Abstract
LTR (long terminal repeat) retrotransposons are the main components of higher plant genomic DNA. They have shaped their host genomes through insertional mutagenesis and by effects on genome size, gene expression and recombination. These Class I transposable elements are closely related to retroviruses such as the HIV by their structure and presumptive life cycle. However, the retrotransposon life cycle has been closely investigated in few systems. For retroviruses and retrotransposons, individual defective copies can parasitize the activity of functional ones. However, some LTR retrotransposon groups as a whole, such as large retrotransposon derivatives and terminal repeats in miniature, are non-autonomous even though their genomic insertion patterns remain polymorphic between organismal accessions. Here, we examine what is known of the retrotransposon life cycle in plants, and in that context discuss the role of parasitism and complementation between and within retrotransposon groups.
Similar content being viewed by others
Introduction
Transposable elements are the primary hitchhikers of the cell, and comprise the bulk of higher plant genomes, ranging from 15% of the nuclear DNA in Arabidopsis thaliana to more than 90% in some Liliaceae. The majority of these elements are the Class I LTR (long terminal repeat) retrotransposons, which transpose via an RNA intermediate in a ‘Copy-and-Paste’ mechanism. Because retrotransposons use cellular resources and their own enzymes to replicate independently of the genome as a whole, they have been considered as ‘selfish DNA’ and nuclear parasites. They have thereby become in many cases more predominant than the cellular genes. The process by which they replicate is called retrotransposition (Figure 1), because the RNA transcript is reverse-transcribed (regarding the normal direction as specified by the Central Dogma) into DNA. It is thought to share many features with the internal life cycle of retroviruses such HIV (lentiviruses). However, the exact correspondence between the retroviral and retrotransposon life cycles has been explicitly examined in very few systems. Moreover, whereas at least a few of the retroviruses arriving in an organism during an infection must be functional in order for the infection to proceed, some LTR retrotransposon families appear to completely lack active members even if their genomic insertion patterns remain polymorphic between organismal accessions.
Theoretical life cycle of LTR retrotransposons. (a) Transcription of the mRNA, starting from the 5′ R region to the 3′ R region. (b) Translation and protein synthesis of active elements in GAG and POL; POL is further internally cleaved by AP in AP, RT-RNAseH and IN. (c) Dimerization of RNA before or during packaging, using a ‘kissing-loop’ mechanism based on DIS recognition (see text). (d) Packaging of RNA and start of reverse transcription. The GAGs polymerize to form the VLP, in which the reverse transcription is performed by the dual protein RT-RNaseH. This allows the synthesis of the first strand of the cDNA using the packaged RNA as matrix. (e) Degradation of the RNA matrix and initiation of synthesis of the second strand of the cDNA. (f) Completion of the double-stranded cDNA synthesis and linkage of the IN to the LTRs. (g) Double-stranded break and integration of the newly synthesized copy in a new genomic location.
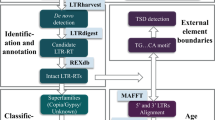
As their name indicates, the LTR retrotransposons are delimited by LTRs (Figure 2a), which contain signals needed for transcription. The process of reverse transcription renders the LTRs identical at the moment of integration of a new retrotransposon copy. They flank an internal sequence, which may or not code for the proteins necessary for carrying out retrotransposition. These proteins are encoded in two primary open reading frames (ORFs), which may in some elements be fused into one: the Gag, encoding the structural protein involved in nucleocapsid formation; the Pol, specifying the activities for the reverse transcription and integration of new copies. The Pol is a polyprotein and contains domains for an AP (aspartic proteinase), responsible for the post-translational processing of the Pol ORF product, RT (reverse transcriptase) and RNAseH, which, as a bifunctional polypeptide, carries out reverse transcription and IN (integrase), which inserts the new LTR retrotransposon copy into the genome. The LTR retrotransposons are generally divided into gypsy and copia groups, following the organization of the Pol ORF (Figure 2a) and named according to the type elements of Drosophila melanogaster. Some elements, generally closer to the gypsy group, possess a third potential ORF very similar to the env (envelope domain) ORF from retroviruses. This third ORF is involved in retroviral infectivity by mediating membrane–membrane fusion. Nevertheless, some LTR retrotransposons partly or entirely lack ORFs, and thus have had to be classified into other groups, named large retrotransposon derivatives (LARDs) (Kalendar et al., 2004), terminal repeats in miniature (TRIMs) (Witte et al., 2001) and Morgane (Sabot et al., 2006).
(a) LTR retrotransposon structure and retrotransposon groups. The groups are separated according to the presence or absence of the Gag and Pol ORFs. (b) Autonomy and non-autonomy. Non-autonomous groups lack coding capacity for GAG and POL. Autonomous groups encode GAG and POL, which may be nevertheless inactive owing to mutations. The parasitic families (italics) are proposed to use the machinery of host elements (bold) in a cis- (dashed green arrows) or a trans- (dashed red arrows) mode.
