Abstract
Genetic recombination is usually considered to facilitate adaptive evolution. However, recombination prevents the reliable cotransmission of interacting gene combinations and can disrupt complexes of coadapted genes. If interactions between genes have important fitness effects, restricted recombination may lead to evolutionary responses that are different from those predicted from a purely additive model and could even aid adaptation. Theory and data have demonstrated that phenomena that limit the effectiveness of recombination via increasing homozygosity, such as inbreeding and population subdivision and bottlenecks, can temporarily increase the additive genetic variance available to these populations. This effect has been attributed to the conversion of nonadditive to additive genetic variance. Analogously, phenomena such as chromosomal inversions and apomictic parthenogenesis that physically restrict recombination in part or all of the genome may also result in a release of additive variance. Here, we review and synthesize literature concerning the evolutionary potential of populations with effectively or physically restricted recombination. Our goal is to emphasize the common theme of increased short-term access to additive genetic variance in all of these situations and to motivate research directed towards a more complete characterization of the relevance of the conversion of variance to the evolutionary process.
Similar content being viewed by others
Introduction
Genetic recombination is traditionally regarded as a mechanism that aids adaptive evolution. The link between recombination and adaptation can be traced to Weismann (1904), who proposed that the advantages of recombination lie in its ability to unite favorable alleles from separate lineages into one lineage more quickly than mutation alone (also see Fisher, 1930; Muller, 1932). Recombination also allows selection to operate more effectively on individual beneficial alleles segregating on a polymorphic genetic background (Hill and Robertson, 1966; Birky and Walsh, 1988). These lines of reasoning underlie the widely held assumption that recombination facilitates the evolutionary response to selection. Recombination, however, also breaks up complexes of interacting genes and disrupts favorable gene combinations (Maslin, 1968; White, 1973). Theory and empirical research suggest that these consequences of recombination can act as a constraint upon adaptive evolution. Here, we review and synthesize evidence that a phenomenon associated with limits on recombination, the conversion of nonadditive, epistatic genetic variance to additive genetic variance, may help explain why evolutionary responses can be different from predicted and short-term adaptive evolution can occur more efficiently than expected when recombination is restricted.
Introduction of concepts
Additive and nonadditive variance
The additive or average effect of an allele is defined as the expected deviation from the population mean genotypic value caused by a single copy of an allele (Fisher, 1918; Lynch and Walsh, 1998). The sum of the additive effects of alleles across all loci is the breeding value of an individual. This breeding value represents the expected genotypic value of offspring of the individual. Given random mating, the additive genetic variance for a trait is simply the variance of breeding values of individuals for that trait in a population (Falconer and Mackay, 1996; Lynch and Walsh, 1998).
The genotypic value of an individual, however, is not merely the sum of the additive effects of its genes, because each allele can also have nonadditive, interaction effects. These nonadditive effects include interactions with other alleles at the same locus (ie dominance), or interactions with alleles at other loci (ie epistasis) (Falconer and Mackay, 1996; Lynch and Walsh, 1998). The deviation of the genotypic value of two alleles at the same locus from the expected genotypic value based on the average effect of each allele is the dominance deviation. In a randomly mating population, the variance in dominance deviations across all loci is the dominance variance. Similarly, the deviation of the genotypic value of two alleles at different loci from the expected genotype is the interaction deviation. The variance of interaction deviations is the epistatic variance (Falconer and Mackay, 1996; Lynch and Walsh, 1998).
It is important to note that just because two alleles at the same locus display physiological dominance, meaning that they interact nonadditively such that one allele is dominant and the other recessive, does not mean that the population will exhibit any dominance variance. Similarly, just because alleles at different loci show physiological epistasis (interact nonadditively) does not mean that the population will exhibit epistatic variance (Cheverud and Routman, 1995; Wade et al, 2001). Allelic interaction will only contribute to genetic variance if there is allelic polymorphism at the interacting loci (Goodnight, 1988). Furthermore, deviations from additivity due to dominance and epistasis may occur at particular sets of loci, but dominance variance and epistatic variance are based on the sum of dominance and epistatic deviations across all loci (Phillips et al, 2000).
Sexually produced diploid offspring only inherit one allele per locus from each parent, and unrestricted recombination disrupts the coinheritance of interacting alleles at different loci. This means that neither dominance deviations nor interaction deviations are reliably transmitted from parents to offspring (Wright and Cockerham, 1985; Falconer and Mackay, 1996; Gibson, 1996; Wade, 2001). Rather, it is only the additive effects of alleles that are predictably inherited by sexually derived offspring. Accordingly, evolutionary responses to individual level selection are expected to occur only when there is additive genetic variance.
Just as single-locus genotypes can be described in terms of additive and dominance genotypic values, so can multiple-locus genotypes. For example, two-loci epistatic systems are often described in terms of additive-by-additive epistasis, additive-by-dominance epistasis, dominance-by-additive epistasis, and dominance-by-dominance epistasis. Epistatic systems involving more than two loci are described analogously as additive-by-additive-by-additive epistasis, etc. These terms are used because they fit well into the existing quantitative genetic theory (Crow and Kimura, 1970), though they ultimately remain statistical constructs. In reality, allelic and genic interactions are likely to include combinations of these types of epistasis (Cheverud, 2000; Carlborg and Haley, 2004). In this review, we will focus mainly on a simple system of additive-by-additive epistasis (Figure 1) because such a system is most conducive to the conversion of epistatic to additive variance, as we detail below. Furthermore, conversion dynamics in systems of additive-by-additive epistasis have been well-described analytically (eg Goodnight, 1987; Cheverud and Routman, 1995; Cheverud, 2000).
Fitness landscape showing pure additive-by-additive epistasis. The horizontal axis shows the allele frequencies at loci A and B, and the vertical axis shows the mean fitness of a population with those allele frequencies. Note that neither allele at either locus is unconditionally favored at all allele frequencies; the effect of alleles at the A and B loci depends on the frequency of alleles at the alternate locus. In the flat central region of the fitness surface, pA1≈pB1≈0.5, the fitness of all genotypes is approximately 0 and neither allele at either locus is favored, but as the allele frequencies tend away from this region, one allele becomes favored. For example, as pA1 tends towards 1.0 (right side of figure), locus B acts more additively, and B1 is strongly favored over B2. This corresponds to a conversion of epistatic to additive variance (after Whitlock et al, 1995).
Conversion of variance by restricted effective recombination
The additive effects and interaction deviations of alleles, and hence the additive and nonadditive genetic variance of populations, are dependent on the allele frequencies of populations. As allele frequencies change, so do additive effects, interaction deviations, and the amount of additive and nonadditive genetic variance. For example, as selection or drift causes the loss or fixation of alleles, heterozygosity decreases and levels of additive genetic variance are also expected to decrease. Changes in allele frequencies can also alter the magnitudes of dominance and epistatic deviations, consequently affecting the relative amounts of nonadditive and additive variance (Goodnight, 1988; Cheverud and Routman, 1995; Barton and Turelli, 2004). One example of this phenomenon is provided by dominance deviations, which decrease as recessive alleles become more common and the proportion of heterozygotes decrease. Loci that formerly had large dominance deviations then act more additively (Wang et al, 1998b). Similarly, when one or more of the epistatic partners of an allele in epistatic systems with additive-by-additive components approach loss or fixation, the interaction deviation decreases and the allele acts more additively (Figures 1 and 2; Goodnight, 1988). In general, the effect of an allele is more predictable (ie is more additive) when the genetic background with which it interacts is more nearly constant.
