Abstract
We characterized the population genetics of two Neotropical dioecious palm species of Chamaedorea with contrasting life strategies from the region that is both the northernmost extent and most species rich of the genus. Chamaedorea tepejilote is a common, wind-pollinated arboreal understory palm. Although most adult plants reproduce each year, only a few individuals produce the majority of flowers and seeds. Chamaedorea elatior, conversely, is an uncommon climbing subcanopy palm with entomophilous flowers (insect-pollinated characteristics). Most of the mature palms do not reproduce in consecutive years and fruiting is episodic. Isozymes with a total of 107 alleles for 27 loci of 17 enzymes from six populations were assessed. For both species, co-occurrence of high levels of genetic variation and homozygosity was observed (C. tepejilote: He: 0.385–0.442, f: 0.431–0.486; C. elatior: He: 0.278–0.342, f: 0.466–0.535). Genetic differentiation of C. elatior was much lower (θ=0.0315) than that for C. tepejilote (θ=0.152). The contrast in differentiation may be influenced by differences in the spatial scale of the genetic neighborhoods of the two species. The simultaneous maintenance of inbreeding and of a large number of alleles within the populations is attributable to the low and variable number of mating pairs. Demographic studies indicate that this pattern could be explained by low reproductive frequency among individuals and over years in C. elatior and by reproductive dominance in C. tepejilote.
Similar content being viewed by others
Introduction
The genetic reservoirs that are present in remnant tropical rainforests have been strongly influenced by the constantly changing status of their populations over relatively recent time periods as well as by the long-term evolutionary history of the species. Genetic variation patterns have resulted from diverse processes at different spatial and temporal scales. In Neotropical rainforests, genetically distinct regional populations of some plant species have been found among areas that differ in biogeographic history, suggesting long-term regional isolation (Aide and Rivera, 1998; Hamilton, 1999; Dick et al., 2003; Novick et al., 2003). Geographically peripheral populations have sometimes shown distinctive genetic patterns (Lavin et al., 1991; Chase et al., 1995; Gillies et al., 1999; Cavers et al., 2003). The evolutionary connectedness between populations depends on their location within the species range. For example, edge effects, such as low genetic variation or geographic distinctiveness, have been reported for most northerly populations of species that migrated into North America from South America (Chase et al., 1995; Chamberlain, 1998; Cavers et al., 2003). Conversely, species in groups that diversified in North Mesoamerica could be predicted to show higher levels of genetic variation than conspecific populations resulting from range expansion toward the South. At a local scale, the low genetic differentiation observed among conspecific populations of many Neotropical plants (Loveless, 1992; Hamrick, 1994; Murawski, 1995; Nason et al., 1996a; White et al., 1999; Lemes et al., 2003) indicates that either the populations have been derived from one another in the fairly recent past or that there are high levels of gene flow. Thus, both regional genetic differentiation and high genetic similarity at local scales depend on biogeographic history and gene flow (Caron et al., 2000; Novick et al., 2003; Dick et al., 2003).
For a single species, population subdivision could increase when evolutionary and ecological independence among populations is induced by landscape heterogeneity. In addition, low density or rare plants with discontinuous populations with a wide regional distribution are expected to show lower levels of genetic differentiation than high-density populations of common plants—given that the comparisons are made at similar spatial scales (Ellstrand, 1992; Cole, 2003; Degen et al., 2004). For example, Hall et al (1994a) reported substantial differentiation at a small geographic scale for the abundant tree Pentaclethra macroloba in comparison to Cordia alliodora (Chase et al., 1995) in areas where the overall abundance of the latter species was substantially lower than that of the former. In general, because biogeographic history and habitat heterogeneity are main determinants of genetic structure among populations, the genetic assessment of congeneric species in areas of overlap could clarify the influence of geographical barriers, population density or neighborhood size on population genetic differentiation.
The genetic diversity of a population is generally reorganized through mating and recombination following colonization, and then structured through localized genetic drift and possibly selection (Wright, 1978; Lefèvre et al., 2004). After a species' initial establishment, most recruitment is from local sources (Olivieri and Gouyon, 1997) resulting in an enduring genetic patch structure (Wright, 1943). One major effect of such structure is the establishment of genetic isolation by distance, which depends on the effective population size (Malécot, 1948; Sokal and Wartenberg, 1983; Epperson et al., 1999). In obligate outcrossing species, homozygote excess within populations ensues from mating between neighboring relatives (Epperson and Álvarez-Buylla, 1997), temporal reproductive isolation (Robertson, 1964; Hendry and Day, 2005), episodic reproductive events (Epperson and Chung, 2001) and nonrandom dispersal (Garant et al., 2005; Postma and Noordwijk, 2005). Thus, all those forms of nonrandom mating, and their implications for recombination and gene movement, have a major effect on genetic variation patterns of outcrossing plants. Even for populations having high numbers of reproductive adults, effective population sizes may be small due to asynchronous reproduction (for example, Hall et al., 1994a; Chase et al., 1996; Degen et al., 2004).
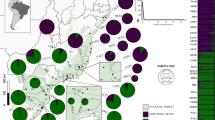
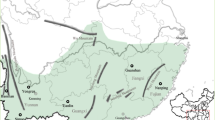
This article reports a population genetic study of two demographically well-characterized species of Chamaedorea, the most speciose genus of neotropical palms (Henderson et al., 1995). The study species, Chamaedorea elatior and Chamaedorea tepejilote, are both long-lived, dioecious obligate-outcrossers, but they differ in growth form, spatial, and geographical distributions, genetic neighborhood size and individual reproductive periodicity over years. Few comparisons of this type have been made in non-woody tropical rainforest species (Sarukhán et al., 1984). The research reported is part of an ongoing project on the evolutionary biology and conservation of Chamaedorea palms in Mexican tropical rainforest. Based on fossil pollen, it was concluded that the genus has been in the region since at least the Pliocene (Graham and Dilcher, 1998). A highly valued and overexploited non-timber resource at a world-wide scale since the 19th century (Hodel, 1992; Oyama, 1992; Vovides and Garcia-Bielma, 1994), local Chamaedorea populations have been going extinct from both habitat destruction and a long anthropogenic history of extraction of seeds, leaves and entire individuals from otherwise undisturbed forests (Oyama, 1992, 1993). Because effects of selective harvesting on population viability for a given species depend on conditions before management practices, an assessment of their genetic effects requires information about organization of genetic variation of pristine populations (Hall et al., 1994b; Nason et al., 1996b). In addition, the habitat heterogeneity associated with mountains has been suggested to be a major factor in regional differentiation in Mesoamerican rain forests (Dick et al., 2003). Accordingly, we sampled the most northerly undisturbed mountainous area (Figure 1) within the larger of the two centers of distribution of the genus (Hodel, 1992).