Both the retrotransposon and retrovirus life cycles are inherently error-prone and mutagenic. Once a copy is replicated and inserted, it likely displays neutral or nearly neutral rates of decay over time as a component of the genome. Hence, various members of a retrovirus or LTR retrotransposon family will display sequence divergence in their LTRs, processing signals or coding regions, including the occurrence of stop codons. For retroviruses, this has given rise to the concept of ‘quasispecies,’ in which a few infecting virons give rise to widely variable, related groups of retroviruses in the infected individual. The concept is appropriate for LTR retrotransposons as well (Casacuberta et al., 1995). Individuals of retrovirus or LTR retrotransposon quasispecies can vary in their competence for the various steps of replication. Functions for which individual elements are not competent may be complemented by parasitism on active elements (Escarmis et al., 2006; Holland, 2006; Sevilla and de la Torre, 2006).
Here, we will explore what is known about the various steps of the retrotransposon life cycle (Figure 1) within the framework of the retroviral model, and consider the causes and possible consequences of lack of function at each stage. These steps will be reviewed in their order of occurrence: transcription, translation, dimerization and packaging, reverse transcription and integration. We will also examine the potential workarounds used by the non-autonomous elements to override the various possible blocks to their life cycle.
Autonomy, non-autonomy, complementation and parasitism
For Class II elements, autonomy refers to the competence of an individual element within a family to express transposase and thereby catalyze its own transposition. Non-autonomous derivatives can be generated by deletion or mutation. For example, an autonomous maize Ac element that suffers a deletion in its transposase ORF becomes a non-autonomous Ds element. However, because Class I elements transpose replicatively, the question of autonomy is more complex for them. Individual non-autonomous Class I elements may give rise to groups of closely related but non-autonomous elements over time if they are able to be replicated despite their lack of autonomy. As described below, this appears to have occurred. Therefore, we apply the terms autonomous and non-autonomous on the group and family level for Class I elements (Figure 2b).
Families of retrotransposons containing individuals with an internal domain able to code for the requisite proteins can be said to be autonomous. Individual copies may be, to varying degrees, transcriptionally or translationally competent (translation leading to a functional protein) or active. Active elements may complement the life cycle blocks of inactive or incompetent members of the same family in cis and of other families or groups in trans. To the extent that the complementation reduces the ability of the active element to propagate, the inactive element is parasitic on the active one. This is conceptually similar to negative interfering viruses and their parasitism on otherwise virulent viruses (Hu et al., 1997). One can envisage a translationally incompetent element that is particularly successful at propagation emerging as a new subfamily and ultimately family of non-autonomous elements.
Recent findings have identified large, structurally uniform retrotransposon groups in which no member contains the Gag, Pol or Env internal domains. These groups are non-autonomous, yet individual elements may be active or inactive transcriptionally. Examples of non-autonomous groups are LARD, TRIM and Morgane. LARDs contain long LTRs and a long, conserved internal domain that shows no protein coding capacity. TRIMs are highly reduced, with short LTRs and internal domains that contain only the signals for reverse transcription. Morgane elements are intermediate between being autonomous and fully non-autonomous, and possess non-functional and small remnants of the Pol ORF. No trans-activating (trans-parasitic) element has been demonstrated yet for any of these groups.
Expression and translation of the LTR retrotransposons
Most of the plant LTR retrotransposons that have been investigated produce larger pools of transcripts in response to stress, biotic as well as abiotic. Examples of stresses shown to increase expression of various transcriptionally active LTR retrotransposons include chilling, infection, mechanical damage, in vitro regeneration, hybridization and generation of doubled haploids (Hirochika, 1995; Wendel and Wessler, 2000; Grandbastien et al., 2005). Consistent with the view that this represents transcriptional activation, numerous endogenous stress promoters share strong sequence similarities with LTRs (White et al., 1994; Dunn et al., 2006). Moreover, some transcripts of retrotransposon origin can be detected also under normal, non-stressed conditions, especially in active tissues such as embryos, root tips and buds. This has been seen for abundant elements such as BARE-1 (barley retroelement 1, Manninen and Schulman, 1993) or Sukkula (Kalendar et al., 2004). The selective advantages (for the retrotransposon or for the cell or plant) of stress induction remain unclear. The retrotransposon may benefit by exploiting a conserved and necessary, but sporadic, cellular response because it may be difficult for the plant to simultaneously silence transcription of the retrotransposon and maintain the response. For the plant, recruitment of solo LTRs containing stress response elements to roles as cellular promoters may provide a ready coordinating system for coping with stress. In any case, it is interesting that retroviruses such as HIV are stress induced as well (Nakamura et al., 2002).
Plant LTR retrotransposons are thought to be expressed by a classical polII promoter mechanism. The 5′ LTR drives this expression, which starts just before the 5′ R region (downstream of the TATA box) and extends until at least the 3′ R region within the 3′ LTR (Figures 1a and 2a; reviewed by Kumar and Bennetzen, 1999). The same bicistronic messenger RNA (mRNA) thus encodes at least the two ORFs, Gag and Pol. Retrotransposons and their mRNA are normally free of introns, and the RT acts on a mature, spliced mRNA. There are nevertheless exceptions such as the Ogre elements, which harbor an intron (Neumann et al., 2003). Expression from the LTRs, as for any polII promoter, is thus dependent on the host factors involved in mRNA synthesis. In this sense, the retrotransposon is parasitic on the cellular transcription mechanism. The process is expected to yield a polyadenylated mRNA if the LTR contains an efficient polyadenylation signal (Besansky, 1990; Suck and Traut, 2000). The mRNA is transferred to the cytoplasm, as is every cellular polII mRNA.
