is now largely restricted to small, fragmented and often degraded stands, usually surrounded by agricultural areas. Natural regeneration is severely reduced in many stands by the effects of fire and the activity of grazing animals (Zamora-Serrano and Velasco-Fiscal, 1977; Dvorak and Brouard, 1987; Richardson and Rundel, 1998). Concern about the threats to this species has led to the establishment of germplasm banks in countries such as Brazil, Colombia and South Africa, most notably by the CAMCORE Cooperative, North Carolina State University (Dvorak and Brouard, 1987;Dvorak et al, 1996, Wright et al, 1996).
This study was designed to assess the pattern of genetic variation within and between populations of P. chiapensis from throughout the natural range of the species. This was achieved through the use of random amplified polymorphic DNA (RAPD) and mtDNA restriction fragment length polymorphism (RFLP) markers. These techniques have increasingly been applied to the study of tree species in recent years, to provide information in support of conservation planning and management (Newton et al, 1999).
Materials and methods
Sample collection
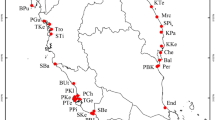
Range-wide seed collections of P. chiapensis were made in Mexico and Guatemala between 1983–1986 as part of the genetic conservation activities undertaken by the CAMCORE cooperative (Dvorak and Brouard, 1987; Dvorak and Donahue, 1992). Seeds were collected from individual mother trees, with a minimum of 100 m between selected trees within each population, and kept separate. After collection, seeds were air-dried and stored at 4°C. Ten populations from this collection were included in the present analyses (A–J, Table 1, Figure 1). Samples were thereby obtained from throughout the entire distributional range of the species. An additional two populations were sampled (by RF del Castillo) during 1997–1998 in Mexico (K–L, Table 1), using the same approaches (Figure 1). Overall, the altitude of the collection sites ranged from 470 to 2025 m, and annual rainfall varied between approximately 1300 to 2500 mm.
Map illustrating the position of Pinus chiapensis populations sampled for assessment of genetic variation. (For details of sample locations, see Table 1.)
DNA isolation
DNA was isolated from single embryos for analysis, and was therefore generally of very low yield. Although sufficient quantities of DNA were obtained from individual seeds for RAPD analyses, mtDNA RFLPs required larger amounts of DNA than these methods, and therefore collections of seed from individual mother trees were pooled (20–50 seeds per pooled sample). Because mtDNA is maternally inherited in Pinus, this provided a combined sample of maternal mtDNA for each family. By using this method sufficiently large quantities (approximately 0.2 mg) of DNA were obtained.
DNA was isolated using the CTAB extraction method adapted from Doyle and Doyle (1990). Excised seed embryos were ground with a mortar and pestle in a 1.5 ml eppendorf tube under liquid nitrogen to a fine powder. After allowing liquid nitrogen to evaporate, 0.5 ml CTAB isolation buffer was added (2% CTAB (cetyltrimethylammonium bromide), 1.4 M NaCl, 20 mM EDTA, 1% PEG 8000, 100 mM Tris-HCl (pH 9.5)) and incubated at 55°C for 1 h. The mixture was treated twice by shaking with 0.5 ml chloroform/isoamyl alcohol (24:1) followed by centrifugation to separate phases and removal of the aqueous layer. A 0.5 ml of isopropanol was added to the final aqueous extract and mixed gently to precipitate the DNA. The precipitate was then centrifuged to pellet DNA and washed in 70% ethanol. Finally the DNA was dissolved in 100 μl TE buffer (10 mM tris base, 1 mM EDTA, adjusted to pH 8.0 with HCl) and stored at 4°C. DNA concentration was determined by comparison to standards on agarose gels and dilutions made in TE buffer to give 25 ng DNA per μl for RAPD reactions. For mtDNA studies, the method was scaled up by a factor of 10 using 20–50 seeds and DNA was further purified using the Hybaid Recovery kit (Hybaid, UK) to facilitate digestion by restriction enzymes. Checks of DNA quality and replicability of the results indicated that the DNA was of satisfactory quality for analysis, despite the age of the source material.