In this study, patterns of population genetic variation of the two study species were used to address the following questions (1) What are the levels of genetic variation of C. elatior and of C. tepejilote in a mountainous region within the area where the genus diversified? (2) What do allelic patterns reveal about the maintenance of population genetic differentiation at a local scale? (3) Do their contrasting reproductive dynamics support similar deviation levels from Hardy–Weinberg equilibrium? (4) Since dioecy within the study populations is promoting outcrossing, but there are other factors of life history that favor nonrandom mating, do populations harbor simultaneously high levels of genetic variation and inbreeding?
Materials and methods
Study system
Chamaedorea is the largest, terminal and most speciose genus of a chain of related genera of palms (Uhl and Dransfield, 1987). In this genus, vegetative characters (size, color, leaves or growth form) are highly variable among species, within populations and within individuals. Species and subgenera have been mostly diagnosed using discrete inflorescence and flower attributes (Hodel, 1992). Chamaedorea pinnatifrons and C. tepejilote are two of the more widely distributed species showing a high degree of morphological variation over an ecologically differentiated gradient with uncertain taxonomic status. Quero (1994) recognized two different species (Chamaedorea concolor and C. pinnatifrons) within the morphological variation that Hodel (1992) categorized as C. pinnatifrons. Hodel (1992) distinguished two species (Chamaedorea alternans and C. tepejilote) within the morphological variation that Henderson et al. (1995) considered to be one species, C. tepejilote. Recently, Bacon and Bailey (2006) showed amplified fragment length polymorphisms differentiation between these two morphological groups. They also suggested that C. alternans and C. tepejilote are not sister species (Thomas et al., 2006).
In this study, we selected two species showing contrasting traits of pollination, growth form, plant density and individual reproductive periodicity. They are C. tepejilote and C. elatior. Since the status of C. tepejilote and C. alternans is unresolved, we will use the name C. tepejilote for the latter (on the basis of nomenclatural priority).
C. tepejilote is a dioecious, arboreal palm distributed from Mexico to Colombia and reaches up to 5 m in height. Its small and greenish-yellow flowers are wind-pollinated (Otero-Arnaiz and Oyama, 2001). Several species of birds and bats eat their fruits (Trejo-Pérez, 1989) and may disperse the seeds, but fruits are mainly dispersed by gravity. Most seeds are eaten by seed predators (Oyama, 1991, 1997). Most mature individuals produce reproductive structures during consecutive years in permanent study sites established previously in the lowland rainforest Los Tuxtlas of Mexico, where C. tepejilote is one of the most abundant understory plants (Oyama, 1990). In contrast, C. elatior is a dioecious, climbing species distributed from Mexico to Honduras (Aguilar, 1986; Henderson et al., 1995), and reaches over 20 m in length. Insect pollination is indicated by the sticky pollen and the scented small greenish-yellow male flowers of this species (Henderson, 1986; Oyama, 1997). Pooled 4-year demographic data of one C. elatior population in Los Tuxtlas showed high mortality at adult stages (approximately 20%). Most reproductive individuals produced flowers or fruits only once during the study period (Luna, 1999).
Study populations and field methods
In the mountainous area of Los Tuxtlas Region (Mexico), we explored the two main massifs, Santa Marta Mountain and the San Martin Volcano (Figure 1). Annual rainfall is approximately 4700 mm and annual mean temperature is about 23°C. Full descriptions of the region are given by González et al. (1997). This region has extensive areas of pristine rainforest and the highest number of Chamaedorea species on the Atlantic coast of Mexico (Zarco-Espinosa, 1999). For each species, three undisturbed tropical rainforest areas in Los Tuxtlas were selected. For each species, individuals taller than 50 cm (juvenile and adults) located at least 5 m away from each other were sampled from rainforest patches of several hundred hectares, where a large population of either or both of the two studied species occurred. In total, 346 individuals of C. elatior and 347 of C. tepejilote were sampled. The C. elatior samples consisted of 197 individuals from the Los Tuxtlas Field Station ‘eFS’ (95°06′44″W, 18°35′22″N, 150–250 m a.s.l.), 78 from Santa Marta ‘eSM’ (94°58′44″W, 18°25′31″N, 1224 m a.s.l.) and 71 from San Martin Pajapan ‘eMP’ (94°44′28″W, 18°18′43″N, 700 m a.s.l.). For C. tepejilote, we collected exclusively individuals that Hodel (1992) defined as C. alternans, but the most recent taxonomic status sensu Henderson et al. (1995) is used here because this includes the morphological variation previously considered as C. alternans. Thus, 219 palms from Los Tuxtlas Field Station ‘tFS’ (95°06′16″W, 18°35′45″N, 150 m a.s.l.), 63 from Santa Marta ‘tSM’ (94°59′45″W, 18°25′25″N, 1224 m a.s.l.) and 65 from San Martin Tuxtla ‘tMT’ (95°10′49″W, 18°33′08″N, 1200 m a.s.l.) were sampled. From each individual, three pinnae from the youngest leaf were washed, placed in polyethylene bags, frozen in liquid nitrogen and stored at −70°C until processed for enzyme analysis.
At the Field Station, numbers of palms surpassed those sampled in the other localities because every palm of either C. tepejilote or C. elatior had been recorded and mapped within permanent study sites during a 4-year period (Oyama, 1990; Luna, 1999). Therefore, to standardize both the strategy and size of sampling, resampling (N=63) was carried out within the long-term study sites. Thus, the first series of statistical analyses is termed ‘unequal sizes’ and the second ‘equal sizes’.