The level of LTR retrotransposon expression, even following stress induction, is generally much lower than for ‘classical’ genes (Wessler et al., 1995; Jääskeläinen et al., 1999). This is due to the LTR itself being a weak promoter, or to transcriptional or post-transcriptional repression, with possible mechanisms including methylation, heterochromatin formation and RNA interfernce (RNAi) (Okamoto and Hirochika, 2001). Retrotransposons sometimes may be driven also by endogenous ‘classical’ promoters (‘Master Copy’ theory, Deragon et al., 1996) following a ‘promoter-trap-like’ insertion. Furthermore, run-off transcription from LTRs can lead to overexpression or suppression of nearby genes (Kashkush et al., 2002, 2003). Some specific elements also may have an alternative means of expression, mediated by a polIII promoter. This is the case for the Cassandra TRIM elements, which can also be expressed using the 5S promoter within their LTR sequence (Kalendar et al., unpublished data).
Translation: only for the autonomous elements
In all non-autonomous groups studied so far (LARDs, TRIMs and Morganes), no ORFs potentially generating the polypeptides needed for transposition have been detected. Although only highly corrupted remnants have been found in Morgane (Sabot et al., 2006), no ORFs are found in the LARDs (Kalendar et al., 2004; Sabot et al., unpublished data). Hence, these groups of elements must parasitically exploit the polypeptides of other retrotransposons in order to replicate and propagate.
In autonomous retrotransposon groups, the bicistronic RNA is translated into GAG and POL proteins using the corresponding ORFs. The translational shift between the two ORFs can be accomplished in various ways. The ribosome entry site for the Gag ORF is the conventional, upstream one, but a less efficient, internal site can also serve Pol (IRES) (Meignin et al., 2003). This would lead to more GAG than POL products being synthesized during translation. For retroviruses, the greater amount of GAG produced by frameshifting is thought to match the stoichiometry required for particle assembly (Briggs et al., 2004). Furthermore, internal ribosome entry allows individual elements with stop codons in the Gag region to express Pol products nonetheless.
Alternatively, the sequence between Gag and Pol ORF can contain a small repetitive motif (such as AAAAA) that induces slippage of the ribosome, which then allows the translation of the second ORF by frameshifting (Jin and Bennetzen, 1989; Gao et al., 2003; Kovalchuk et al., 2005). This mechanism, including pseudoknot formation, is very common among plant viruses (Giedroc et al., 2000). Another possible means is the use of a specific and rare transfer RNA (tRNA), causing ribosomal stalling and slippage and allowing entry into the second ORF (reviewed by Hull and Covey, 1995). Despite the seeming logic of two ORFs for the sake of stoichiometric balancing of GAG with the non-structural products, it appears that some plant retrotransposons use the more ‘wasteful’ approach of a single ORF. For example, in BARE-1, there is only one ORF, leading to the synthesis of a single, large polyprotein including GAG and POL, which is later cleaved into functional units. Not only does the DNA sequence not reveal a putative frameshift, but also Western immunoblots probed with anti-GAG antibodies are consistent with a single ORF (Jääskeläinen et al., 1999; Jääskeläinen and Schulman, unpublished data). Post-translational processing of POL (and between GAG and POL for BARE-1) protein is performed endoproteolytically by the AP domain of the POL (Figure 1b; Jääskeläinen et al., 1999).
Nucleocapsid formation, packaging and dimerization
The retroviral GAG protein possesses three functional domains (in many investigated cases, cleaved into separate polypeptides from a GAG precursor), which are shared by the GAG of LTR retrotransposons. These are: the Capsid (polymerization); the Nucleocapsid itself, harboring the Zn-fingers and basic residues (nucleic acid interactions); the Matrix domain (association with the envelope protein; Adamson and Jones, 2004). The GAG of plant LTR retrotransposons appears to contain both the Capsid and Nucleocapsid domains, and possesses relatively little similarity to the Matrix domain (Jääskeläinen et al., 1999 and unpublished data).
Virus-like particles (VLPs) are formed to allow the reverse transcription of a (theoretically) specific RNA. The VLP results from GAG polymerization, mediated by the Capsid domain. The mechanism by which the RNA is internalized within the VLP is called Packaging. Packaging is generally selective for the RNA corresponding to the GAG that forms the VLP. In retroviruses, this selectivity is directed by the PSI sequence (packaging signal), a secondary RNA structure specifically recognized by either the Zn-fingers or the basic residues of the Nucleocapsid domain of the GAG proteins (reviewed by Harrison et al., 1995; Evans et al., 2004). The PSI sequence is generally located just after the PBS (primer-binding site) but before the Gag AUG. For HIV- and SIV-like retroviruses, the important and selective components of the PSI are an RCC sequence within a 7-base loop, followed or preceded by a less specific GAYC loop with a GC-rich stem (Harrison et al., 1995; Clever et al., 2002). Accessory stem–loop formations ensure a high level of specificity in packaging. For the LTR retrotransposons, the location of the PSI sequence has not yet been verified. Nevertheless, the high level of RNA structural conservation near the PBS (i.e., at the putative position of a PSI), as well as the family-specific motifs in this area, lead us to suppose a common mechanism for the retrovirus and retrotransposon packaging (Sabot and Schulman, unpublished data).