mtDNA RFLPs
DNA for 6–8 samples from selected populations (Figure 2) for which sufficient material was available were digested with single restriction enzymes SspI, HaeIII and AluI and products were transfered to nylon membranes by Southern blot as described in Sambrook et al(1989). Blots were hybridised to 32P αCTP random prime labelled mitochondrial fragments of CoxI (Sinclair et al, 1998) and nad4/3-4 at 65°C. Following hybridisation, membranes were washed twice in 0.1X SSC, 0.05% SDS, at 65°C, blotted dry and exposed to X-ray film. Both probes were amplified from P. chiapensis DNA using the following PCR protocol, all reactions were performed in Perker–Elmer GeneAmp 9700 Thermal Cycler, using 10 μl reaction volume: approx. Twenty-five ng template DNA, 5 pmol each primer, 0.5 U Taq polymerase (Bioline UK), 100 mM each dNTP (Sigma), 1.5 mM MgCl2 with 1× Taq buffer (16 mM (NH4)2SO4, 67 mM Tris-HC1 (pH 8.8), 0.01% Tween 20) 35 polymerase chain reaction (PCR) cycles, consisting of 1 min denaturation at 94°C, 2 min primer annealing at 58°C, and 2 min extension at 72°C were used. Samples were maintained at 4°C after the termination of the programme.
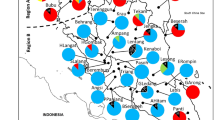
Map illustrating the distribution of two mitotypes of Pinus chiapensis detected by RFLP analysis. Populations represented by filled circles were fixed for mitotype A, populations represented by empty circles were fixed for mitotype B (for definition of mitotypes, see text). The populations represented by cross-hatched circles contained both mitotypes, but in differing proportions: the frequency of mitotype B was 0.125 in population H, and 0.875 in population L.
RAPD reactions
RAPD analyses were undertaken on individual seeds from separate mother trees. The number of individuals included in the analysis differed between populations, ranging from five to 35 (with an overall mean of 13.8), reflecting variation in the size of the population and the number of seeds available (Table 1). The total number of individuals screened was 138.
The PCR constituent concentrations and conditions were optimised for representative samples of P. chiapensis to give repeatable markers. All reactions were performed in Perker–Elmer GeneAmp 9700 Thermal Cycler, using 10 μl reaction volume. Optimum reactions contained: approx. 25 ng template DNA, 5 pmol primer, 0.5 U Taq polymerase (Bioline, UK), 100 mM each dNTP (Sigma, UK), 1.5 mM MgCl2 with 1× Taq buffer (16 mM (NH4)2SO4, 67 mM Tris-HC1 (pH 8.8), 0.01% Tween 20). Forty-five PCR cycles, consisting of 1 min denaturation at 94°C, 1 min primer annealing at 40°C, and 2 min extension at 72°C were used. Samples were maintained at 4°C after the termination of the programme. Primer kits OPK and OPAL, a total of 40 (Operon Technologies, Alameda, CA, USA), were screened with a subset of P. chiapensis DNA samples for production of clear RAPD profiles. RAPD products were separated on 6% polyacrylamide gels (15:1 acrylamide:bis-acrylamide), stained with 10 μg/ml ethidium bromide and photographed over UV light.
Only RAPD bands that could be unequivocally scored were counted in the analysis. Generally, only bands in size ranges between clearly visible monomorphic bands were scored. This greatly reduced the potential number of bands that could have been scored but, importantly, avoided mis-scoring of bands for which co-migration could not be certain.