Protein electrophoresis and statistical methods
A 1 cm2 leaf tissue sample per individual was finely sliced and ground in an extraction buffer consisting of three parts YO buffer (Yeh and O'Malley, 1980) and one part Veg II buffer (Cheliak and Pitel, 1984). Enzyme analyses were carried out following the standard procedures for starch gel electrophoresis (Soltis and Soltis, 1989; Kephart, 1990). Seventeen enzymes were dyed in three buffer systems. (1) C system (Stuber et al., 1988) resolved: DIA (E.C. 1.6.99), G6PD (E.C. 1.1.1.49), GDH (E. C. 1.4.1.3), EST (E.C. 3.1.1.), LAP (E.C. 3.4.11.1), GOT (E.C. 2.6.1.1), CPX (E.C. 1.11.1.7), APX (E.C. 1.11.1.7) and RUB (E.C. 4.1.1.39). (2) D system (Stuber et al., 1988) resolved MDH (E.C. 1.1.37), PGM (E.C. 5.4.2.2), ME (E.C. 1.1.1.40) and PGI (E.C. 5.3.1.9). (3) Morpholin-citrate (Wendel and Weeden, 1989) resolved ACPH (E.C.3.2.3.2), IDH (E.C. 1.1.1.41), 6PGD (E.C. 1.1.1.44) and SDH (E.C. 1.1.1.25). Staining protocols were modified to obtain optimal resolution of genotypes (Luna, 1999). Assignment of genotypes was consistent with the known enzyme structure (Soltis and Soltis, 1989; Wendel and Weeden, 1989). Putative loci and alleles were designated sequentially. Loci and alleles that were most anodally migrating were denoted in sequence by ‘1’, ‘2’, and so on.
The genetic variation of C. elatior and C. tepejilote was characterized by measuring the number of alleles (A), total alleles (AT), exclusive alleles (AX), effective alleles per locus (Ae=1/Σpi2, where pi is the frequency of allele i (Nei, 1987)) and the percent of polymorphic loci (P). Comparisons between A and Ae among populations and between species were performed using the Wilcoxon test (Sokal and Rohlf, 1979). Genetic diversity was assessed by calculating the observed heterozygosity (Ho), the expected heterozygosity (He) under Hardy–Weinberg equilibrium ‘HW’ (Levene, 1949), and the Nei (1987) measures: within populations (HS), among populations (DST), the total diversity (HT) and the genetic differentiation coefficient (GST). Differences in levels of genetic diversity between species were tested for significance using Mann–Whitney U-tests (Sokal and Rohlf, 1979). An inbreeding coefficient per population was estimated by f=(He−Ho)/He, and significant deviations from HW were determined by X2=f2N(k−1), d.f.=[k(k−1)]/2, where N is the sample size and k the number of alleles (Li and Horvitz, 1953). Nei's (1972, 1978) genetic differentiation between pairs of populations was summarized in an unweighted pair group method with arithmetic mean (UPGMA) phenogram (Sneath and Sokal, 1973). Fixation indices per locus and population differentiation were tested for significance by following Markov Chain Monte–Carlo methods (Raymond and Rousset, 1995) using the tools for population genetic analyses (TFPGA) program, version 1.3 (Miller, 1997). Genetic structure was measured by f, F and θ statistics (Weir and Cockerham, 1984), which estimate FIS, FIT and FST (Wright, 1951), respectively. Thus, excess of homozygosity is indicated by statistically significant positive values of f and F and deficits by negative values. The θ values may indicate from equal (θ=0) up to entirely different (θ=1) allele frequencies among populations. Li and Horvitz's (1953) X2 statistic was used to test if f and F values per locus were significantly different from zero or not, and significance of FST per locus was determined by X2=2N FST (k−1), d.f.=(k−1)(s−1), s is the number of subpopulations (Workman and Niswander, 1970). Confidence intervals at 95% for the mean f, F and θ statistics were calculated by a bootstrap procedure and the variances of each statistic was estimated by jackknifing over loci, using the TFPGA (Miller, 1997) computer program. Gene flow was indirectly estimated from Nm=((1/FST)−1)/4 (Wright, 1951) and also from GST. Populations were geo-referenced and pairwise geographical distances were calculated in ArcView and ILWIS programs.
Results
Genetic variation
The 17 enzymes stained for gave a total of 28 banding zones (loci), and all but one of these were observed in both C. elatior and C. tepejilote. In part because samples for both species were done simultaneously and hence under the exact same gel and staining conditions, and because of consistency in inheritance patterns (Wendel and Weeden, 1989), we presume that the banding zones are the same allozyme loci. The only locus that was not shared, EST-4, was excluded from all statistical analyses. For the 27 remaining loci, 107 alleles from a total of 693 individuals in the samples of unequal sizes, and 102 alleles from 378 palms in the equalized samples were recorded. For unequal sample sizes, C. tepejilote showed 101 total alleles which included 13 exclusive alleles, whereas AT=94 and AX=6 from C. elatior were recorded, respectively. For equalized samples, AT and AX were equal to 94 and 13 for C. tepejilote; and 89 and 8 for C. elatior. AT and AX decreased from unequal to equal sampling size, as would be expected since more than one-half of the Field Station's exclusive alleles (AX) were omitted. The distribution of exclusive, total and shared alleles is illustrated in Venn diagrams in Figure 2. Most loci had three or four alleles. After sample size standardization, the number of loci having three alleles increased but loci with four or five alleles decreased (Figure 3). Ae was substantially lower than A for each population and species (P<0.0001). Thus, alleles with low frequencies were numerous and contributed substantially to allelic diversity (AT). In the case of C. tepejilote, approximately 49–52.5% of the alleles had frequencies less than 0.21 and only 4–5% of the alleles had frequencies greater than 0.81 (Figure 4). Forty-three or 44 alleles (44 or 46%, respectively) had frequencies between 0.21 and 0.8. For C. elatior, the majority of alleles (57 or 62%) occurred at low frequencies, with 26–27% being higher than 0.61 and few alleles (11 or 15) with intermediate frequencies (Figure 4). The A values did not differ significantly between species, but those for Ae did (Z=4.52, P<0.0001). Mean Ae value was about 1.5 for C. elatior and 2.0 for C. tepejilote. The percent of polymorphic loci was very high for every population (Table 1). Only one locus, RUB-1, was monomorphic in all populations. In C. tepejilote, polymorphism ranged from 92.6 to 96.3%; and in C. elatior from 81.5 to 88.9. The highest population-specific level of monomorphism was observed in the eMP population: six out of 27 loci were monomorphic. High gene diversity in total population (HT) and within populations (HS) was observed for both C. elatior (HT=0.340 and HS=0.329) and C. tepejilote (HT=0.483 and HS=0.426), although both HT and HS were significantly greater in C. tepejilote (Mann–Whitney U-test, P<0.01).