The packaging mechanism is quite specific for retroviruses, because the PSI sequence is necessary for the packaging of the RNA to proceed. If the PSI sequence is located upstream of a reporter gene, the reporter mRNA is efficiently packaged within the corresponding retroviral nucleocapsid (Guan et al., 2000). In the same way, if the PSI is missing, the retroviral infection is highly reduced. However, there is some room for flexibility or errors in packaging, as attested by the ‘retroprocessed pseudogenes’ (Hu and Leung, 2006). These sequences originate from spliced mRNA that is accidentally retrotransposed in the genome. Most of these sequences are inactive because of the lack of an associated promoter.
Nonetheless, non-autonomous groups of elements appear to be more efficiently propagated than are individual retroprocessed sequences (Witte et al., 2001; Kalendar et al., 2004; Sabot et al., 2005a, 2006; Antonius-Klemola et al., 2006). Therefore, non-autonomous groups may share common, specific PSI sequences with their active partners. Alternatively, generalist PSI sequences may allow packaging into a variety of VLPs. The observation that non-autonomous groups such as TRIMs and LARDs are propagated at all indicates that it is possible to escape from the constraints of protein expression, so long as the other steps including packaging are maintained.
The retroviral RNA is generally dimeric within the VLP; this dimerization either occurs before or simultaneously with packaging (Figure 1c and d; Brunel et al., 2002). For LTR retrotransposons, only the Ty1 element from the bakers’ yeast Saccharomyces cereviseae has been shown to be dimeric (Feng et al., 2000). In addition to Ty1, some complex elements (via template switching, see below), which are abnormally integrated, provide indirect proof for the dimeric state of the packaged RNA (Vicient et al., 2005; Sabot et al., 2005b and unpublished data). Dimerization occurs via the dimerization initiation signal (DIS), which allows the recognition and the interaction of the two RNAs, even in the absence of proteins (Darlix et al., 1990; Roy et al., 1990; Marquet et al., 1991). The signal is formed by a symmetrical loop near the PSI (reviewed by Paillart et al., 2004). This non-covalent, symmetrical intermolecular interaction is called a ‘kissing-loop complex’ for retroviruses, and is further stabilized by a more extended duplex (Paillart et al., 2004, and references within).
In a way analogous to that for the PSI sequence, the dimerization mechanism is assumed to be specific. Thus, elements of the non-autonomous groups must either harbor the same DIS as their active partners (forming specific heterodimers), or their competitive packaging efficiency must allow them to be preferentially packaged and therefore strictly homodimeric. Alternatively, they may be able to dimerize with RNAs bearing a variety of other DIS signals (nonspecific heterodimers). No specific bias in chimeras (complex elements, see below) between non-autonomous and autonomous elements has been found so far, leading to the idea that either competitive homodimers or nonspecific heterodimers can be formed.
Reverse transcription and cDNA formation
Our understanding of retroelement complementary DNA (cDNA) synthesis derives largely from the retroviruses and from the LTR retrotransposons of yeast (Figure 1d–f). Once packaged, the RNA forms a ‘buckle’, using ‘R-R’ pairing. The PBS sequence is primed generally by a tRNA (or a structural RNA) to allow the (−)-strand DNA synthesis by the RT, using the RNA as a template. Initial (−)-strand transcription proceeds to the 5′ end of the LTR, which lies inside the R domain, forming the ‘strong-stop’ (−)-strand cDNA. As the (−)-strand is synthesized, the RNaseH degrades the RNA template. This exposes the cDNA and allows a template switch to occur, with the (−)-strand cDNA being transferred to the R domain present at the 3′ end of the second template. Synthesis of the (−)-strand proceeds to include the PPT (polypurine tract) located just upstream of the 3′ LTR. Following RNAseH degradation of the RNA template at the PPT, (+)-strand synthesis is initiated, using small RNA oligonucleotides as the primers and the (−)-strand DNA as a template. Thus, a double-stranded cDNA is synthesized from the original mRNA, with two identical LTRs (Wilhelm and Wilhelm, 2001; Le Grice, 2003).
Reverse transcription itself is not specific for a particular RNA template, and any packaged RNA can be reverse transcribed as soon as it is primed. Therefore, elements of the non-autonomous groups do not require any specific features to be reverse transcribed beyond the PPT and PBS priming sites. This step is thus not a critical limiting part of the life cycle for them. Although there is some variation in the PBS motif, the great majority of LTR retrotransposons use the initiator-methionyl tRNA as the primer. Hence, examination of the PBS motif in the non-autonomous groups gives scant indication of what families of elements may serve as their hosts for reverse transcription. The reverse transcription reaction occurs with an error rate (2.5 × 10−5 errors/nucleotide/cycle) 100- to 1000-fold higher than that of the cellular DNA polymerase. The RT has a higher error rate because it can continue sequence extension even after misincorporating a nucleotide, has a low discrimination for incorrect ribonucleotides, lacks 3′ and 5′ exonuclease activity and suffers from the sliding of primers with respect to the template at long repetitive runs of nucleotides (Preston, 1996; Boutabout et al., 2001).