Data analysis
The mating system of P. chiapensis in the populations studied is not known and therefore calculation of allele frequencies by compensating for deviation from Hardy–Weinburg equilibrium was not attempted as has been described elsewhere (Lynch and Milligan, 1994). In the absence of more detailed characterisation of RAPD bands, their use as genetic phenotypes would appear prudent. Genetic distance methods were therefore used. In the case of RAPD data, amplified DNA marker bands were scored in a binary manner as either present (1) or absent (0) and entered into a binary data matrix. The shared absence of a band was not scored as a shared character. A simple pair-wise distance measure (Jaccard’s) was calculated from the binary RAPD data using the following formula (Sneath and Sokal, 1973):

where S is the total number of shared present band positions and T is the total number of band positions between ith and jth individuals. It is important to note that T is the number of positions that for i and j at least one band is present, therefore a shared absence is not scored as a possible band position. This reduces overestimation of distance where individuals may have fewer RAPD bands owing to artifactual PCR failure.
An analysis of molecular variance (AMOVA) was conducted using the Jaccard’s distance measure to examine differences between and within populations (Excoffier et al, 1992). The AMOVA programme generates Φ statistics (Excoffier et al, 1992), which are analogous to Wright’s FST (Wright, 1951). This approach has been widely adopted in the analysis of RAPD data (eg, Gillies et al, 1997, 1999; Allnutt et al, 1999; Nybom and Bartish, 2000; Ehtisham-ul-Haq et al, 2001; Bekessy et al, 2002), allowing comparison of results with a variety of other studies. Comparison of between-population diversity estimated with AMOVA-derived ΦST and with Nei’s GST, in a wide range of RAPD studies, indicated that results from the two approaches are highly correlated, and often almost identical (Nybom and Bartish, 2000). Pair-wise ΦST values were used to construct a UPGMA dendrogram using PHYLIP (Phylogeny Inference Package Version 3.57c,Felsenstein, 1993), to illustrate statistical similarity between the RAPD profiles of different populations. Jaccard distances were also used to perform a principal co-ordinate analysis (PCO), which provides a graphical representation of the similarity between individuals based on RAPD phenotypes.
Shannon’s diversity estimates (Lewontin, 1972) were calculated to provide a relative estimate of the degree of variation within each population by using the formula:

where pi is the frequency of presence or absence of each RAPD band (treating each RAPD band as a single locus according to the approach of Lewontin (1972) for isozymes). Note that we here denote Shannon’s measure as ‘S’, not ‘H’ so as to avoid confusion with other measures of diversity such as heterozygosity, to which Shannon’s is not directly comparable. Following conventions for heterozygosity we denoted Shannon’s estimate across the total sample as St, and for each (sub)population as Ss.
Results
mtDNA RFLPs
Mitochondrial DNA probes amplified from P. chiapensis DNA were of the expected size originally reported (Sinclair et al, 1998). Blots probed with nad4/3-4 were monomorphic for all populations. For CoxI, HaeIII and AluI, digests were monomorphic across all samples whereas in SspI digests, two mitotypes were observed, ‘A’ consisting of a single 3500 bp fragment and ‘B’ consisting of a single 2800 bp fragment. These two mitotypes showed geographically distinct patterns of distribution (Figure 2). Populations from Chiapas and Guatemala (A, B, D) at the eastern end of the range of the species, were all fixed for mitotype A, as were those at the northwestern limit of those populations sampled (F, G, K). Two geographically intermediate populations, E and J, were fixed for mitotype B. The other two populations contained both mitotypes, but in differing proportions. Population H had a low frequency of mitotype B (0.125) and population L displayed a high frequency (0.875) of mitotype B (Figure 2).
RAPDs
From a set of 25 primers screened, eight produced clear RAPD patterns consisting of a total of 49 scorable markers, 12 of which were polymorphic (100% criterion) (Table 2). Screening of the entire set of samples was repeated with two of the primers to assess repeatability of RAPD profiles, and identical RAPD patterns were obtained. It was observed that the proportion of monomorphic bands was unusually high (low % polymorphism) compared with other RAPD studies using comparable methods (Table 3). No population specific bands were observed.
Calculated pair-wise distances were used as input for principal co-ordinate and AMOVA analyses. The first two principal co-ordinates of RAPD distances described 7.26 and 5.79% of the total variance respectively. When the two principal coordinates explaining the most variation were considered, individuals plotted as a broad scatter, with no population clustering independently from all of the others (Figure 3). However, samples from individual populations displayed a tendency to cluster together, as illustrated by the PCO (Figure 3). For example, samples from population E overlapped only with individuals from populations A and D. Similarly, samples from population J overlapped only with those from populations F, G, H, and I. The difference between populations E and J as indicated by this analysis is surprising, given their relative geographic proximity.