Genetic structure
Allele frequency heterogeneity per locus among populations was significant at P<0.05 for most loci: 21 of the 26 loci from the three populations of C. elatior, and 23 of the 27 for those of C. tepejilote. Exact tests for population differentiation over all populations and loci were significant for each species at P<0.0001. The analyses of all possible pairwise population comparisons (1 vs 2, 1 vs 3 and 2 vs 3) also showed high levels of significance (P<0.0001). For C. elatior, 10 non-significant ‘ns’ loci and 16 significant ‘s’ loci between eFS and eSM populations were found; 10 ns and 16 s, between eFS and eMP; and 21 ns and 5 s for eSM and eMP. In the case of C. tepejilote, the non-significant and the significant numbers of loci for each pairwise comparison were: 7 ns and 20 s between tFS and tSM, 12 ns and 15 s between tFS and tMT, and 12 ns and 15 s between tSM and tMT. C. tepejilote showed higher values of Nei's (1978) genetic distance than did C. elatior (Table 1), the lowest value (0.007) of which occurred between eSM and eMP. This node was supported by 22 loci (81.5%). For C. tepejilote, the pair of tFS and tMT populations showed the lowest distance (0.037), with a consistency index of about 37% and 10 loci supporting this node. Nei's (1972) genetic similarity supports the idea that genetic heterogeneity between populations of C. tepejilote is larger than that between populations of C. elatior (Figure 5). For instance, the smallest genetic distance (0.0371) between the two geographically closest populations of C. tepejilote is higher than that between the farthest populations of C. elatior (0.0371 vs 0.0246), even when Euclidean distance (9481 vs 49 849 m) is almost fivefold longer for the latest populations (Table 1).
Nei's (1972) genetic similarity among populations of C. elatior and C. tepejilote at Los Tuxtlas, Mexico.
Based on a total of 156 fixation indexes calculated, excess homozygosity was indicated by significantly positive f estimates in 62 (79.5%) of 78 cases for the C. elatior populations and 58 (74.4%) of 78 for the C. tepejilote populations. Overall significant deficits of heterozygotes were found for every population. For both species, excess of homozygous individuals was further indicated by the positive FIT and FIS values of most loci (Table 2). For the C. elatior populations, θ=0.0315±0.005 and for the C. tepejilote populations, θ=0.152±0.035. In addition, the absolute difference among populations (DST) and the coefficient of relative differentiation (GST) were 0.011 and 0.033, respectively, for C. elatior; and 0.057 and 0.118 for C. tepejilote. Thus, most genetic variation (96.7 and 88.2%, respectively) was found within populations. The mean estimates of gene flow using GST and FST were from 1.4 to 1.9 for C. tepejilote and from 7.3 to 7.7 for C. elatior. Pairwise comparisons of DST, GST and Nm between species were significant (Mann–Whitney U-test, P<0.05).
Discussion
C. tepejilote and C. elatior showed very high levels of isozyme variation in comparison to values in other plants. Polymorphism and expected heterozygosity for C. tepejilote (P: 92.6–96.3, He: 0.385–0.442) and C. elatior (P: 81.5–88.9, He: 0.278–0.342) substantially exceed the corresponding average values (P: 65.5 and He: 0.180) observed for other long-lived perennial outcrossing species (Hamrick and Godt, 1997). Both high numbers of exclusive alleles and very large numbers of total alleles within populations were also found. Deficits of heterozygotes were also observed for most loci in all study populations. In contrast, less than one-tenth of the genetic variation occurred among populations, which would normally be taken to indicate high levels of gene flow. It is probably that other life history factors, such as the influence of biogeographic history and current gene dispersal have affected the standing genetic patterns of C. elatior and C. tepejilote.
Both theoretical and empirical studies provide reasons to expect that dioecy, perenniality and wide distributions contribute to minimize population differentiation and slow the processes of genetic erosion in C. elatior and C. tepejilote. C. tepejilote showed higher values of genetic variation within and among populations than C. elatior. Our findings are in agreement with both the highest P and the lowest GST values reported for long-lived outcrossing plants, as well as lower GST values documented for tall plants with low population density relative to plants with the opposite traits (Hamrick and Godt, 1997; Takeuchi et al., 2004). Similar to C. elatior, Neotropical populations of Swietenia humilis (White et al., 1999), Carapa guianensis (Dayanandan et al., 1999) and Symphonia globulifera (Aldrich et al., 1998) have also been shown to have low genetic differentiation and high allelic diversity. In addition to C. tepejilote, moderate genetic differentiation and high genetic variation within populations were reported for P. macroloba (Hall et al., 1994a), Swietenia macrophylla (Gillies et al., 1999; Lemes et al., 2003) and Caryocar brasiliense (Collevatti et al., 2001a). The larger number of exclusive alleles within C. tepejilote populations is consistent with the GST measures in indicating more-restricted gene flow in comparison to that of C. elatior.
It is important to compare these results to earlier findings on a very different spatial scale between contiguous adults within localized populations. It was found that while genetic autocorrelations were low in both species, values for C. elatior were somewhat greater than C. tepejilote (Luna et al., 2005). Consequently, at the localized scale (on the order of 100's of meters to a few kilometers), there is greater genetic differentiation for C. elatior than C. tepejilote, opposite the present results. The values corresponded to those expected for Wright's neighborhood sizes of 100 reproductive individuals for C. elatior and 300 reproductive individuals for C. tepejilote. If density were ignored, this finding would imply that dispersal distance is more limited in C. elatior than C. tepejilote. However, Wright's neighborhood is always relative to density, which differed greatly between the two species, being much higher in C. tepejilote. The areas occupied by neighborhoods are estimated at 1.0 Ha for C. tepejilote vs 13.2 Ha for C. elatior. Thus in the sense used by Wright (1943), the area over which there is effective gene flow is much larger in C. elatior. If one projects these results to the much greater spatial distances between the study populations examined here, then they are consistent with observed lesser differentiation for C. elatior. In other words, the almost threefold higher FST value among the populations of C. tepejilote can be explained by differences in the spatial scale over which the two species' neighborhood sizes are harbored. Finally, we note that criteria for obtaining unbiased values of population genetic differentiation for among-species comparisons remained unclear (Cole, 2003), although one relative measure of population genetic differentiation has been recently developed (Hedrick, 2005).
The spatial scale of sampling is typically related to the estimated magnitude of population structuring. Low genetic differentiation has been found for many tropical tree species sampled within a single natural population or among multiple populations separated by less than 100 km (O'Malley and Bawa, 1987; Hamrick and Loveless, 1989; Murawski and Bawa, 1994; Boshier et al., 1995; Hall et al., 1994b, 1996; Nason et al., 1996b), but high population differentiation has also been detected at similar geographical scales (Hall et al., 1994a). At a regional level, C. alliodora showed significant differences between Atlantic and Pacific populations, although the overall amount of genetic structure was low (Chase et al., 1995). For C. elatior and C. tepejilote, the significant genetic heterogeneity among populations supports the idea that local demes may deviate significantly from the population mean allele frequency even if genetic differentiation equals 0.5 (Wright, 1978).