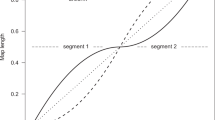
The additive and epistatic variance for an additive-by-additive epistatic system. In (a) and (b), the horizontal axes show the allele frequencies at loci A and B, and the vertical axis shows the additive (a) or epistatic (b) variance for those allele frequencies. (c) shows how epistatic (solid line) and additive (dotted line) variances change with allele frequencies when pA1=pB1. Note that epistatic variance is at a maximum and additive variance at a minimum when the allele frequencies pA1=pB1=0.5, which corresponds to the flat saddle region in Figure 1. As allele frequencies at loci A and B tend away from 0.5, the loci act more additively and epistatic variance is converted to additive variance. Additive variance for each locus is at a maximum when p=0.5 at one locus and p=0 or 1 at the alternate locus. That is, when there is no variance at one locus, the alternate locus acts completely additively (after Whitlock et al, 1995; Cheverud and Routman, 1996; Cheverud, 2000).
Changes in allele frequencies, and consequently, the amount of dominance and interaction deviation, can cause a ‘conversion’ of dominance or epistatic variance to additive variance (Goodnight, 1987, 1988; Whitlock et al, 1993; Figure 2). For example, whenever the ratio of additive-by-additive epistatic variance to additive variance in a panmictic ancestral population exceeds 1:3, the average within-deme amount of additive variance will increase for all values of FST, Wright's measure of population genetic subdivision, due to conversion (Goodnight, 1988). In this way, as homozygosity increases, the conversion of nonadditive to additive variance may slow the expected decline in additive variance.
Populations that have undergone inbreeding, population bottlenecks, or population subdivision will be increasingly homozygous and there may be substantial conversion of nonadditive to additive variance (Wade and Goodnight, 1998). The response to selection in these populations may be different from or greater than under purely additive expectations (Lynch, 1984b; Carson, 1990; Wade, 1996, 2001; Willis, 1996; Barton and Turelli, 2004). Mechanistically, the increased homozygosity caused by allelic loss and fixation decreases the effects of recombination, because recombination between homozygous loci does not increase genotypic diversity. Rather, it merely swaps identical alleles. Thus, the ability of recombination to inhibit the co-inheritance of sets of interacting alleles is diminished by inbreeding, population bottlenecks, and population subdivision (Wade, 1996), all of which increase homozygosity in the population.
While all forms of epistasis can increase the average additive genetic variance through conversion, the conversion of additive-by-additive variance results in a greater boost in additive variance than other types of epistasis (Cheverud and Routman, 1996; Naciri-Graven and Goudet, 2003). Moreover, unlike epistatic variance with dominance components, additive-by-additive epistatic variance cannot contribute to inbreeding depression (Falconer and Mackay, 1996). Thus, conversion of additive-by-additive variance seems more likely to facilitate adaptive evolution than the conversion of other types of nonadditive genetic variance.
Conversion of variance by physical restriction of recombination
The physical inhibition of recombination, as in apomictic asexual reproduction and some types of chromosomal rearrangements, can be considered to effectively convert epistatic to additive genetic variance. When recombination becomes physically restricted, groups of interacting genes that had previously been disrupted every generation by recombination are now ‘seen’ by selection as single alleles because recombination does not break them up. This is analogous to the situation described above, where increased homozygosity decreases the effectiveness of recombination at disrupting the coinheritance of interacting genes. In this way, phenomena that physically restrict recombination across previously freely recombining genomic regions may also convert nonadditive to additive variance. In the following sections, we will expand our discussion of how the appearance of chromosomal inversions and transitions from sexual to asexual reproduction can result in the conversion or release of nonadditive to additive variance that may facilitate adaptive evolution.
Is nonadditive variance important to adaptive evolution?
In order for the conversion of nonadditive to additive variance to be an important evolutionary process, a substantial amount of the genetic variance underlying fitness-related quantitative traits in natural populations must be nonadditive, that is, generated either by dominance or by epistasis (Goodnight, 1987, 1988; Cockerham and Tachida, 1988; Whitlock et al, 1993; Wang et al, 1998a). The relative amount of nonadditive variance is notoriously difficult to estimate (eg Frankel and Schork, 1996; Wade, 2002; Barton and Turelli, 2004), and the amount and evolutionary importance of nonadditive variance has been a topic of considerable debate (eg Coyne et al, 1997; Wade and Goodnight, 1998). Theoretically, traits more closely associated with fitness are expected to have a relatively high proportion of nonadditive to additive variance because strong directional selection is expected to erode away additive variance, leaving nonadditive variance (eg Wright, 1977; Roff, 1997). These expectations are generally supported by the available data (Falconer, 1981; Merilä and Sheldon, 1999). Thus, it is possible that in natural populations there is little additive variance but substantial nonadditive variance for fitness traits. Conversion of nonadditive to additive variance could potentially release additive variance so that fitness traits could continue to respond to directional selection, whereas the evolution of fitness traits without conversion could be constrained (Goodnight, 1988).
Dominance versus epistatic variance
Although both dominant and epistatic gene actions are nonadditive, the evolutionary consequences of their conversion to additive genetic variance are likely to be quite different (Goodnight, 1987, 1988; Whitlock et al, 1993; Whitlock, 1995; Wade and Goodnight, 1998; Barton and Turelli, 2004). Theory has shown that the conversion of dominance to additive variance that occurs with inbreeding, population bottlenecks, and population subdivision can increase the additive variance, and thus heritability, above original levels (Robertson, 1952; Cockerham and Tachida, 1988; Willis and Orr, 1993; Wang et al, 1998a; López-Fanjul et al, 2000). However, most of the increase in additive variance will involve the expression of deleterious recessive alleles and is thus likely to lead to purging rather than to adaptive evolution (Willis and Orr, 1993; Wade et al, 1996). In contrast, if much of the nonadditive variance is epistatic, the fixation or near fixation of different epistatic partners will cause the remaining segregating epistatic partners to act more additively as long as there is an additive component to the basic gene action of the still-segregating locus(i). Selection may then act on these additively acting alleles. Thus, the fixation or near fixation of epistatic partners associated with inbreeding, population bottlenecks, and population subdivision can expose alleles otherwise hidden from selection in large, panmictic populations (Wright, 1977; Wade et al, 1996; Goodnight and Stevens, 1997; Meffert, 2000; Wade, 2000; Barton and Turelli, 2004).
Several authors maintain that the widespread nature of inbreeding depression and the ability of models including only dominance variance to explain most empirical examples of conversion of nonadditive to additive variance means that the relative contribution of dominance variance to conversion is high and thus that conversion of variance will not play a significant role in adaptive evolution (eg Barton and Turelli, 1989, 2004; Willis and Orr, 1993; Coyne et al, 1997; López-Fanjul et al, 2004). However, others contest these conclusions on the grounds that we still lack the theoretical, statistical, and empirical tools to accurately determine the nature of underlying gene action, particularly if epistasis is involved (eg Bryant and Meffert, 1996; Wade et al, 1996; Wang et al, 1998b; Meffert, 2000; Naciri-Graven and Goudet, 2003).
Potential of epistatic variance to contribute to the conversion process
Most evolutionary biologists from Fisher and Wright onward agree that physiological epistasis, defined as nonadditive interactions between genes, is ubiquitous. Indeed, this type of epistasis is integral to a wide range of evolutionary theories, from the maintenance of sex, recombination, and genetic variation, to speciation (reviewed in Wolf et al, 2000). However, there is much disagreement regarding whether epistatic genetic variance, or statistical epistasis, accounts for a significant proportion of the total phenotypic variance of natural populations (Cheverud and Routman, 1995). It is this question that is relevant to evaluating the potential evolutionary importance of the conversion of nonadditive to additive variance.