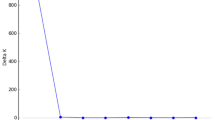
Plot of the first two principal coordinates of pair-wise distances derived from RAPD analysis of Pinus chiapensis populations (describing 7.26% and 5.79% of the variation respectively). Letters A–J refer to populations (see Table 1 for details). Populations E and J are outlined.
Examination of the RAPD results by AMOVA (Table 4) indicated that although most of the variation (77.4%) was found within populations, a significant proportion (P < 0.002, tested using a 500 replication bootstrap) was attributable to differences between populations. The estimate of ΦST was also similar to values of GST obtained from Shannon’s diversity index (0.219).
Pair-wise ΦST values derived from AMOVA highlighted a large number of significant (P < 0.05) differences between populations, when individual pairs of populations were compared (Table 5). Populations were significantly different from each other in every case (P < 0.05), with the exception of population C when compared to populations A, E, F, H; and populations D and E were also not significantly different from each other (Table 5). These non-significant differences may be attributed to the relatively low sample numbers for populations C, D and E.
When Φst values were used to construct a UPGMA dendrogram in order to examine relationships between populations (Figure 4), two main groups were apparent: the first including populations A, C, D and E, the second including populations G, H, I and J. Although populations B and F occupied an intermediate position between these two groups, both were significantly different from each other. These two groupings relate closely to their geographical distribution. Populations A (Guatemala), B, C and D (Chiapas) all lie towards the eastern end of the range of the species, whereas the other populations all lie further to the north and west (Oaxaca, Veracruz). The two most geographically distant populations, A and I, were also among the most distant genetically, according to the UPGMA. However, although population E grouped with those populations from Chiapas and Guatemala on the UPGMA analysis, geographically it is closest to population J, lying as it does in Oaxaca. Diversity estimates indicated by Shannon’s index differed significantly between populations, values ranging from 0.441 (J) to 0.755 (F) (overall mean, St, 0.544) (Table 6).
UPGMA diagram constructed using pair-wise ΦST values derived by RAPD analysis for populations of Pinus chiapensis. Populations were significantly different from each other in every case (P < 0.05), with the exception of population C and populations A, E, F, H, and populations D and E were not significantly different from each other (Table 5). For details of populations sampled, see Table 1.
Discussion
Previous analyses of mtDNA variation in pine species have provided valuable insights into recent evolutionary history. For example, two common mitotypes were detected in RFLP analysis of mtDNA in Scottish populations of P. sylvestris (Sinclair et al, 1998). Although one mitotype was present in all sites, the second was confined to three western populations, which was interpreted as indicating a separate origin for these populations during post-glacial migration (Sinclair et al, 1998). The current results highlight a similar degree of variation in P. chiapensis, with two mitotypes detected.
Mitotype A was the most widespread, occurring in eight out of the 10 populations of P. chiapensis sampled. In contrast, mitotype B was restricted to a narrower geographical region towards the centre of the species’ range. The two populations fixed for mitotype B, J and E, both lie in the mountain range to the north of the state of Oaxaca, the Sierra Norte. Populations H and L, which contained both mitotypes, are geographically isolated from these mountains by relatively arid regions; as P. chiapensis is a moisture-demanding species (Dvorak and Brouard, 1987), such aridity acts as a significant barrier to dispersal and colonisation. Population L (Chimalapas) is of particular geographic interest, as it is an area of relatively low elevation lying between the mountain ranges of Oaxaca (to the west) and Chiapas (to the east). This population is therefore geographically isolated at the current time, but according to these results, displays genetic affinities to neighbouring populations both to the east and west. These results are consistent with the hypothesis that populations of P. chiapensis were more contiguous in the past and have since been fragmented as a result of human impact and climate change.