One of the most notable ‘accidents’ in the reverse transcription step is template switching. Usually, the switching occurs between the two packaged RNA transcripts during standard reverse transcription, but it can also occur between two unrelated RNAs packaged in the same nucleocapsid (reviewed by Mikkelsen and Pedersen, 2000). This leads to chimeric products such as the Veju_L (Sabot et al., 2005a) and BARE-2 (Vicient et al., 2005) elements. In the same way, template switching can promote the formation of ‘complex’ elements. These can take such forms as LTR-internal sequence-LTR-internal sequence-LTR, flanked by the same TSD (target-site duplication), and are often encountered in large Triticeae genomes (Vicient et al., 2005; Sabot et al., 2005b and our unpublished data). They generally span two blocks of elements, but they can be comprised of three or more blocks. The LTRs of the complex are highly similar to each other, as are the internal structures, testifying to a common origin.
Given the preceding discussion, the RNA plays two distinct and to some extent contradictory roles in the life cycle of a retrotransposon: as the template for translation and as the template for reverse transcription. Because the RNA is degraded during reverse transcription, if a single template copy serves both purposes translation must precede reverse transcription. Binding of GAG would likely exclude newly entering ribosomes and serve as a branch point leading to consequent packaging and reverse transcription. The question remains, however, if the same RNA actually serves both functions. Because translationally incompetent copies seem to be able to cis-parasitize the copies with ORFs, authors have supposed that there is a possible distinction between RNAs for translation and RNAs for reverse transcription. For retroviruses, even if most of them are subject to splicing in order to obtain the various mature mRNAs (and so the various proteins), the ‘genomic’ RNA that serves as the RT template is generally though to be the same as the mRNA used for translation (cis-preference; Poon et al., 2002). Nevertheless, recent experimental data suggest that the HIV-1 packaging may occur mainly in trans (Nikolaitchik et al., 2006).
Integration of the newly synthesized copy
The VLP is ultimately localized to the nucleus and the double-stranded cDNA transferred into the nucleus. However, no analyses show clearly whether localization occurs with the RNA templates or the partially completed cDNA still within the VLPs and associated with RT or following reverse transcription, with the double-stranded DNA bound to the IN only. The IN is bound to the LTR, in a specific way, based on the sequence of the LTR and particularly on the motifs of the borders. The IN makes an asymmetric, double-stranded break in the genomic DNA at the target site, which is generally 2–16 bp long. The nature of the target site, such as whether it is heterochromatic or whether other retrotransposons are already inserted, affects the propensity for integration (reviewed by Sabot et al., 2004). The double-stranded break is then repaired by the endogenous DNA repair system, leading to a TSD. The enzymology of the IN reaction, which does not require exogenous ATP or energy intermediates, is conserved in the retroviruses and in bacterial transposases such as that of bacteriophage Mu (Rice et al., 1996; Figure 1f).
In the Poaceae, and especially in large genome plants such as maize, wheat and barley, LTR retrotransposons are found mainly outside of gene space. In maize, for instance, where they compose more than 70% of the genomic DNA, very few gene mutations are associated with retrotransposon insertions (Kumar and Bennetzen, 1999, 2000; Bennetzen, 2000). In wheat and barley, the long genomic sequences analyzed so far do not show a large number of insertions into genes, even though >50% of the genomic DNA is derived from retrotransposons (Keller and Feuillet, 2000; Feuillet and Keller, 2002; Schulman and Kalendar, 2005; Sabot et al., 2005b). Of course, in such large genomes, genes compose less than 10% of genomic DNA, but even taking this low level into account, it seems that there is either a bias for insertion of LTR retrotransposons outside of genic regions or a highly efficient counter-selection against such insertions.
The non-autonomous groups therefore have two possible trans-parasitic routes for gaining integration function, as they do for packaging. They may either share the same specific IN recognition motif in their LTRs with an autonomous partner or they may possess generalist motifs allowing them to capture and use IN from a wide range of sources. A third scenario involves partial domestication by the cell, with selective pressure on a non-autonomous family to counteract and limit the spreading of autonomous elements. Here, the non-autonomous elements need only override autonomous and active ones during packaging and reverse transcription. Integration would not be a critical step, could be nonspecific and would only need to be efficient enough to counteract the decay of existing genomic copies. Nonspecific and fairly inefficient integration of double-stranded DNA is possible, as it has been shown for transformation both by Agrobacterium-mediated T-DNA and by vector bombardment. Some plant DNA viruses, particularly of the geminivirus, badnavirus and caulimovirus groups, also may integrate without encoding an IN function (Hull et al., 2000). The LARD group may adhere to this third mode. Their integrations are often atypical, either lacking definite TSDs or a complete LARD sequence, as might be expected from a nonspecific integration event. In contrast, TRIM integration appears to generate TSDs that are canonical for LTR retrotransposons (Witte et al., 2001; Sabot et al., 2005a), as does integration of Morgane elements (Sabot et al., 2006).
Conclusions
Our current understanding of the life cycle of the LTR retrotransposons, presented here, is more idealized and diagrammatic than dynamic. It does not connect activity rates for each reaction and event of the life cycle either with the sizes of steady-state pools of VLP components or with the ultimate integration frequency of a particular family of elements. For example, LARD elements are currently the most actively expressed ones in barley (Kalendar et al., 2004) but not necessarily the most frequently integrated (Sabot et al., 2005b). Some measurements currently can be made. The RNA pool size can be estimated directly from EST (expressed sequence tag) database analysis or by means of hybridization or quantitative polymerase chain reaction. The integration rate, reflecting the ability of an element to increase its prevalence, the ultimate goal of ‘selfish replication’, can be calculated using the Activity Index AI (defined as by Sabot et al., 2005b). This estimates the ratio between the complete (‘new’) and the fragmented or interrupted (older) copies.