At equilibrium, large, panmictic populations are expected to be at or close to mutation-selection balance at all loci. Slightly deleterious mutant alleles may be segregating at two or more epistatically interacting loci, but because these alleles are held at low frequency by selection, most of this polymorphism will contribute to additive rather than to epistatic variance. Moreover, new mutations that are beneficial on average across the experienced environments and genetic backgrounds will quickly sweep to fixation. It is unlikely that these positively selected alleles will remain segregating long enough for another beneficial mutant to arise at an epistatically interacting locus. Thus, positively selected mutants will sequentially fix across a relatively uniform genetic background, and there will be very little epistatic variance (reviewed in Brodie, 2000).
Substantial conversion of additive-by-additive epistatic variance only occurs when the allelic frequencies of interacting loci are intermediate before the conversion process begins, meaning that there is substantial genetic variation at all interacting loci (López-Fanjul et al, 1999, 2000). Some believe that the maintenance of this type of genetic variation requires conditions not easily conceived in natural populations experiencing directional selection (eg López-Fanjul et al, 1999, 2000).
However, genotype–environment interaction as well as phenomena such as population structure and admixture, frequency-dependent selection, and balancing selection due to environmental heterogeneity or antagonistic pleiotropy are believed to be important in the maintenance of intermediate allele frequencies at single loci (eg Roff, 1997) and may also help maintain variation at multiple loci (eg Gillespie and Turelli, 1989; Wade and Goodnight, 1998; Peters and Lively, 1999). In addition, stabilizing selection may facilitate the maintenance of variation at interacting loci relative to directional selection because it operates on the properties of linkage groups rather than on the effects of individual alleles (Lynch and Walsh, 1998). There is broad empirical support for the existence of many of these phenomena in natural populations (eg genotype–environment interaction: Sgro and Hoffman, 2004; Meffert and Hagenbuch, 2005; population structure: Hastings and Harrison, 1994; Wade and Goodnight, 1998; metapopulation structure: Hanski and Gaggioti, 2004; frequency-dependent selection: Kassen, 2002; Sinervo and Svensson, 2002).
Empirical evidence for epistatic variance
Despite the notorious difficulty of detecting epistatic variance (eg Cheverud and Routman, 1995; Goodnight, 1995; Whitlock, 1995; Frankel and Schork, 1996; Wade and Goodnight, 1998; Cheverud, 2000; Wade, 2001), epistatic variance can and does contribute to fitness-related traits (eg Bryant and Meffert, 1996; Long et al, 1996; Routman and Cheverud, 1997; Cheverud et al, 1999; Meffert, 2000; Kover and Caicedo, 2001; Bonhoeffer et al, 2004; Sanjuán et al, 2004). Recent QTL studies provide additional, more direct evidence that genes important to fitness do interact epistatically (Brockmann et al, 2000; Rüppell et al, 2004; reviewed in Cheverud, 2000). Moreover, there is a growing body of evidence supporting the possibility that substantial epistatic variance can exist within and between natural populations and subpopulations (Hard et al, 1992; Goodnight and Stevens, 1997; Lair et al, 1997; Cheverud, 2000; Fenster and Galloway, 2000; Templeton, 2000; Bradshaw et al, 2005).
Importantly, the specific types of epistatic interactions that are most likely to contribute to substantial conversion of variance and least likely to cause inbreeding depression often underlie traits important to fitness. For example, Routman and Cheverud (1997) found substantial evidence for epistatic interactions between QTL for juvenile body size in mice, including additive-by-additive epistasis. Interestingly, they determined that the types of epistasis that are predicted to make the largest contribution to the conversion of variance, additive-by-additive and dominance-by-additive epistasis, were twice as common and had more profound effects on additive variance than types of epistasis that do not make as substantive of a contribution to the conversion of variance. This is particularly impressive given that additive-by-additive epistasis is notably difficult to detect in genetic mapping studies (Routman and Cheverud, 1997). Many other recent QTL studies have detected additive-by-additive epistasis for fitness-related traits (reviewed in Cheverud, 2000), including litter size in mice (Peripato et al, 2004), flowering time in Arabidopsis thaliana (Juenger et al, 2005), and disease resistance in spring wheat (Yang et al, 2005).
Evidence for the conversion process
Conversion with restricted effective recombination
It is well-established that genetic diversity and heterozygosity decrease when a formerly outcrossing population undergoes inbreeding, population bottlenecks, or population subdivision (eg Wright, 1951; Robertson, 1952; Crow and Kimura, 1970; Lande, 1980; Whitlock and Fowler, 1999). However, the effect of these phenomena on additive genetic variance and thus on heritability and the ability of a population to respond to selection is more complex and less predictable (Wright and Cockerham, 1985; Barton and Turelli, 2004). If most of the genetic variation underlying quantitative traits is additive, the amount of additive genetic variation within a population will decrease proportionately as FIS, the inbreeding coefficient of the population, increases (Wright, 1951; Robertson, 1952; Lande, 1980).
These simple relationships between the inbreeding coefficient and additive genetic variance no longer hold if phenotypic trait values deviate from additivity (reviewed in Moreno, 1994). If much of the genetic variation for a trait is due to nonadditive variance within and/or between loci, the fixation or near fixation of allelic and epistatic partners that can result from inbreeding, population bottlenecks, or population subdivision may actually cause an increase in additive genetic variation (eg Goodnight, 1988; Whitlock et al, 1993; Wang et al, 1998a; Saccheri et al, 1999).
The first study to empirically demonstrate the release of additive genetic variance following a population bottleneck showed that additive genetic variation in housefly populations occasionally increased following a bottlenecking event (Bryant et al, 1986). In a later experiment, the lines were subjected to five additional founder-flush events and assayed for fitness (Bryant et al, 1990). The authors found that the lines had fully recovered from any inbreeding depression caused by bottlenecking and were able to quickly respond to selection to novel environments (Bryant et al, 1990; also reviewed in Meffert, 2000; Meffert and Hagenbuch, 2005). Similarly, additive genetic variance increased following a bottlenecking event in the Hawaiian fruit fly Drosophila silvestris (Carson and Wisotzkey, 1989) and a forced inbreeding event in D. melanogaster (López-Fanjul and Villaverde, 1989). In the latter study, the authors determined that the increase in additive variation for viability was best explained under a model that included the conversion of epistatic to additive variance.
More recently, Bryant and Meffert (1995) showed that the conversion of epistatic variance contributed to increased heritability for morphometric traits in bottlenecked relative to nonbottlenecked lines of houseflies (also see Meffert, 2000). Furthermore, Regan et al (2003) found that bottlenecked housefly lines had a more diverse set of responses to selection than nonbottlenecked lines. These results suggest that, contrary to the expectations of the additive model, bottlenecks may not always reduce evolutionary potential. Similar results were obtained by Wade et al (1996), and Pray and Goodnight (1997), who tracked the effect of inbreeding on phenotypic variance for traits important to fitness in Triboleum castaneum, and by Cheverud et al (1999), who studied the effects of inbreeding on additive variation in mice. Of particular note is a 1998 study of allozyme and quantitative genetic variation for fitness-related traits in North American populations of the pitcher-plant mosquito (Wyeomyia smithii) across a latitudinal gradient (Armbruster et al, 1998). The authors found that while allozyme heterozygosity declined precipitously with increasing latitude, additive genetic variance for preadult development time and critical photoperiod actually increased with latitude. Armbruster et al (1998) attributed this result to the conversion of nonadditive variance via the bottlenecking and population subdivision that accompanied W. smithii's spread into northern regions following the retreat of glaciers at the conclusion of the Pleistocene glaciation. This study was the first to document an evolutionary response linked to the conversion of nonadditive to additive variance in a natural system.