In marked contrast to the general expectation and usual observation that the lowest levels of inbreeding in plants occur in obligate outcrossing species, large excesses of homozygotes were observed for both species, and these occurred consistently among loci and populations. Moreover, the levels of inbreeding are far higher than those that would be expected from simple forms of mating by spatial proximity (Epperson, 1990). Detailed demographic data from the permanent sites at the Field Station have demonstrated that both species blossom once a year (Oyama, 1990; Luna, 1999). In the case of C. tepejilote, although almost all adult plants reproduced every year, only a few individuals produced most of seeds and did so consistently during four successive years (Oyama, 1990). Moreover, strong winds that transport the copious pollen that is released during the storm season could enhance mating-pair heterogeneity. In any case, the strong reproductive dominance displayed by small numbers of individuals may greatly increase the genetic similarity among contemporary offspring. A contrasting pattern was apparent in C. elatior. Almost all fruits that developed were on palms that reproduced only one specific year. Most of reproductive palms (68%) flowered only once during the 4-year study period. The majority of seeds were produced by about one-tenth of the female plants. Given the small number of maternal plants, the seeds produced each year are highly related and, differ from other years. This mating pattern would contribute to mating-pair heterogeneity in the long term, which is otherwise unexpected for populations having small effective sizes.
The role of male reproduction is also important. For C. tepejilote, there is a unimodal temporal distribution of pollination activity (Otero-Arnaiz and Oyama, 2001) and some male individuals remained reproductively dominant (that is, produce larger numbers of male flowers) over the 4-year period (at least). These effects will combine to produce unequal individual contributions to the population genetic structure and even greater inbreeding. Such temporal restriction on gene flow within populations composed of a mixture of individuals that reproduce at different—and often heritable—times has been called ‘isolation by time’ (Hendry and Day, 2005).
Robertson (1964) examined the case where a population effectively splits into sublines and there is some mixing of the sublines. Within sublines, inbreeding and genetic drift are increased, but genetic drift of the entire population is decreased over the long term. The demographic data indicate that this type of population structure may occur in both study species, for slightly different reasons. The sublines would maintain both high levels of polymorphism and explain the homozygote excess (Sonesson and Meuwissen, 2001).
Various environmental heterogeneity and genetic factors have been proposed to explain interindividual variability in growth, survivorship and reproduction (Kohyama, 1981; Solbrig, 1981; Primack and Antonovics, 1982; Sarukhán et al., 1984; Oyama, 1990; Peters, 1991; Hendry and Day, 2005). Studies of five tropical arboreal species (Bullock, 1982; Piñero and Sarukhán, 1982) found high reproductive output for a minority of individuals. It was suggested that other individuals would later emerge as frequent reproducers. In the case of the local abundant species P. macroloba, Hall et al. (1994a) hypothesized that asynchronous reproduction resulted in a small genetic neighborhood. Degen et al. (2004) also found low effective density (1.3–1.6 trees/Ha) within a population of S. globulifera having a high adult density. In that study, and in Acacia melanoxylon (Muona et al., 1991), non-random mating enhanced by a limited number of pollen donors was suggested. For Pithecellobium elegans (Chase et al., 1996), temporal variation of individual flowering was reported, in which the pollen pool was dominated by some particular individuals that produced massive flowers during one period, but then were minimal contributors to the pool in other years. Recently, high levels of polymorphism together with significant levels of inbreeding have been demonstrated in microsatellite studies of the tropical plants S. globulifera (Aldrich et al., 1998; Degen et al., 2004), S. humilis (White et al., 1999), Melaleuca alternifolia (Rossetto et al., 1999), S. macrophylla (Lemes et al., 2003; Novick et al., 2003) and within juvenile cohorts of Elaeocarpus grandis (Rossetto et al., 2004). In addition, heterozygote deficits in tropical plants have been found in populations of Piper amalago (Heywood and Fleming, 1986), Austromyrtus spp. (Shapcott and Playford, 1996), Syzygium nervosum (Shapcott, 1998), Ancistrocladus korupensis (Foster and Sork, 1997), Capsicum annuum (Hernández-Verdugo et al., 2001), Escontria chiotilla (Tinoco et al., 2005) and the dioecious species Chamaedorea tuerckheimii, Carica papaya and Carica cauliflora (Oyama et al., unpublished data). Inbreeding was also detected in populations of the trees Pithecellobium pedicellare (O'Malley and Bawa, 1987) and Bertholletia excelsa (O'Malley et al., 1988). Inbreeding has often been attributable to self-fertilization or kin mating. Within populations showing flowering synchrony, a Wahlund effect as generator of nonrandom mating has sometimes been discounted (Novick et al., 2003). However, as we have suggested here, synchronization at the population level does not exclude reproductive variability at the individual level. In other cases, an excess of homozygotes was not initially attributed to inbreeding (Collevatti et al., 2001a), but was later confirmed (Collevatti et al., 2001b). Accordingly, long-term studies that assess the effects of individual reproductive variation on population genetic structure can reveal critical aspects of the mating system and their effects on the structure of genetic variation. As comparable analyses become available, the importance of reproductive isolation by time and its influence on levels of identity by descent should be clarified.
References
Aguilar AR (1986). El Género Chamaedorea Willd (Palmae) en el Estado de Veracruz. BSc thesis, Universidad Veracruzana, Mexico.
Aide TM, Rivera E (1998). Geographic patterns of genetic diversity in Poulsenia armata (Moraceae): implications for the theory of Pleistocene refugia and the importance of riparian forest. J Biogeogr 25: 695–705.
Aldrich PR, Hamrick JL, Chavarriaga P, Kochert G (1998). Microsatellite analysis of demographic genetic structure in fragmented populations of the tropical tree Symphonia globulifera. Mol Ecol 7: 933–944.
Bacon CD, Bailey CD (2006). Taxonomy and conservation: a case study from Chamaedorea alternans. Ann Botany 98: 755–763.
Boshier DH, Chase MR, Bawa KS (1995). Population genetics of Cordia alliodora (Boraginaceae), a neotropical tree. 2. Mating system. Am J Bot 82: 476–483.
Bullock SH (1982). Population structure and reproduction in the neotropical dioecious tree Compsoneura sprucei. Oecologia 55: 238–242.