A lot of endogenous activities and factors can affect the life cycle. These include heterochromatinization or methylation of the actively transcribed copies and repression by RNAi post-transcriptional mechanisms. In addition, the rate of accumulation of point mutations and small insertions or deletions, which would give rise to non-functional proteins, also affects the dynamics of a family of elements. The role of cis-parasitism (inactive copies of autonomous families on active copies) and trans-parasitism (non-autonomous on autonomous), as well as the likelihood of nested insertions of one element within the ORF or LTR of another, inactivating the genomic copy, will affect the population dynamics of both individual element families and families in host–parasite pairs in the genome over time.
A full understanding of the role of retrotransposons in genome evolution, therefore, must take into account not only the interaction of the retrotransposons with the genic fraction and the bulk effect of their integration, but also the secondary interactions between active and inactive copies within retrotransposon families and among autonomous and non-autonomous families of elements. Each level of interaction leaves room for both parasitism and selection. Moreover, the idea of the cis- and trans-parasitism, despite its acceptance and use for the Class II elements and the non-LTR retrotransposons, is not well developed for the LTR retrotransposons. It may change our view of the life and ‘half-life’ of LTR retrotransposons, as it was described by Ma and Bennetzen (2004) and Vitte and Panaud (2003). Hitchhikers should travel light, and only signals, not ORFs, may be enough for hitchhikers in the genome. The emerging picture shows that life of a LTR retrotransposon does not stop with its translational death, but rather when all its signals are dead, including those for transcription, translation, packaging (PSI), dimerization (DIS) and integration. Indeed, as the character of Lovecraft novels (1926), the mad Arabian poet Abdul Alhazred, said in the Necronomicon,
‘That is not dead which can eternal lie, And with strange aeons even death may die.’
References
Adamson CS, Jones IM (2004). The molecular basis of HIV capsid assembly – five years of progress. Rev Med Virol 14: 107–121.
Antonius-Klemola K, Kalendar R, Schulman AH (2006). TRIM retrotransposons occur in apple and are polymorphic between varieties but not sports. Theor Appl 112: 999–1008.
Bennetzen JL (2000). Transposable element contributions to plant gene and genome evolution. Plant Mol Biol 42: 251–269.
Besansky NJ (1990). A retrotransposable element from the mosquito Anopheles gambiae. Mol Cell Biol 10: 863–871.
Boutabout M, Wilhelm M, Wilhelm F-X (2001). DNA synthesis fidelity by the reverse transcriptase of the yeast retrotransposon Ty1. Nucleic Acids Res 29: 2217–2222.
Briggs JA, Simon MN, Gross I, Krausslich HG, Fuller SD, Vogt VM et al. (2004). The stoichiometry of GAG protein in HIV-1. Nat Struct Mol Biol 11: 672–675.
Brunel C, Marquet R, Romby P, Ehresmann C (2002). RNA loop-loop interactions as dynamic functional motifs. Biochimie 84: 925–944.
Casacuberta JM, Vernhettes S, Grandbastien MA (1995). Sequence variability within the tobacco retrotransposon Tnt1 population. EMBO J 14: 2670–2678.
Clever JL, Mirandar Jr D, Parslow TG (2002). RNA structure and packaging signals in the 5' leader region of the human immunodeficiency virus type 1 genome. J Virol 76: 12381–12387.
Darlix JL, Gabus C, Nugeyre MT, Clavel F, Barre-Sinoussi F (1990). Cis elements and trans-acting factors involved in the RNA dimerization of the human immunodeficiency virus HIV-1. J Mol Biol 216: 689–699.
Deragon JM, Gilbert N, Rouquet L, Lenoir A, Arnaud Ph, Picard G (1996). A transcriptional analysis of the S1Bn (Brassica napus) family of SINE retroposons. Plant Mol Biol 32: 869–878.
Dunn CA, Romanish MT, Gutierrez LE, van de Lagemaat LN, Mager DL (2006). Transcription of two human genes from a bidirectional endogenous retrovirus promoter. Gene 366: 335–342.
Escarmis C, Lazaro E, Manrubia SC (2006). Population bottlenecks in quasispecies dynamics. Curr Top Microbiol Immunol 299: 141–170.
Evans MJ, Bacharach E, Goff SP (2004). RNA sequences in the Moloney murine leukemia virus genome bound by the GAG precursor protein in the yeast three-hybrid system. J Virol 78: 7677–7684.
Feng YX, Moore SP, Garfinkel DJ, Rein A (2000). The genomic RNA in Ty1 virus-like particles is dimeric. J Virol 74: 10819–10821.
Feuillet C, Keller B (2002). Comparative genomics in the grass family: molecular characterization of grass genome structure and evolution. Ann Bot (London) 89: 3–10.
Gao X, Havecker ER, Baranov PV, Atkins JF, Voytas DF (2003). Translational recoding signals between gag and pol in diverse LTR retrotransposons. RNA 9: 1422–1430.