The maintenance and evolution of ‘supergenes,’ defined as closely linked loci that control phenotypic polymorphism (reviewed by Kelly, 2000), may also occur through a conversion of variance mechanism. Well-studied examples of supergenes include cases of color polymorphism in butterflies (eg Clarke and Sheppard, 1971; Naisbit et al, 2003) and land snails (Murray and Clarke, 1976), selfish genetic elements in mice (Silver, 1993), and self-incompatibility in flowering plants (reviewed in Castric and Vekemans, 2004). Supergenes are expected to evolve when balancing selection for the maintenance of certain gene combinations is strong relative to the recombination rate between the loci involved (Kelly, 2000). These conditions are facilitated when recombination between interacting loci is limited, when the effects of recombination are limited by high homozygosity, as with inbreeding (reviewed in Charlesworth, 2003), and when interacting loci are physically linked or occur within chromosomal inversions (reviewed in Wang et al, 2003). Supergenes thus evolve when interacting alleles can be co-inherited and act additively, perhaps via a conversion of variance process.
Conversion with physical limits on recombination
Chromosomal inversions
The idea that linked blocks of genes could play an important role in evolution largely stems from Dobzhansky's work on selectively maintained chromosomal inversions in natural populations of D. pseudoobscura (eg Dobzhansky, 1951, 1956; reviewed in Powell, 1997). Kimura (1956) further showed that if certain gene combinations increase fitness, an ‘inversion or other cross-over reducing mechanism’ (p 283) will be selectively favored and will often spread (also see Turner, 1967; Lande, 1984). This is because recombination between the genes in an inversion is often physically inhibited or even entirely blocked (Coyne et al, 1993; Navarro and Ruiz, 1997), which means that favorable gene combinations can be coinherited.
Allelic variation at genes spanned by a new favorable inversion will initially be mainly epistatic, because free recombination between the genes will still occur in most individuals. As the inversion spreads, this epistatic variance will be converted to additive variance between karyotypes. In this situation, there will essentially be two allelic types: (1) chromosomes carrying the inversion that have higher than the mean fitness of the ancestral population because they contain a combination of favorably interacting genes, and (2), chromosomes with free recombination that have the same mean fitness of the ancestral population. Effectively, individuals without the inversion are stuck in the flat saddle region of Figure 1, with a mean fitness of 0, because recombination breaks up any favorable (or unfavorable) gene combinations that may occur. In contrast, individuals with the inversion are at a fitness peak, with a fitness of +1. Additive variance will be at a maximum in this scenario when the inversion is at a frequency of 0.5.
Thus, in an analogous sense to the phenomena discussed above that cause reduced effective recombination, physically inhibited or blocked recombination caused by chromosomal inversions can enable evolutionary responses to selection via the conversion of epistatic to additive variance. As stated earlier, certain types of epistatic interactions, especially additive-by-additive epistasis, are most likely to contribute to the conversion of variance when the efficacy of recombination between interacting genes is reduced. However, when recombination is physically blocked between a whole set of genes, as in the case of genes spanned by inversions, the whole set of genes is inherited as a unit. This means that all types of epistatic interactions between these genes can contribute to the conversion of variance.
The question of whether chromosomal inversions can contribute to adaptation has received considerable empirical attention (reviewed in Powell, 1997). While a comprehensive review of this field is beyond the scope of this paper, we will provide a brief summary of the substantial body of research suggesting a link between inversions, recombination suppression, conversion of epistatic variance, and adaptation.
One important early study of ethological isolation from founder populations in chromosomally rearranged lineages of D. suboobscura found the most pronounced isolation for the only line homozygous for a chromosomal inversion, in which recombination is meaningless (Dodd and Powell, 1985). More recent work on D. suboobscura has indicated that the frequencies of certain inversions within both natural and laboratory populations are strongly correlated with different temperature regimes and seasons (Orengo and Prevosti, 1996; Kamping and Van Delden, 1999). Clines in inversion frequency related to variation in life history, morphological, and fitness-related traits and correlated to different climatic conditions have been characterized in D. suboobscura (Prevosti et al, 1988; Orengo and Prevosti, 1996; Schaeffer et al, 2003), other fruit fly species (Inoue et al, 1984; Rodriguez et al, 2000), and in mosquitoes (Coluzzi et al, 1979). Interestingly, Rodriguez et al (2000) found that only inversions associated with positive effects on trait values or fitness were involved in a latitudinal cline of D. buzzatti; inversions that were associated with neutral and negative effects on trait values or fitness did not show latitudinal patterns.
Some intriguing evidence linking chromosomal rearrangements and adaptive evolution comes from several recent studies of inversions in flies. Cáceres et al (1999) determined that common and/or fixed inversions tend to be long in both physical and map length (recombinational frequency) terms, and thus suppress recombination to a greater extent than rarer inversions (also see Cáceres et al, 1997). The authors interpreted this result to mean that longer inversions may have a greater chance of becoming common because the longer the inversion, the greater the chance that it includes interacting genes. Most recently, Schaeffer et al (2003) found that strong selection appeared to have acted on inversions in D. suboobscura, and argued that the inversions may have evolved as an adaptation to suppress recombination within coadapted gene complexes strongly linked to fitness.
The suppression of recombination linked to chromosomal inversions may also play a role in speciation. The common observation that closely related species are often fixed for different chromosomal inversions has led researchers to consider the possibility of a link between inversions and speciation (eg White, 1978; Rieseberg et al, 1999; Noor et al, 2001; Rieseberg, 2001; Navarro and Barton, 2003a, 2003b). Classical models for chromosomal speciation suggested that inversions facilitate speciation by acting as barriers to gene flow because inversion heterozygotes are often semisterile (reviewed by Rieseberg, 2001). More recent models assume that heterozygotes do not experience semisterility because recombination is suppressed in the region of inversion (see Coyne et al, 1993; Navarro and Ruiz, 1997) so that inversions are nearly neutral and can fix by drift (Noor et al, 2001; Rieseberg, 2001; Navarro and Barton, 2003a; Livingstone and Rieseberg, 2004). Subsequent to fixation by drift, inversions can act as a barrier to gene flow and populations can diverge at loci in the inverted region despite gene flow across the rest of the genome (reviewed in Livingstone and Rieseberg, 2004). Alternatively, as described above, inversions may be actively favored by selection if they suppress recombination between sets of favorably interacting alleles.
Recent studies provide further empirical evidence for a potential link between inversions and speciation. For example, many different inversions are fixed in humans and chimpanzees. A study of protein divergence between these two species found that the rate of divergence in inverted regions of the genome is more than double the rate of divergence in collinear regions (Navarro and Barton, 2003b; but see Bowers, 2003; Lu et al, 2003; Zhang et al, 2004). Furthermore, research on fruit flies has found that traits linked to reproductive divergence are often located in inverted regions (Noor et al, 2001; Machado et al, 2002). These studies suggest that evolutionary divergence during speciation may originate more commonly or be better-maintained in inverted regions. The conversion of variance process described above could be involved in the initial spread of these chromosomal inversions.
In general, there is a wide variety of support for the notion that the inhibition of recombination caused by chromosomal inversions may be selectively favored because beneficial gene combinations can be selected as an intact unit, though a definitive connection between inversions, increased access to additive variance, and a greater ability to respond to selection has yet to be shown. In our view, the conversion of epistatic to additive genetic variance that is likely to accompany the creation and fixation of chromosomal inversions may be an important factor in adaptive evolution and speciation.