A somewhat contrasting pattern of genetic variation was recorded by the RAPD analyses. Here, populations towards the southern and eastern limit of the range of the species (Chiapas and Guatemala) tended to cluster together. Some other geographically proximate populations also displayed a degree of genetic affinity, such as F and G. However, the neighbouring populations J and E, which were the only two populations fixed for the rarer mitotype B, were highly differentiated in the RAPD analyses. Results derived from RAPD and mtDNA analyses should be compared with caution, as they sample different parts of the genome. In addition, the fact that seeds were to characterise the genetic variation of populations should also be considered in the interpretation of results. Whereas mtDNA analysis of seeds provides information relating solely to the the population of seed trees, this is only partially true for the RAPD markers, where long-distance pollen flow may have influenced the patterns detected. Ideally, the haploid endosperm would have been used to examine the inheritance of RAPD bands, enabling such processes to be evaluated, but sample sizes were too small to permit this.
The main feature of the RAPD results was the high degree of differentiation recorded between populations. The estimate of ΦST derived here for P. chiapensis (0.226) is substantially higher than values derived for other tree species using similar methods (Isabel et al, 1995; Allnutt et al, 1999, 2001; Gillies et al, 1999; Bekessy et al, 2002). This value is also high compared to values for other pine species using a variety of different markers. In the 66 studies listed by Ledig (1998), only four recorded values of GST greater than 0.2 (20%), namely P. ayacahuite, P. cembra, P. muricata and P. torreyana. Of these, the populations of P. cembra were specifically chosen because they were isolated and widely separated, and P. torreyana has only two extant populations. From a review of 195 isozyme studies of long-lived perennial woody taxa, Hamrick et al (1992) recorded an overall mean GST value of 8.4%. However, comparison of the estimates of population differentiation recorded here with these results from other studies should be made with caution, particularly those that were based on the use of isozyme markers. RAPD analyses of tree species have often detected a higher degree of population differentiation than isozyme analyses (Newton et al, 1999), which may be attributed to a contribution of maternally inherited organelle DNA to the RAPD phenotypes observed (Aagard et al, 1995), or artifacts of statistical analysis (Isabel et al, 1995).
Such a high degree of genetic differentiation implies that populations are reproductively isolated, or have been isolated in the past, resulting in differentiation by inbreeding and genetic drift, or through local adaptation. Populations of P. chiapensis are highly fragmented, both as a result of both human impact and the discontinuous distribution of suitable habitat. As noted earlier, the species is restricted to relatively humid sites, and it is therefore conceivable that low-lying arid areas, together with topographic features such as the many mountain ranges in the region, might constitute significant barriers to gene flow, and may thereby have promoted genetic differentiation between populations of this species. Furthermore, distinct differences in flowering times between populations and trees within populations have been noted for P. chiapensis when planted as an exotic species in South Africa (Dvorak, personal observation). Such differences may also exist to some degree in natural populations and limit gene flow from one area to another.
Historical factors may also have had a major influence on genetic structure of P. chiapensis. Limited information is available concerning forest history in Mexico during the Holocene, which is considered to have begun around 8500–9000 year BP (Heine and Ohngemach, 1976). The changes in climate associated with glaciation events are likely to have resulted in the southern migration of pine taxa along mountain ranges such as the Sierra Madre Occidental (Perry et al, 1998). Postglacial increases in temperature were associated with northward migration of some Mexican conifers, and increased population fragmentation. For example, during the last glaciation Chihuahuan spruce (Picea chihuahuana) is known to have occurred at least 700 km further south than it does today. Analysis of isozyme variation within this species detected an exceptionally high degree of population differentiation (FST: 24.8%), which was attributed to the effects of genetic drift occurring in populations which became increasingly isolated and fragmented as a result of climatic warming during the postglacial period (Ledig et al, 1997).