Giedroc DP, Theimer CA, Nixon PL (2000). Structure, stability and function of RNA pseudoknots involved in stimulating ribosomal frameshifting. J Mol Biol 298: 167–185.
Grandbastien MA, Audeon C, Bonnivard E, Casacuberta JM, Chalhoub B, Costa AP et al. (2005). Stress activation and genomic impact of Tnt1 retrotransposons in Solanaceae. Cytogenet Genome Res 110: 229–241.
Guan Y, Whitney JB, Diallo K, Wainberg MA (2000). Leader sequences downstream of the primer binding site are important for efficient replication of simian immunodeficiency virus. J Virol 74: 8854–8860.
Harrison GP, Hunter E, Lever AM (1995). Secondary structure model of the Mason-Pfizer monkey virus 5' leader sequence: identification of a structural motif common to a variety of retroviruses. J Virol 69: 2175–2186.
Hirochika H (1995). Activation of plant retrotransposons by stress. In: Oono K, Takaiwa H (eds). Modification of Gene Expression and Non-Mendelian Inheritance. NIAR: Japan. pp 15–21.
Holland JJ (2006). Transitions in understanding of RNA viruses: a historical perspective. Curr Top Microbiol Immunol 299: 371–401.
Hu KJ, Leung PC (2006). Complete, precise, and innocuous loss of multiple introns in the currently intronless, active cathepsin L-like genes, and inference from this event. Mol Phylogenet Evol 38: 685–696.
Hu WS, Bowman EH, Delviks KA, Pathak VK (1997). Homologous recombination occurs in a distinct retroviral subpopulation and exhibits high negative interference. J Virol 71: 6028–6036.
Hull R, Covey SN (1995). Retroelements: propagation and adaptation. Virus Genes 11: 105–118.
Hull R, Harper G, Lockhart B (2000). Viral sequences integrated into plant genomes. Trends Plant Sci 5: 362–365.
Jääskeläinen MJ, Mykkänen AH, Arna T, Vicient CM, Suoniemi A, Kalendar R et al. (1999). Retrotransposon BARE-1: expression of encoded proteins and formation of virus-like particles in barley cells. Plant J 20: 413–422.
Jin YK, Bennetzen JL (1989). Structure and coding properties of Bs1, a maize retrovirus-like transposon. Proc Natl Acad Sci USA 86: 6235–6239.
Kalendar R, Vicient CM, Peleg O, Anamthawat-Jonsson K, Bolshoy A, Schulman AH (2004). LArge retrotransposon derivatives: abundant, conserved but non-autonomous retroelements of barley and related genomes. Genetics 166: 1437–1450.
Kashkush K, Feldman M, Levy AA (2002). Gene loss, silencing and activation in a newly synthesized wheat allotetraploid. Genetics 160: 1651–1659.
Kashkush K, Feldman M, Levy AA (2003). Transcriptional activation of retrotransposons alters the expression of adjacent genes in wheat. Nat Genet 33: 102–106.
Keller B, Feuillet C (2000). Colinearity and gene density in grass genomes. Trends Plant Sci 5: 246–251.
Kovalchuk A, Senam S, Mauersberger S, Barth G (2005). Tyl6, a novel Ty3/gypsy-like retrotransposon in the genome of the dimorphic fungus Yarrowia lipolytica. Yeast 22: 979–991.
Kumar A, Bennetzen JL (1999). Plant retrotransposons. Annu Rev Genet 33: 479–532.
Kumar A, Bennetzen JL (2000). Retrotransposons: central players in the structure, evolution and function of plant genomes. Trends Plant Sci 5: 509–510.
Le Grice SF (2003). ‘In the beginning’: initiation of minus strand DNA synthesis in retroviruses and LTR-containing retrotransposons. Biochemistry 42: 14349–14355.
Lovecraft HP (1926). The call of Chtulhu. Weird Tales 11: 159–178.
Ma J, Bennetzen JL (2004). Rapid recent growth and divergence of rice nuclear genomes. Proc Natl Acad Sci USA 101: 12404–12410.
Manninen I, Schulman AH (1993). Analysis of BARE-1, a barley transposon. In: The Nordic Cereal Industry in an Integrating Europe. Proceedings from the 25th Nordic Cereal Congress. (Finnish Society of Cereal Technology, Helsinki, Finland).
Marquet R, Baudin F, Gabus C, Darlix JL, Mougel M, Ehresmann C et al. (1991). Dimerization of human immunodeficiency virus (type 1) RNA: stimulation by cations and possible mechanism. Nucleic Acids Res 19: 2349–2357.
Meignin C, Bailly JL, Arnaud F, Dastugue B, Vaury C (2003). The 5′ untranslated region and Gag product of Idefix, a long terminal repeat-retrotransposon from Drosophila melanogaster, act together to initiate a switch between translated and untranslated states of the genomic mRNA. Mol Cell Biol 23: 8246–8254.
Mikkelsen JG, Pedersen FS (2000). Genetic reassortment and patch repair by recombination in retroviruses. J Biomed Sci 7: 77–99.
Nakamura H, Masutani H, Yodoi J (2002). Redox imbalance and its control in HIV infection. Antioxid Redox Signal 4: 455–464.