Asexuality
As Sewall Wright (1977) first realized, the increased fidelity of genotypic transmission from generation to generation by asexual reproduction means that the response to selection might depart significantly from that predicted for a similarly diverse sexual population. More specifically, Wright suggested that mass selection upon a collection of genomes that are transmitted intact from generation to generation via asexual reproduction will act upon the composite properties of each genome as a whole (also see Lynch and Gabriel, 1983; Lynch, 1984a, 1984b and 1984c). This type of selection is potentially more efficient than selection upon sexual organisms, which, because genotypes are broken up each generation by genetic recombination, acts largely upon the average additive effects of alleles rather than on groups of interacting loci. In turn, the faithful nature of asexual reproduction can constitute an advantage if the particular niche(s) to which a clone is well-adapted remains constant over time. In this situation, sexual reproduction will be at a disadvantage in terms of creating well-adapted offspring. This constitutes a cost of sex known as recombinational load, defined as the breakup of beneficial gene combinations by recombination (Maslin, 1968; White, 1973).
The creation of an apomictic (nonrecombinational) asexual lineage from an existing sexual population inherently involves a population bottleneck of one individual and immediate fixation of all allelic combinations within each new lineage. In fact, apomictic asexuality is the most effective way of converting nonadditive variance to additive variance because the entire genome is transmitted as a whole, thereby ensuring that even the most complex allelic/genic combinations remain intact (Lynch and Gabriel, 1983; Lynch, 1984b). All types of epistatic variance (including dominance components) are converted to among-lineage additive variance upon the creation of multiple asexual lineages from sexual progenitors. Thus, the expectation that asexual evolution is ineffectual relative to evolution in sexual competitors is only supported if most of the underlying genetic variance is truly additive (Lande, 1976; Lynch and Gabriel, 1983). Otherwise, the generation of multiple asexual lineages will result in the conversion of variance that is nonadditive for the sexual population to variance that is additive at the level of an assemblage of coexisting asexual lineages fixed for different combinations of genes (Wright, 1977; Lynch and Gabriel, 1983; Lynch, 1984b). The increased access to additive variance for an asexual assemblage is analogous to the conversion of epistatic to additive variance within a sexual population and could enable adaptive evolution for the asexual population in a direction that is unavailable to the sexual population (Lynch and Gabriel, 1983; Lynch, 1984b, 1984c), though the ubiquity of sexual reproduction suggests that avoidance of recombinational load does not reliably counter the selective advantages of sex.
Nevertheless, a variety of studies have found evidence for evolution and adaptation occurring in a surprisingly effective manner in asexual populations (eg Parker, 1979a; Williamson, 1981; Glazier, 1992; Christensen et al, 1992; Toline and Lynch, 1994; Andrade and Roitberg, 1995; Sunnucks et al, 1998; Weeks and Hoffman, 1998; Wilson et al, 1999, 2003). These works cite such evidence as diverse and closely adapted clonal arrays (eg Parker, 1979a, 1979b and 1979c; Weeks and Hoffman, 1998; Wilson et al, 1999), natural clonal assemblages shown to have high (comparable to sexual) heritabilities for life history, morphological, and fitness-related traits (Stratton, 1991, 1992), and parthenogenetic genotypes that seem to outcompete sympatric sexual forms (eg Browne, 1992; Christensen et al, 1992; Weeks and Hoffman, 1998). Insight can also come from cyclical parthenogens, as demonstrated by Lynch (1984c). He determined that conversion of epistatic to additive variance occurred following episodes of sex in cyclically parthenogenetic Daphnia, and showed that this release of hidden genetic variance could lead to heritabilities higher than those achievable via obligate sexuality.
One last example comes from the literature considering the evolutionary mechanisms underlying the repeated transition of a recombining sex chromosome to a largely nonrecombining sex chromosome (Fisher, 1931; Bull, 1983; Rice, 1987; Charlesworth, 1991; Ellegren and Carmichael, 2001; Nicolas et al, 2005). There is a general consensus that this transition occurs because selection will favor reduced recombination between the sex-determining locus and alleles that increase fitness in the heterogametic sex, but are detrimental to the opposite sex in the sex chromosome marking the heterogametic some sex.
In summary, there seems to be a wide range of evidence lending support to the possibility that the inheritance of intact genomes across generations plays a role in facilitating asexual adaptation. It is reasonable to conclude that the ecological success of some asexual populations, contrary to the intuitive expectations of a largely additive world, may be in part linked to the increased access to additive genetic variance of genetically diverse asexual assemblages.
Conclusions
There is a variety of evidence suggesting that limits on recombination may facilitate adaptive evolution and that a more extensive exploration of the genetic mechanisms underlying ecological and evolutionary success in situations where recombination is effectively or actually limited is warranted. In general, further study of adaptive evolution in recombinationally limited populations could provide insight into key issues, such as the evolutionary relevance of nonadditive genetic variance and the selective value of recombination. Following the lead of earlier work that emphasizes the potential importance of nonadditive variance (eg Lynch and Gabriel, 1983; Ritchie and Kyriacou, 1994), we stress that an understanding of the nature and prevalence of epistatic variance may often be a key step in elucidating the evolution and adaptation of populations and species.
References
Andrade MCB, Roitberg BD (1995). Rapid response to intraclonal selection in the pea aphid (Acyrthosiphon pisum). Evol Ecol 9: 397–410.
Armbruster P, Bradshaw WE, Holzapfel CM (1998). Effects of postglacial range expansion on allozyme and quantitative genetic variation of the pitcher-plant mosquito, Wyeomyia smithii. Evolution 52: 1697–1704.
Barton NH, Turelli M (1989). Evolutionary quantitative genetics: how little do we know? Ann Rev Genet 23: 337–370.
Barton NH, Turelli M (2004). Effects of genetic drift on variance components under a general model of epistasis. Evolution 58: 2111–2132.
Birky Jr CW, Walsh JB (1988). Effects of linkage on rates of molecular evolution. Proc Natl Acad Sci USA 85: 6414–6418.
Bonhoeffer S, Chappey C, Parkin NT, Whitcomb JM, Petropoulos CJ (2004). Evidence for positive epistasis in HIV-I. Science 306: 1547–1550.
Bowers EJ (2003). Chromosomal speciation. Science 301: 764–765.
Bradshaw WE, Haggerty BP, Holzapfel CM (2005). Epistasis underlying a fitness trait within a natural population of the pitcher-plant mosquito, Wyeomyia smithii. Genetics 169: 485–488.
Brockmann GA, Kratzsch J, Haley CS, Renne U, Schwerin M, Karle S (2000). Single QTL effects, epistasis and pleiotropy account for two-thirds of the phenotypic F2 variance of growth and obesity in DU6i × DBA/2 mice. Genome Res 10: 1941–1957.
Brodie III ED (2000). Why evolutionary genetics does not always add up. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 3–19.
Browne RA (1992). Population genetics and ecology of Artemia: insights into parthenogenetic reproduction. Trends Ecol Evol 7: 232–237.
Bryant EH, McCommas SA, Combs LM (1986). The effect of an experimental bottleneck upon quantitative genetic variation in the housefly. Genetics 114: 1191–1211.
Bryant EH, Meffert LM (1995). An analysis of selectional response in relation to a population bottleneck. Evolution 49: 626–634.
Bryant EH, Meffert LM (1996). Nonadditive genetic structuring of morphometric variation in relation to a population bottleneck. Heredity 77: 168–177.