The RAPD data presented here provide evidence that similar processes may have occurred with Pinus chiapensis. The percentage polymorphism of RAPD loci was substantially lower than in any of the other tree species for which comparable data are available (Chalmers et al, 1992; Isabel et al, 1995; Yeh et al, 1995; Gillies et al, 1997, 1999; Allnutt et al, 1999, 2001; Bekessy et al, 2002; see Table 3). Low genetic diversity has been recorded in other Mexican pines: for exanmple, isozyme analysis of P. maximartinezii indicated very low percentage polymorphism (30.3%), which was interpreted as evidence of a population bottleneck, probably within the last 1000 years (Ledig et al, 1997).
The results of this study have a number of implications for both the in situ and ex situ conservation of P. chiapensis. The high degree of differentiation recorded between populations suggests that populations throughout the range of the species should be incorporated into both in situ and ex situ conservation plans. This conclusion is supported by the detection of variation in growth traits between provenances of this species (Dvorak et al, 1996; Wright et al, 1996). Conservation decisions should be based on quantitative genetic variation as well as patterns of variation in molecular markers, which are generally assumed to be selectively neutral. Collection of additional provenances and their maintenance in cultivation could make a significant contribution to conserving the genetic base of this species; in particular, populations from the state of Guerrero are not well represented in existing germplasm collections (Dvorak et al, 1996).
These data could also be used to define appropriate units for in situ conservation (Newton et al, 1999). For example, on the basis of mtDNA variation, the populations in the eastern part of the Sierra Norte in Oaxaca (populations E and J) might usefully be considered separately from neighbouring populations to the west; the difference in mitotypes recorded is likely to represent an ancient evolutionary event (Sinclair et al, 1998), which it might be desirable to recognize in conservation planning (Newton et al, 1999). However, the RAPD analyses highlighted significant genetic differences between many of the populations sampled, and therefore any approach aiming to conserve the full breadth of variation within the species should aim to incorporate as many populations as possible. The same conclusion has been reached by the CAMCORE in its testing and ex situ conservation programmes.
To date, no integrated plan for the conservation of P. chiapensis has been developed in southern Mexico and northwestern Guatemala, although a logging ban is now in force, at least in some areas. As noted by Wright et al (1996), without intervention by forest authorities in Mexico and Guatemala, much of the gene pool of this species will disappear; protection of remaining forest is urgently required to maintain it as a viable resource.
References
Aagaard, JE, Vollmer, SS, Sorensen, FC, Strauss, SH (1995). Mitochondrial DNA products among RAPD profiles are frequent and strongly differentiated between races of Douglas-fir. Mol Ecol, 4: 441–447.
Allnutt, TR, Newton, AC, Lara, A, Premoli, A, Armesto, JJ, Vergara, R et al. (1999). Genetic variation in Fitzroya cupressoides (alerce), a threatened South American conifer. Mol Ecol, 8: 975–987.
Allnutt, TR, Courtis, JR, Gardner, M, Newton, AC (2001). Genetic variation in wild Chilean and cultivated British populations of Podocarpus salignus D. Don (Podocarpaceae). Edinb J Bot, 58: 459–473.
Allnutt, TR, Newton, AC, Premoli, A, Lara, A (2002). Genetic variation in the threatened conifer, Pilgerodendron uviferum. Biol Conserv. In press.
Bekessy, SA, Allnutt, TR, Premoli, AC, Lara, A, Ennos, RA, Burgman, MA et al. (2002). Genetic variation in the vulnerable and endemic Monkey Puzzle tree, detected using RAPD. Heredity, 88: 243–249.
Chalmers, KJ, Waugh, R, Sprent, JI, Simons, AJ, Powell, W (1992). Detection of genetic variation between and within populations of Gliricidia sepium and G. maculata using RAPD markers. Heredity, 69: 465–472.
Cornelius, J (1994). Heritabilities and additive genetic coefficients of variation in forest trees. Can J For Res, 24: 372–379.
Del Castillo, RF, Acosta, S (2001). Ethnobotanical notes on Pinus strobus var. chiapensis. Boletin Sociedad Botánica de Mexico. (In press).
Doyle, JJ, Doyle, JL (1990). Isolation of plant DNA from fresh tissue. Focus, 12: 13–15.