Caron H, Dumas S, Marque G, Messier C, Bandou E, Petit RJ et al. (2000). Spatial and temporal distribution of chloroplast DNA polymorphism in a tropical tree species. Mol Ecol 9: 1089–1098.
Cavers S, Navarro C, Lowe AJ (2003). Chloroplast DNA phylogeography reveals colonization history of a neotropical tree, Cedrela odorata L. in Mesoamerica. Mol Ecol 12: 1451–1460.
Chamberlain JR (1998). Isozyme variation in Calliandra calothyrsus (Leguminosae): its implications for species delimitation and conservation. Am J Bot 85: 37–47.
Chase MR, Boshier DH, Bawa KS (1995). Population genetics of Cordia alliodora (Boraginaceae), a neotropical tree. 1. Genetic variation in natural populations. Am J Bot 82: 468–475.
Chase MR, Moller C, Kesseli R, Bawa KS (1996). Distant gene flow in tropical trees. Nature 383: 398–399.
Cheliak WM, Pitel JA (1984). Techniques for Starch Gel Electrophoresis of Enzymes from Forest Tree Species Information Report PI-X-42. Petawawa National Forestry Institute: Chalk River, Ontario. pp 1–49.
Cole CT (2003). Genetic variation in rare and common plants. Annu Rev Ecol Evol S 34: 213–237.
Collevatti RG, Grattapaglia D, Hay JD (2001a). Population genetic structure of the endangered tropical tree species Caryocar brasiliense, based on variability at microsatellite loci. Mol Ecol 10: 349–356.
Collevatti RG, Grattapaglia D, Hay JD (2001b). High resolution microsatellite based analysis of the mating system allows the detection of significant biparental inbreeding in Caryocar brasiliense, an endangered tropical tree species. Heredity 86: 60–67.
Dayanandan S, Dole J, Bawa K, Kesseli R (1999). Population structure delineated with microsatellite markers in fragmented populations of a tropical tree, Carapa guianensis (Meliaceae). Mol Ecol 8: 1585–1592.
Degen B, Bandou E, Caron H (2004). Limited pollen dispersal and biparental inbreeding in Symphonia globulifera in French Guiana. Heredity 93: 585–591.
Dick CW, Abdul-Salim K, Bermingham E (2003). Molecular systematic analysis reveals cryptic tertiary diversification of a widespread tropical rain forest tree. Am Nat 162: 691–703.
Ellstrand NC (1992). Gene flow among seed plant populations. New For 6: 241–256.
Epperson BK (1990). Spatial autocorrelation of genotypes under directional selection. Genetics 124: 757–771.
Epperson BK, Álvarez-Buylla ER (1997). Limited seed dispersal and genetic structure in life stages of Cecropia obtusifolia. Evolution 51: 275–282.
Epperson BK, Chung MG (2001). Spatial genetic structure of allozyme polymorphisms within populations of Pinus strobus (Pinaceae). Am J Bot 88: 1006–1010.
Epperson BK, Huang Z, Li TQ (1999). Measures of spatial structure in samples of genotypes for multiallelic loci. Genet Res 73: 251–261.
Foster PF, Sork VL (1997). Population and genetic structure of the West African rain forest liana Ancistrocladus korupensis (Ancistrocladaceae). Am J Bot 84: 1078–1091.
Garant D, Kruuk LEB, Wilkin TA, McCleery RH, Sheldon BC (2005). Evolution driven by differential dispersal within a wild bird population. Nature 433: 60–65.
Gillies ACM, Navarro C, Lowe AJ, Newton AC, Hernández M, Wilson J et al. (1999). Genetic diversity in Mesoamerican populations of mahogany (Swietenia macrophylla), assessed using RAPDs. Heredity 83: 722–732.
González E, Vogt R, Dirzo R (1997). La Historia Natural de Los Tuxtlas. UNAM: Mexico.
Graham A, Dilcher DL (1998). Studies in neotropical paleobotany. XII. A palynoflora from the Pliocene Rio Banano formation of Costa Rica and the Neogene vegetation of Mesoamerica. Am J Bot 85: 1426–1438.
Hall P, Chase MR, Bawa KS (1994a). Low genetic variation but high population differentiation in a common tropical forest tree species. Conser Biol 8: 471–482.
Hall P, Orrell LC, Bawa KS (1994b). Genetic diversity and mating system in a tropical tree, Carapa guianensis (Meliaceae). Am J Bot 81: 1104–1111.
Hall P, Walker S, Bawa KS (1996). Effect of forest fragmentation on genetic diversity and mating system in a tropical tree, Pithecellobium elegans. Conser Biol 10: 757–768.
Hamilton M (1999). Tropical tree gene flow and seed dispersal. Nature 401: 129–130.
Hamrick JL (1994). Distribution of genetic diversity in tropical tree populations: implications for the conservation of genetic resources. In: Lambeth CC, Dvorak W (eds). Resolving Tropical Forest Resource Concerns through Tree Improvement, Gene Conservation and Domestication of New Species. North Carolina State University Press: Raleigh, NC. pp 74–82.
Hamrick JL, Godt MJ (1997). Effects of life history traits on genetic diversity in plant species. In: Silvertown J, Franco M, Harper JL (eds). Plant Life Histories. Cambridge University Press: Cambridge. pp 102–118.
Hamrick JL, Loveless MD (1989). The genetic structure of tropical tree populations: associations with reproductive biology. In: Bock JH, Linhart YB (eds). Evolutionary Ecology of Plants. Westview Press: Boulder, CO. pp 129–146.
Hedrick PW (2005). A standardized genetic differentiation measure. Evolution 59: 1633–1638.
Henderson A, Galeano G, Bernal R (1995). Field Guide to the Palms of the Americas. Princeton University Press: New Jersey.
Henderson A (1986). A review of pollination studies in the Palmae. Bot Rev 52: 221–259.
Hendry AP, Day T (2005). Population structure attributable to reproductive time: isolation by time and adaptation by time. Mol Ecol 14: 901–916.
Hernández-Verdugo S, Luna-Reyes R, Oyama K (2001). Genetic structure and differentiation of wild and domesticated populations of Capsicum annuum (Solanaceae) from Mexico. Plant Syst Evol 226: 129–142.
Heywood JS, Fleming TH (1986). Patterns of allozyme variation in three Costa Rican species of Piper. Biotropica 18: 208–213.
Hodel DR (1992). Chamaedorea Palms: The Species and Their Cultivation. Allen Press: Kansas.