Bryant EH, Meffert LM, McCommas SA (1990). Fitness in serially bottlenecked populations of the house-fly. Am Nat 136: 542–549.
Bull JJ (1983). Evolution of Sex Determining Mechanisms. Benjamin/Cummings: Menlo Park.
Cáceres M, Barbadilla A, Ruiz A (1997). Inversion length and breakpoint distribution in the Drosophila buzzati species complex: is inversion length a selected trait? Evolution 51: 1149–1155.
Cáceres M, Barbadilla A, Ruiz A (1999). Recombination rate predicts inversion size in Diptera. Genetics 153: 251–259.
Carlborg Ö, Haley CS (2004). Opinion: Epistasis: too often neglected in complex trait studies? Nat Rev Genet 5: 618–625.
Carson HL (1990). Increased genetic variance after a population bottleneck. Trends Ecol Evol 5: 228–230.
Carson HL, Wisotzkey RG (1989). Increase in genetic variance following a population bottleneck. Am Nat 134: 668–673.
Castric V, Vekemans X (2004). Plant self-incompatibility in natural populations: a critical assessment of recent theoretical and empirical advances. Mol Ecol 13: 2873–2889.
Charlesworth B (1991). The evolution of sex chromosomes. Science 251: 1030–1033.
Charlesworth D (2003). Effects of inbreeding on the genetic diversity of populations. Philos Trans Roy Soc B 358: 1051–1070.
Cheverud JM (2000). Detecting epistasis among quantitative trait loci. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 58–81.
Cheverud JM, Routman EJ (1995). Epistasis and its contribution to genetic variance components. Genetics 139: 1455–1461.
Cheverud JM, Routman EJ (1996). Epistasis as a source of increased additive genetic variance at population bottlenecks. Evolution 50: 1042–1051.
Cheverud JM, Vaughn TT, Pletscher LS, King-Ellison K, Bailiff J, Adams E et al (1999). Epistasis and the evolution of additive genetic variance in populations that pass through a bottleneck. Evolution 53: 1009–1018.
Christensen B, Hvilsom M, Pedersen BV (1992). Genetic variation in coexisting sexual diploid and parthenogenetic triploid forms of Fridericia galba (Enchytraeidae, Oligochaeta) in a heterogeneous environment. Hereditas 117: 153–162.
Clarke CA, Sheppard PM (1971). Further studies on the genetics of the mimetic butterfly Papilio memnon L. Philos Trans Roy Soc B 263: 35–70.
Cockerham CC, Tachida H (1988). Permanency of response to selection for quantitative characters in finite populations. Proc Natl Acad Sci USA 85: 1563–1565.
Coluzzi M, Sabatini A, Petrarca V, Di Deco MA (1979). Chromosomal differentiation and adaptation to human environments in Anopheles gambiae complex. Trans Roy Soc Trop Med H 73: 483–497.
Coyne JA, Barton NH, Turelli M (1997). Perspective: a critique of Sewall Wright's shifting balance theory of evolution. Evolution 51: 643–671.
Coyne JA, Meyers W, Crittenden AP, Sniegowski P (1993). The fertility effects of pericentric inversions in Drosophila melanogaster. Genetics 134: 487–496.
Crow JF, Kimura M (1970). An Introduction to Population Genetics Theory. Harper and Row: New York.
Dobzhansky T (1951). Genetics and the Origin of Species. Columbia University Press: New York.
Dobzhansky T (1956). Genetics of natural populations. XXV. Genetic changes in populations of Drosophila pseudoobscura and Drosophila persimilis in some localities in California. Evolution 10: 82–92.
Dodd DMB, Powell JR (1985). Founder-flush speciation: an update of experimental results with Drosophila. Evolution 39: 1388–1392.
Ellegren H, Carmichael A (2001). Multiple and independent cessation of recombination between avian sex chromosomes. Genetics 158: 325–331.
Falconer DS (1981). Introduction to Quantitative Genetics, 2nd edn. Longman Group Ltd: Harlow.
Falconer DS, Mackay TFC (1996). Introduction to Quantitative Genetics, 4th edn. Longman Group Ltd: Harlow.
Fenster CB, Galloway LF (2000). The contribution of epistasis to the evolution of natural populations: a case study of an annual plant. In: Wolf JB, Brodie ED III, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 232–244.
Fisher RA (1918). The correlation between relatives on the supposition of Mendelian inheritance. Trans Royal Soc Edinburgh 52: 399–433.
Fisher RA (1930). The Genetical Theory of Natural Selection. Clarendon Press: Oxford.
Fisher RA (1931). The evolution of dominance. Biol Rev 6: 346–368.
Frankel WN, Schork NJ (1996). Who's afraid of epistasis? Nat Genet 14: 371–373.
Gibson PT (1996). Correcting for inbreeding in parent–offspring regression estimates of heritability with nonadditive and genotype × environment effects present. Crop Sci 36: 594–600.
Gillespie JH, Turelli M (1989). Genotype-environment interactions and the maintenance of polygenic variation. Genetics 121: 129–138.
Glazier DS (1992). Effects of food, genotype, and maternal size and age on offspring investment in Daphnia magna. Ecology 73: 910–926.
Goodnight CJ (1987). On the effect of founder events on epistatic genetic variance. Evolution 41: 80–91.
Goodnight CJ (1988). Epistasis and the effect of founder events on the additive genetic variance. Evolution 42: 441–454.
Goodnight CJ (1995). Epistasis and the increase in additive genetic variance: Implications for phase I of Wright's shifting-balance process. Evolution 49: 502–511.
Goodnight CJ, Stevens L (1997). Experimental studies of group selection: what they tell us about group selection in nature. Am Nat 150: S59–S79.
Hanski I, Gaggioti OE (2004). Metapopulation biology: Past, present, and future. In: Hanski I, Gagiotti OE (eds) Ecology, Genetics, and Evolution of Metapopulations. Elsevier Academic Press: Oxford. pp 3–22.
Hard JJ, Bradshaw WE, Holzapfel CM (1992). Epistasis and the genetic divergence of photoperiodism between populations of the pitcher-plant mosquito, Wyeomyia smithii. Genetics 131: 389–396.
Hastings A, Harrison S (1994). Metapopulation dynamics and genetics. Ann Rev Ecol Syst 25: 167–188.
Hill WG, Robertson A (1966). The effect of linkage on limits to artificial selection. Genet Res 8: 269–294.
Inoue Y, Watanabe T, Watanabe TK (1984). Evolutionary change of the chromosomal polymorphism in Drosophila melanogaster populations. Evolution 38: 753–765.
Juenger TE, Sen S, Stowe KA, Simms EL (2005). Epistasis and genotype–environment interaction for quantitative trait loci affecting flowering time in Arabidopsis thaliana. Genetica 123: 87–105.
Kamping A, Van Delden W (1999). A long-term study on interactions between the Adh and αGpdh allozyme polymorphisms and the chromosomal inversion In(2L)t in a semi-natural population of D. melanogaster. J Evol Biol 12: 809–821.
Kassen R (2002). The experimental evolution of specialists, generalists, and the maintenance of diversity. J Evol Biol 15: 173–190.
Kelly JK (2000). Epistasis, linkage and balancing selection. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 146–157.
Kimura M (1956). A model of a genetic system which leads to closer linkage by natural selection. Evolution 10: 278–287.
Kover PX, Caicedo AL (2001). The genetic architecture of disease resistance in plants and the maintenance of recombination by parasites. Mol Ecol 10: 1–16.