Dvorak, WS, Brouard, J (1987). An evaluation of Pinus chiapensis as a commercial plantation species for the tropics and subtropics. Comm For Rev, 66: 165–176.
Dvorak, WS, Donahue, JK (1992). CAMCORE Cooperative Research Review 1980-1992. College of Forest Resources, North Carolina State University, USA, 93pp.
Dvorak, WS, Donahue, JK, Vasquez, JA (1996). Provenance and progeny results for the tropical white pine, Pinus chiapensis, at five and eight years of age. New For, 12: 125–140.
Excoffier, L, Smouse, PE, Quattro, JM (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics, 131: 479–491.
Ehtisham-Ul-Haq, M, Allnutt, TR, Smoth-Ramirez, C, Gardner, M, Armesto, JJ, Newton, AC (2001). Patterns of genetic variation in in- and ex-situ populations of the threatened vine Berberidopsis corallina, detected using RAPD markers. Ann Bot, 87: 813–821.
Farjon, A, Page, CN (1999). Conifers – status survey and conservation action plan. IUCN/SSC Conifer Specialist Group: Cambridge, UK.
Felsenstein, J (1993). PHYLIP (Phylogeny Inference Package). Department of Genetics, University of Washington, Seattle.
Gillies, ACM, Cornelius, JP, Newton, AC, Navarro, C, Hernández, M, Wilson, J (1997). Genetic variation in Costa Rican populations of the tropical timber tree Cedrela odorata L. (Spanish cedar), assessed using RAPDs. Mol Ecol, 6: 1133–1145.
Gillies, ACM, Navarro, C, Lowe, AJ, Newton, AC, Hernández, M, Wilson, J et al. (1999). Genetic diversity in Mesoamerican populations of mahogany (Swietenia macrophylla), assessed using RAPDs. Heredity, 83: 722–732.
Hamrick, JL, Godt, MJW (1990). Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant Population Genetics, Breeding and Genetic Resources, Sinauer Associates: Sunderland, Mass pp 43–63.
Hamrick, JL, Godt, MJW, Sherman-Broyles, SL (1992). Factors influencing levels of genetic diversity in woody plant species. New For, 6: 95–124.
Heine, K, Ohngemach, D (1976). Die Pleistozän/Holozän-Grenze in Mexiko. Münstersche Forschungen zur Geologie und Paläonontologie, 38/39: 229–251.
Isabel, N, Beaulieu, J, Bousquet, J (1995). Complete congruence between gene diversity estimates derived from genotypic data at enzyme and random amplified polymorphic DNA loci in black spruce. Proc Nat Acad Sci USA, 92: 6369–6373.
Ledig, FT (1998). Genetic variation in Pinus. In: Richardson DM (ed) Ecology and Biogeography of Pinus. Cambridge University Press: Cambridge. pp 251–280.
Ledig, FT, Conkle, MT, Bermejo-Velázquez, B, Eguiluz-Piedra, T, Hodgskiss, PD, Johnson, DR et al. (1999). Evidence for an extreme bottleneck in a rare Mexican endemic pinyon: genetic diversity, disequilibrium, and the mating system of Pinus maximartinezii. Evolution, 53: 91–99.
Ledig, FT, Jacob-Cervantes, V, Hodgskiss, PD, Eguilez-Piedra, T (1997). Recent evolution and divergence among populations of a rare Mexican endemic, Chihuahua spruce, following Holocene climatic warming. Evolution, 51: 1815–1827.
Lewontin, RC (1972). The apportionment of human diversity. Evol Biol, 6: 381–398.
Liston, A, Robinson, WA, Piñero, D, Alvarez-Buylla, ER (1999). Phylogenetics of Pinus (Pinaceae) based on nuclear ribosomal DNA internal transcribed spacer region sequences. Mol Phylogenet Evol, 11: 95–109.
Lynch, M, Milligan, BG (1994). Analysis of population genetic structure with RAPD markers. Mol Ecol, 3: 91–99.