Kephart S (1990). Starch gel electrophoresis of plant isozymes: a comparative analysis of techniques. Am J Bot 77: 693–712.
Kohyama T (1981). Studies on the Abies populations of Mt. Shimagare. II. Reproductive and life history traits. Bot Mag Tokyo 95: 167–181.
Lavin M, Mathews S, Hughes C (1991). Chloroplast DNA variation in Gliricidia sepium (Leguminosae): intraspecific phylogeny and tokogeny. Am J Bot 78: 1576–1585.
Lefèvre F, Fady B, Fallour-Rubio D, Ghosn D, Bariteau M (2004). Impact of founder population, drift and selection on the genetic diversity of a recently translocated tree population. Heredity 93: 542–550.
Lemes MR, Gribel R, Proctor J, Grattapaglia D (2003). Population genetic structure of mahogany (Swietenia macrophylla King, Meliaceae) across the Brazilian Amazon: implications for conservation. Mol Ecol 12: 2875–2883.
Levene H (1949). On a matching problem arising in genetics. Ann Math Stat 20: 91–94.
Li CC, Horvitz DG (1953). Some methods of estimating the inbreeding coefficient. Am J Hum Genet 5: 107–117.
Loveless MD (1992). Isozyme variation in tropical trees: patterns of genetic organization. New Forests 6: 67–94.
Luna R (1999). Demografía y Genética Poblacional de Chamaedorea elatior en la Selva de Los Tuxtlas, Veracruz. BSc thesis, UNAM, Mexico.
Luna R, Epperson BK, Oyama K (2005). Spatial genetic structure of two sympatric neotropical palms with contrasting life histories. Heredity 95: 298–305.
Malécot G (1948). Les Mathématiques de l'Hérédité. Masson: Paris.
Miller MP (1997). Tools for Population Genetic Analyses (TFPGA) Version 1.3. Web http://herb.bio.nau.edu/~miller/tfpga.htm. Department of Biological Sciences, Northern Arizona University: Flagstaff, AZ, USA.
Muona O, Moran GF, Bell JC (1991). Hierarchical patterns of correlated mating in Acacia melanoxylon. Genetics 127: 619–626.
Murawski DA (1995). Reproductive biology and genetics of tropical trees from a canopy perspective.In: Lowman MD, Nadkarni NM (eds). Forest Canopies. Academic Press: San Diego, CA, USA. pp 457–493.
Murawski DA, Bawa KS (1994). Genetic structure and mating system of Stemonoporus oblongifolius (Dipterocarpaceae) in Sri Lanka. Am J Bot 81: 144–160.
Nason JD, Aldrich PR, Hamrick JL (1996b). Dispersal and the dynamics of genetic structure in fragmented tropical tree populations. In: Laurence WF, Bierregaard RO (eds). Tropical Forest Remnants: Ecology, Management, and Conservation of Fragmented Communities. University of Chicago Press: Chicago, IL. pp 304–320.
Nason JD, Herre EA, Hamrick JL (1996a). Paternity analysis of the breeding structure of strangler fig populations: evidence for substantial long-distance wasp dispersal. J Biogeogr 23: 501–512.
Nei M (1972). Genetic distance between populations. Am Nat 106: 283–292.
Nei M (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 9: 583–590.
Nei M (1987). Molecular Evolutionary Genetics. Columbia University Press: New York.
Novick RR, Dick CW, Lemes MR, Navarro C, Caccone A, Bermingham E (2003). Genetic structure of Mesoamerican populations of big-leaf mahogany (Swietenia macrophylla) inferred from microsatellite analysis. Mol Ecol 12: 2885–2893.
Olivieri I, Gouyon PH (1997). Evolution of migration rate and other traits: the metapopulation effect. In: Hanski IA, Gilpin ME (eds). Metapopulation Biology: Ecology, Genetics, and Evolution. Academic Press: San Diego, CA, USA. pp 293–323.
O'Malley DM, Bawa KS (1987). Mating system of a tropical rain forest tree species. Am J Bot 74: 1143–1149.
O'Malley DM, Buckley DP, Prance GT, Bawa KS (1988). Genetics of Brazil nut (Bertholletia excelsa Humb. & Bonpl.: Lecythidaceae). 2. Mating system. Theor Appl Genet 76: 929–932.
Otero-Arnaiz A, Oyama K (2001). Reproductive phenology, seed-set and pollination in Chamaedorea alternans H. A. Wendl. an understorey dioecious palm in a rainforest in Mexico. J Trop Ecol 17: 745–754.
Oyama K (1990). Variation in growth and reproduction in the neotropical dioecious palm Chamaedorea tepejilote. J Ecol 78: 648–663.
Oyama K (1991). Seed predation by a curculionid beetle on the dioecious palm Chamaedorea tepejilote. Principes 35: 156–160.
Oyama K (1992). Conservation and exploitation of tropical resou-rces: the case of Chamaedorea palms. Evol Trend Plant 6: 17–20.
Oyama K (1993). Conservation biology of tropical trees: demographic and genetic considerations. Environment Update 1: 17–32.
Oyama K (1997). Chamaedorea tepejilote. In: González E, Vogt R, Dirzo R (eds). La Historia Natural de Los Tuxtlas. UNAM: Mexico. pp 107–109.
Peters CM (1991). Plant demography and the management of tropical forest resources: a case study of Brosimum alicastrum in Mexico. In: Gómez-Pompa A (ed). Rain Forest Regeneration and Management. UNESCO: Paris. pp 265–272.
Piñero D, Sarukhán J (1982). Reproductive behaviour and its individual variability in a tropical palm, Astrocaryum mexicanum. J Ecol 70: 461–472.
Postma E, Noordwijk AJ (2005). Gene flow maintains a large genetic difference in clutch size at a small spatial scale. Nature 433: 65–68.
Primack RB, Antonovics J (1982). Experimental ecological genetics in Plantago. VII. Reproductive effort in populations of P. lanceolata L. Evolution 36: 742–752.
Quero H (1994). Flora de Veracruz. Listados florísticos de México. Instituto de Ecología AC. UNAM: Mexico.
Raymond ML, Rousset F (1995). An exact test for population differentiation. Evolution 49: 1280–1283.
Robertson A (1964). The effect of non-random mating within inbred lines on the rate of inbreeding. Genet Res 5: 164–167.
Rossetto M, Jones R, Hunter J (2004). Genetic effects of rainforest fragmentation in an early successional tree (Elaeocarpus grandis). Heredity 93: 610–618.