Lair KP, Bradshaw WE, Holzapfel CM (1997). Evolutionary divergence of the genetic architecture underlying photoperiodism in the pitcher-plant mosquito, Wyeomyia smithii. Genetics 147: 1873–1883.
Lande R (1976). Natural selection and random genetic drift in phenotypic evolution. Evolution 30: 314–334.
Lande R (1980). Genetic variation and phenotypic evolution during allopatric speciation. Am Nat 116: 463–479.
Lande R (1984). The expected fixation rate of chromosomal inversions. Evolution 38: 743–752.
Livingstone K, Rieseberg L (2004). Chromosomal evolution and speciation: a recombination-based approach. New Phytol 161: 107–112.
Long AD, Mullaney SL, Mackay TFC, Langley CH (1996). Genetic interactions between naturally occurring alleles at quantitative trait loci and mutant alleles at candidate loci affecting bristle number in Drosophila melanogaster. Genetics 144: 1497–1510.
López-Fanjul C, Fernández A, Toro MA (1999). The role of epistasis in the increase in the additive genetic variance after population bottlenecks. Genet Res 73: 45–59.
López-Fanjul C, Fernández A, Toro MA (2000). Epistasis and the conversion of non-additive to additive genetic variance at population bottlenecks. Theor Popul Biol 58: 49–59.
López-Fanjul C, Fernández A, Toro MA (2004). Epistasis and the temporal change in the additive variance–covariance matrix induced by drift. Evolution 58: 1655–1663.
López-Fanjul C, Villaverde A (1989). Inbreeding increases genetic variance for viability in Drosophila melanogaster. Evolution 43: 1800–1804.
Lu J, Li W-H, Wu C-I (2003). Comment on ‘Chromosomal speciation and molecular divergence–accelerated evolution in rearranged chromosomes’. Science 302: 988.
Lynch M (1984a). Destabilizing hybridization, general-purpose genotypes, and geographical parthenogenesis. Q Rev Biol 59: 257–290.
Lynch M (1984b). The genetic structure of a cyclical parthenogen. Evolution 38: 186–203.
Lynch M (1984c). The limits to life history evolution in Daphnia. Evolution 38: 465–482.
Lynch M, Gabriel W (1983). Phenotypic evolution and parthenogenesis. Am Nat 122: 745–764.
Lynch M, Walsh B (1998). Genetics and Analysis of Complex Traits. University of Arizona Press: Tuscon.
Machado CA, Kliman RM, Markert JA, Hey J (2002). Inferring the history of speciation from multilocus DNA sequence data: the case of Drosophila pseudoobscura and close relatives. Mol Biol Evol 19: 472–488.
Maslin TP (1968). Taxonomic problems in parthenogenetic vertebrates. Syst Zool 17: 219–231.
Meffert LM (2000). The evolutionary potential of morphology and mating behavior: the role of epistasis in bottlenecked populations. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 177–196.
Meffert LM, Hagenbuch KL (2005). The genetic architecture of house fly matingbehavior. Curr Top Dev Biol 66: 189–213.
Merilä J, Sheldon BC (1999). Genetic architecture of fitness and nonfitness traits: empirical patterns and development of ideas. Heredity 83: 103–109.
Moreno G (1994). Genetic architecture, genetic behavior, and character evolution. Ann Rev Ecol Syst 25: 31–44.
Muller HJ (1932). Some genetic aspects of sex. Am Nat 66: 118–138.
Murray J, Clarke B (1976). Supergenes in polymorphic land snails. I. Partula taeniata. Heredity 37: 253–269.
Naciri-Graven Y, Goudet J (2003). The additive genetic variance after bottlenecks is affected by the number of loci involved in epistatic interactions. Evolution 57: 706–716.
Naisbit RE, Jiggins CD, Mallet J (2003). Mimicry: developmental genes that contribute to speciation. Evol Dev 5: 269–280.
Navarro A, Barton NH (2003a). Accumulating postzygotic isolation genes in parapatry: a new twist on chromosomal speciation. Evolution 57: 447–459.
Navarro A, Barton NH (2003b). Chromosomal speciation and molecular divergence-accelerated evolution in rearranged chromosomes. Science 300: 321–324.
Navarro A, Ruiz A (1997). On the fertility effect of pericentric inversions. Genetics 147: 931–933.
Nicolas M, Marais G, Hykelova V, Janousek B, Laporte V, Vyskot B et al (2005). A gradual process of recombination restriction in the evolutionary history of the sex chromosomes in dioecious plants. PLoS Biol 3: 47–56.
Noor MAF, Grams KL, Bertucci LA, Reiland J (2001). Chromosomal inversions and the reproductive isolation of species. Proc Natl Acad Sci USA 98: 12084–12088.
Orengo D-J, Prevosti A (1996). Temporal changes in chromosomal polymorphism of Drosophila suboobscura related to climatic changes. Evolution 50: 1346–1350.
Parker Jr ED (1979a). Ecological implications of clonal diversity in parthenogenetic morphospecies. Am Zool 19: 753–762.
Parker Jr ED (1979b). Phenotypic consequences of parthenogenesis in Cnemidophorus lizards. I. Variability in parthenogenetic and sexual populations. Evolution 33: 1150–1166.
Parker Jr ED (1979c). Phenotypic consequences of parthenogenesis in Cnemidophorus lizards. II. Similarity of C. tesselatus to its sexual parental species. Evolution 33: 1167–1179.
Peripato AC, De Brito RA, Matiolo SR, Pletscher LS, Vaughn TT, Cheverud JM (2004). Epistasis affecting litter size in mice. J Evol Biol 17: 593–602.
Peters AD, Lively CM (1999). The red queen and fluctuating epistasis: a population genetic analysis of antagonistic coevolution. Am Nat 154: 393–405.
Phillips PC, Otto SP, Whitlock MC (2000). Beyond the average: the evolutionary importance of gene interactions and variability of epistatic effects. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 20–40.
Powell JR (1997). Progress and Prospects in Evolutionary Biology. Oxford University Press: New York.
Pray LA, Goodnight CJ (1997). The effect of inbreeding on phenotypic variance in the red flour beetle, Tribolium castaneum. Evolution 51: 308–313.
Prevosti A, Ribo G, Serra L, Aguade M, Balaòa J, Monclus M et al (1988). Colonization of America by Drosophila subobscura: experiment in natural populations that supports the adaptive role of chromosomal-inversion polymorphism. Proc Natl Acad Sci USA 85: 5597–5600.
Regan JL, Meffert LM, Bryant EH (2003). A direct experimental test of founder-flush effects on the evolutionary potential for assortative mating. J Evol Biol 16: 302–312.
Rice WR (1987). The accumulation of sexually antagonistic genes as a selective agent promoting the evolution of reduced recombination between primitive sex chromosomes. Evolution 41: 911–914.
Rieseberg LH (2001). Chromosomal rearrangements and speciation. Trends Ecol Evol 16: 351–357.
Rieseberg LH, Whitton J, Gardner K (1999). Hybrid zones and the genetic architecture of a barrier to gene flow between two sunflower species. Genetics 152: 713–727.
Ritchie MG, Kyriacou CP (1994). Genetic variability of courtship song in a population of Drosophila melanogaster. Anim Behav 48: 425–434.
Robertson A (1952). The effect of inbreeding on the variation due to recessive genes. Genetics 37: 189–207.
Rodriguez C, Piccinali R, Levy E, Hasson E (2000). Contrasting population genetic structure using allozymes and the inversion polymorphism in Drosophila buzzatii. J Evol Biol 13: 976–984.
Roff DA (1997). Evolutionary Quantitative Genetics. Chapman and Hall: New York.