Newton, AC, Allnutt, T, Gillies, ACM, Lowe, A, Ennos, RA (1999). Molecular phylogeography, intraspecific variation and the conservation of tree species. Trends Ecol Evol, 14: 140–145.
Nybom, H, Bartish, IV (2000). Effects of life history traits and sampling strategies on genetic diversity estimates obtained with RAPD markers in plants. Perspect Plant Ecol Evol Systemat, 3/2: 93–114.
Perry, JP, Graham, A, Richardson, DM (1998). The history of pines in Mexico and Central America. In: Richardson DM (ed) Ecology and Biogeography of Pinus. Cambridge University Press: Cambridge pp 137–149.
Price, RA, Liston, AL, Strauss, SH (1998). Phylogeny and systematics of Pinus. In: Richardson DM (ed) Ecology and Biogeography of Pinus. Cambridge University Press: Cambridge pp 49–68.
Richardson, DM, Rundel, PW (1998). Ecology and biogeography of Pinus: an introduction. In: Richardson DM (ed) Ecology and Biogeography of Pinus. Cambridge University Press: Cambridge pp 3–46.
Sambrook, J, Fritsch, EF, Maniatis, T (1989). Molecular Cloning – a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY.
Sinclair, WT, Morman, JD, Ennos, RA (1998). Multiple origins for Scots pine (Pinus sylvestris L.) in Scotland: evidence from mitochondrial DNA variation. Heredity, 80: 233–240.
Sneath, PHA, Sokal, RR (1973). Numerical Taxonomy. W. H. Freeman: San Francisco.
Wright, S (1951). The genetical structure of populations. Annals Eugenics, 15: 323–354.
Wright, JA, Marin, AM, Dvorak, WS (1996). Conservation and use of the Pinus chiapensis genetic resource in Colombia. For Ecol Manag, 88: 283–288.
Yeh, FC, Chong, DKX, Yang, R-C (1995). RAPD variation within and among natural populations of trembling aspen (Populus tremuloides Michx.) from Alberta. J Hered, 86: 454–460.
Zamora-Serrano, C, Velasco-Fiscal, V (1977). Pinus strobus var. chiapensis, una especie en peligro de extincion en el estado de Chiapas. Ciencia Forestal, 2: 3–23.
Acknowledgements
The research was funded by the UK Darwin Initiative and formed part of the SUCRE and BIOCORES Projects funded by the European Commission under Framework IV of DGXII. We are very grateful to staff of CAMCORE and CIIDIR for assistance with sample collection, to Nahum Sanchez and Raul Rivera for help with seed collections, and to colleagues at ECOSUR, Mexico for helpful discussions. The original CAMCORE seed collections were made in collaboration with INIF, Mexico and INAFOR, Guatemala, and were part funded by the Agency for International Development (A.I.D.), USA, Grant no. DPE-5542-6-SS-4017-00. Additional seed were collected with the support of SIBEJ-CONACyT and IPN.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Newton, A., Allnutt, T., Dvorak, W. et al. Patterns of genetic variation in Pinus chiapensis, a threatened Mexican pine, detected by RAPD and mitochondrial DNA RFLP markers. Heredity 89, 191–198 (2002). https://doi.org/10.1038/sj.hdy.6800113
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800113
Keywords
This article is cited by
-
High genetic diversity but spatially restricted genetic connectivity in a tropical montane cloud forest tree (Magnolia schiedeana)
Tree Genetics & Genomes (2023)
-
Morpho-genetic divergence and population structure in Indian Santalum album L.
Trees (2020)
-
Large scale patterns of genetic variation and differentiation in sugar maple from tropical Central America to temperate North America
BMC Evolutionary Biology (2015)
-
Novel Chloroplast Microsatellite (cpSSR) Markers for Genetic Diversity Assessment of Cultivated and Wild Hevea Rubber
Plant Molecular Biology Reporter (2015)
-
A range wide geographic pattern of genetic diversity and population structure of Castanea mollissima populations inferred from nuclear and chloroplast microsatellites
Tree Genetics & Genomes (2013)