Neumann P, Pozarkova D, Macas J (2003). Highly abundant pea LTR retrotransposon Ogre is constitutively transcribed and partially spliced. Plant Mol Biol 53: 399–410.
Nikolaitchik O, Rhodes TD, Ott D, Hu WS (2006). Effects of mutations in the human immunodeficiency virus type 1 gag gene on RNA packaging and recombination. J Virol 80: 4691–4697.
Okamoto H, Hirochika H (2001). Silencing of transposable element in plants. Trends Plant Sci 6: 527–534.
Paillart JC, Shehu-Xhilaga M, Marquet R, Mak J (2004). Dimerization of retroviral RNA genomes: an inseparable pair. Nat Rev Microbiol 2: 461–472.
Poon DT, Chertova EN, Ott DE (2002). Human immunodeficiency virus type 1 preferentially encapsidates genomic RNAs that encode Pr55 (Gag): functional linkage between translation and RNA packaging. Virology 293: 368–378.
Preston BD (1996). Error-prone retrotransposition: rime of the ancient mutators. Proc Natl Acad Sci USA 93: 7427–7431.
Rice P, Craigie R, Davies DR (1996). Retroviral integrases and their cousins. Curr Opin Struct Biol 6: 76–83.
Roy C, Tounekti N, Mougel M, Darlix JL, Paoletti C, Ehresmann C et al. (1990). An analytical study of the dimerization of in vitro generated RNA of Moloney murine leukemia virus MoMuLV. Nucleic Acids Res 18: 7287–7292.
Sabot F, Guyot R, Wicker T, Chantret N, Salse J, Laubin B et al. (2005b). Updating transposable element annotations from large wheat genomic sequences reveals diverse activities and gene association of elements. Mol Genet Genomics 274: 119–130.
Sabot F, Simon D, Bernard M (2004). Plant transposable elements, with an emphasis on grass species. Euphytica 139: 227–247.
Sabot F, Sourdille P, Bernard M (2005a). Advent of a new retrotransposon structure: the long form of the Veju elements. Genetica 125: 325–332.
Sabot F, Sourdille P, Chantret N, Bernard M (2006). Morgane, a new LTR retrotransposon group, and its subfamilies in wheats. Genetica (DOI 10.1007/S10709-006-0016-3).
Schulman AH, Kalendar R (2005). A movable feast: diverse retrotransposons and their contribution to barley genome dynamics. Cytogenet Genome Res 110: 598–605.
Sevilla N, de la Torre JC (2006). Arenavirus diversity and evolution: quasispecies in vivo. Curr Top Microbiol Immunol 299: 315–335.
Suck G, Traut W (2000). TROMB, a new retrotransposon of the gypsy-Ty3 group from the fly Megaselia scalaris. Gene 255: 51–57.
Vicient CM, Kalendar R, Schulman AH (2005). Variability, recombination and mosaic evolution of the barley BARE-1 retrotransposon. J Mol Evol 61: 275–291.
Vitte C, Panaud O (2003). Formation of solo-LTRs through unequal homologous recombination counterbalances amplifications of LTR retrotransposons. Mol Biol Evol 20: 528–540.
Wendel JF, Wessler SR (2000). Retrotransposon-mediated genome evolution on a local ecological scale. Proc Natl Acad Sci USA 97: 6250–6252.
Wessler SR, Bureau TE, White SE (1995). LTR-retrotransposons and MITEs: important players in the evolution of plant genomes. Curr Opin Genet Dev 5: 814–821.
White SE, Habera LF, Wessler SR (1994). Retrotransposons in the flanking regions of normal plant genes: a role for Copia-like elements in the evolution of gene structure and expression. Proc Natl Acad Sci USA 91: 11792–11796.
Wilhelm M, Wilhelm FX (2001). Reverse transcription of retroviruses and LTR retrotransposons. Cell Mol Life Sci 58: 1246–1262.
Witte C-P, Le QH, Bureau TE, Kumar A (2001). Terminal-repeat retrotransposons in miniature (TRIM) are involved in restructuring plant genomes. Proc Natl Acad Sci USA 98: 13778–13783.
Acknowledgements
FS was supported by a fellowship from CIMO and a University of Helsinki Postdoctoral Fellowship. The experiments described here were carried out under a grant the Academy of Finland, Project 106949.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sabot, F., Schulman, A. Parasitism and the retrotransposon life cycle in plants: a hitchhiker's guide to the genome. Heredity 97, 381–388 (2006). https://doi.org/10.1038/sj.hdy.6800903
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800903
Keywords
This article is cited by
-
Homoeologous evolution of the allotetraploid genome of Poa annua L.
BMC Genomics (2023)
-
Comparative genomics reveals low levels of inter- and intraspecies diversity in the causal agents of dwarf and common bunt of wheat and hint at conspecificity of Tilletia caries and T. laevis
IMA Fungus (2022)
-
Full-length LTR retroelements in Capsicum annuum revealed a few species-specific family bursts with insertional preferences
Chromosome Research (2021)
-
Recent advancement of NGS technologies to detect active transposable elements in plants
Genes & Genomics (2021)
-
Genomic abundance and transcriptional activity of diverse gypsy and copia long terminal repeat retrotransposons in three wild sunflower species
BMC Plant Biology (2018)