Routman EJ, Cheverud JM (1997). Gene effects on a quantitative trait: two-locus epistatic effects measured at microsatellite markers and at estimated QTL. Evolution 51: 1654–1662.
Rüppell O, Pankiw T, Page Jr RE (2004). Pleiotropy, epistasis and new QTL: the genetic architecture of honey bee foraging behavior. Genetics 95: 481–491.
Saccheri IJ, Wilson IJ, Nichols RA, Bruford MW, Brakefield PM (1999). Inbreeding of bottlenecked butterfly populations: estimation using the likelihood of changes in marker allele frequencies. Genetics 151: 1053–1063.
Sanjuán R, Moya A, Elena SF (2004). The contribution of epistasis to the architecture of fitness in an RNA virus. Proc Natl Acad Sci USA 101: 15376–15379.
Schaeffer SW, Goetting-Minesky MP, Kovacevic M, Peoples JR, Graybill JL, Miller JM et al (2003). Evolutionary genomics of inversions in Drosophila pseudoobscura: evidence for epistasis. Proc Natl Acad Sci USA 100: 8319–8324.
Sgro CM, Hoffman AA (2004). Genetic correlations, tradeoffs and environmental variation. Heredity 93: 241–248.
Silver LM (1993). The peculiar journey of a selfish chromosome: mouse t haplotypes and meiotic drive. Trend Genet 9: 250–254.
Sinervo B, Svensson E (2002). Correlational selection and the evolution of genomic architecture. Heredity 89: 329–338.
Stratton DA (1991). Life history variation within populations of an asexual plant, Erigeron annuus (Asteraceae). Am J Bot 78: 723–728.
Stratton DA (1992). Life-cycle components of selection in Erigeron annuus: II. Genetic variation. Evolution 46: 107–120.
Sunnucks P, Chisholm D, Turak E, Hales DF (1998). Evolution of an ecological trait in parthenogenetic Sitobion aphids. Heredity 81: 639–647.
Templeton AR (2000). Epistasis and complex traits. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 41–57.
Toline CA, Lynch M (1994). Mutational divergence of life-history traits in an obligate parthenogen. Genome 37: 33–35.
Turner JRG (1967). On supergenes. I. The evolution of supergenes. Am Nat 101: 195–221.
Wade MJ (1996). Adaptation in subdivided populations: kin selection and interdemic selection. In: Rose MR, Lauder G (eds) Evolutionary Biology and Adaptation. Sinauer Associates: Sunderland. pp 381–405.
Wade MJ (2000). Epistasis as a genetic constraint within populations and an accelerant of adaptive divergence among them. In: Wolf JB, Brodie III ED, Wade MJ (eds) Epistasis and the Evolutionary Process. Oxford University Press: New York. pp 213–231.
Wade MJ (2001). Epistasis, complex traits, and mapping genes. Genetica 112–113: 59–69.
Wade MJ (2002). A gene's eye view of epistasis, selection, and speciation. J Evol Biol 15: 337–346.
Wade MJ, Goodnight CJ (1998). Perspective: the theories of Fisher and Wright in the context of metapopulations: when nature does many small experiments. Evolution 52: 1537–1553.
Wade MJ, Shuster SM, Stevens L (1996). Inbreeding: its effect on response to selection for pupal weight and the heritable variance in fitness in the flour beetle, Triboleum castaneum. Evolution 50: 723–733.
Wade MJ, Winther RG, Agrawal AF, Goodnight CJ (2001). Alternative definitions of epistasis: dependence and interaction. Trend Ecol Evol 16: 498–504.
Wang J, Caballero A, Hill WG (1998a). The effect of linkage disequilibrium and deviation from Hardy–Weinberg proportions on the changes in genetic variance with bottlenecking. Heredity 81: 174–186.
Wang J, Caballero A, Keightley PD, Hill WG (1998b). Bottleneck effect on genetic variance: a theoretical investigation of the role of dominance. Genetics 150: 435–447.
Wang Y, Wang X, Skirpan AL, Kao T-H (2003). S-RNase-mediated self-incompatibility. J Exp Bot 54: 115–122.
Weeks AR, Hoffman AA (1998). Intense selection of mite clones in a heterogeneous environment. Evolution 52: 1325–1333.
Weismann A (1904). The Evolution Theory. Edward Arnold: London.
White MJD (1973). Animal Cytology and Evolution. Cambridge University Press: Cambridge.
White MJD (1978). Modes of Speciation. W.H. Freeman: San Francisco.
Whitlock MC (1995). Two-locus drift with sex chromosomes: the partitioning and conversion of variance in subdivided populations. Theor Popul Biol 48: 44–64.
Whitlock MC, Fowler K (1999). The changes in genetic and environmental variance with inbreeding in Drosophila melanogaster. Genetics 152: 345–353.
Whitlock MC, Phillips PC, Moore FB-G, Tonsor SG (1995). Multiple fitness peaks and epistasis. Ann Rev Ecol Syst 26: 601–629.
Whitlock MC, Phillips PC, Wade MJ (1993). Gene interaction affects the additive genetic variance in subdivided populations with migration and extinction. Evolution 47: 1758–1769.
Williamson PG (1981). Palaeontological documentation of speciation in Cenozoic molluscs from Turkana Basin. Nature 293: 437–443.
Willis JH (1996). Measures of phenotypic selection are biased by partial inbreeding. Evolution 50: 1501–1511.
Willis JH, Orr HA (1993). Increased heritable variation following population bottlenecks: the role of dominance. Evolution 47: 949–957.
Wilson AC, Sunnucks P, Hales DF (1999). Microevolution, low clonal diversity and genetic affinities of parthenogenetic Sitobion aphids in New Zealand. Mol Ecol 8: 1655–1666.
Wilson ACC, Sunnucks P, Hales DF (2003). Heritable genetic variation and potential for adaptive evolution in asexual aphids (Aphidoidea). Biol J Linn Soc 79: 115–135.
Wolf JB, Brodie III ED, Wade MJ (eds) (2000). Epistasis and the Evolutionary Process. Oxford University Press: New York.
Wright AJ, Cockerham CC (1985). Selection with partial selfing. I. Mass selection. Genetics 109: 585–597.
Wright S (1951). The genetical structure of populations. Ann Eugen 15: 323–354.
Wright S (1977). Evolution and the Genetics of Populations, Vol 3: Experimental Results and Evolutionary Deductions. University of Chicago Press: Chicago.
Yang ZP, Gilbert J, Fedak G, Somers DJ (2005). Genetic characterization of QTL associated with resistance to Fusarium head blight in a doubled-haploid spring wheat population. Genome 48: 187–196.
Zhang J, Wang X, Podlaha O (2004). Testing the chromosomal speciation hypothesis for humans and chimpanzees. Genome Res 14: 845–851.
Acknowledgements
We would like to thank J Busch, C Goodnight, S Keller, and M Wade for their comments on an earlier version of the manuscript, and C Goodnight for suggestions for figures.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Neiman, M., Linksvayer, T. The conversion of variance and the evolutionary potential of restricted recombination. Heredity 96, 111–121 (2006). https://doi.org/10.1038/sj.hdy.6800772
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800772
Keywords
This article is cited by
-
Species Differentiation on a Dynamic Landscape: Shifts in Metapopulation Genetic Structure Using the Chronology of the Hawaiian Archipelago
Evolutionary Biology (2012)
-
Plant individuality: a solution to the demographer’s dilemma
Biology & Philosophy (2012)
-
Decanalization and the origin of complex disease
Nature Reviews Genetics (2009)