Rossetto M, Slade RW, Baverstock PR, Henry RJ, Lee LS (1999). Microsatellite variation and assessment of genetic structure in tea tree (Melaleuca alternifolia—Myrtaceae). Mol Ecol 8: 633–643.
Sarukhán J, Martínez-Ramos M, Piñero D (1984). The analysis of demographic variability at the individual level and its population consequences. In: Dirzo R, Sarukhán J (eds). Perspectives on Plan Population Ecology. Sinauer Associates: Sunderland, MA, USA. pp 83–106.
Shapcott A (1998). Vagile but inbred: patterns of inbreeding and the genetic structure within populations of the monsoon rain forest tree Syzygium nervosum (Myrtaceae) in northern Australia. J Trop Ecol 14: 595–614.
Shapcott A, Playford J (1996). Comparison of genetic variability in remnant and widespread rainforest understorey species of Austromyrtus (Myrtaceae). Biodivers Conserv 5: 881–895.
Sneath PH, Sokal RR (1973). Numerical Taxonomy. The Principles and Practice of Numerical Classification. Freeman: San Francisco, CA.
Sokal RR, Rohlf FJ (1979). Biometria. H Blume Ediciones: Madrid, Spain.
Sokal RR, Wartenberg DE (1983). A test of spatial autocorrelation analysis using an isolation-by-distance model. Genetics 105: 219–237.
Solbrig OT (1981). Studies on the population biology of the genus Viola. II. The effect of plant size on fitness in Viola sororia. Evolution 35: 1080–1093.
Soltis DE, Soltis PS (1989). Isozymes in Plant Biology. Dioscorides Press: Portland, OR, USA.
Sonesson AK, Meuwissen THE (2001). Minimization of rate of inbreeding for small populations with overlapping generations. Genet Res 77: 285–292.
Stuber CW, Wendel JM, Goodman MM (1988). Techniques and Scoring Procedures for Starch Gel Electrophoresis of Enzymes from Maize (Zea mays) Technical Bulletin 286. North Caroline State University, North Caroline.
Takeuchi Y, Ichikawa S, Konuma A, Tomaru N, Niiyama K, Lee SL et al. (2004). Comparison of the fine-scale genetic structure of three dipterocarp species. Heredity 92: 323–328.
Thomas MM, Garwood NC, Baker WJ, Henderson SA, Russell SJ, Hodel DR et al. (2006). Molecular phylogeny of the palm genus Chamaedorea, based on the low-copy nuclear genes PRK and RPB2. Mol Phyl Evol 38: 398–415.
Tinoco A, Casas A, Luna R, Oyama K (2005). Population genetics of Escontria chiotilla in wild and silvicultural manager populations in the Tehuacán Valley, Central Mexico. Gene Resour Crop Ev 52: 525–538.
Trejo-Pérez L (1989). Diseminación de semillas por aves en Los Tuxtlas, Veracruz. In: Gómez-Pompa A, Del Amo S, Vázquez-Yanes C, Butanda A (eds). Regeneración de Selvas. Continental: Mexico. pp 447–470.
Vovides AP, Garcia-Bielma MA (1994). A study of the in situ situation of four species of threatened understory palms of the genus Chamaedorea in the wild in the state of Veracruz, Mexico. Principes 38: 109–113.
Uhl NW, Dransfield J (1987). Genera Palmarum: A classification of palms based on the work of HE Moore, Jr. International Palm Society and LH Bailey Hortorium: Lawrence.
Weir BS, Cockerham CC (1984). Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wendel J, Weeden N (1989). Visualization and interpretation of plant isozyme. In: Soltis DE, Soltis PS (eds). Isozymes in Plant Biology. Dioscorides Press: Oregon. pp 5–45.
White G, Boshier DH, Powell W (1999). Genetic variation within a fragmented population of Swietenia humilis Zucc. Mol Ecol 8: 1899–1909.
Workman PL, Niswander JD (1970). Population studies on southwestern Indian tribes. II. Local genetic differentiation in the Papago. Am J Hum Gen 22: 24–49.
Wright S (1943). Isolation by distance. Genetics 28: 114–138.
Wright S (1951). The genetical structure of populations. Ann Eugen 15: 322–354.
Wright S (1978). Evolution and the genetics of populations. Variability within and among Natural Populations, Vol. 4. University of Chicago Press: Chicago, IL.
Yeh FCH, O'Malley D (1980). Enzyme variations in natural populations of Douglas fir, Pseudotsuga menziesii (Mub.) Franco, from British Columbia. 1. Genetic variation patterns in coastal populations. Silvae Genet 29: 83–92.
Zarco-Espinosa VM (1999). Patrones Biogeográficos y Filogeográficos del Género Chamaedorea (Palmae). BSc thesis, UNAM, Mexico.
Acknowledgements
We are grateful to Miguel Martínez-Ramos, Faustino Sánchez-Garduño, Mark Olson and two anonymous reviewers for their insightful suggestions, Víctor Zarco-Espinosa and Euler Pedraza-Ortega for providing field collaboration and improving the figures, Daniel Piñero for lending us his laboratory to carry out the initial electrophoresis assays, Gerardo Rivas-Lechuga for assisting in statistical analyses, and Gabriela Guerrero-Pacheco and Luis Giovanni-Ramírez for georeferencing the studied populations. This research was supported by CONABIO and UNAM projects to KO; RL was sponsored by doctoral scholarships from CONACYT and DGEP-UNAM.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luna, R., Epperson, B. & Oyama, K. High levels of genetic variability and inbreeding in two Neotropical dioecious palms with contrasting life histories. Heredity 99, 466–476 (2007). https://doi.org/10.1038/sj.hdy.6801027
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6801027
Keywords
This article is cited by
-
High gene flow through pollen partially compensates spatial limited gene flow by seeds for a Neotropical tree in forest conservation and restoration areas
Conservation Genetics (2021)
-
Contrasting patterns of genetic diversity between two endangered palms with overlapping distributions, Voanioala gerardii (Arecoideae) and Lemurophoenix halleuxii (Arecoideae), from North-east Madagascar
Conservation Genetics (2012)
-
Conservation genetics and ecology of an endemic montane palm on Lord Howe Island and its potential for resilience
Conservation Genetics (2012)
-
Disturbance and Resilience in Tropical American Palm Populations and Communities
The Botanical Review (2011)
-
Population genetics of the understory fishtail palm Chamaedorea ernesti-augusti in Belize: high genetic connectivity with local differentiation
BMC Genetics (2009)