Abstract
Neural tube defects (NTDs), such as spina bifida (SB) or exencephaly, are common congenital malformations leading to infant mortality or severe disability. The etiology of NTDs is multifactorial with a strong genetic component. More than 70 NTD mouse models have been reported, suggesting the involvement of distinct pathogenetic mechanisms, including faulty cell death regulation. In this review, we focus on the contribution of functional genomics in elucidating the role of apoptosis and autophagy genes in neurodevelopment. On the basis of compared phenotypical analysis, here we discuss the relative importance of a tuned control of both apoptosome-mediated cell death and basal autophagy for regulating the correct morphogenesis and cell number in developing central nervous system (CNS). The pharmacological modulation of genes involved in these processes may thus represent a novel strategy for interfering with the occurrence of NTDs
Similar content being viewed by others
Main
Neural tube defects (NTDs) are common (1 in 1000 pregnancies) congenital malformations in humans leading to infant mortality or severe disability. NTD results from failure of complete neurulation during the fourth week of embryogenesis. The etiology of NTDs is complex, with both environmental and genetic contributions.1 In particular, a strong genetic component, accounting for at least 30% of NTDs, has been demonstrated, although only few of the genes involved have been structurally and functionally characterized.2 This is mainly due to the fact that neurulation is a complex multistep process involving precise temporal and spatial regulation of gene expression.3, 4, 5, 6 Animal models are providing insight into the mechanisms participating in correct neurulation or in its failure.7, 8, 9 More than 70 mouse models exhibiting NTDs have been described.10, 11, 12, 13 Mutations display wide locus heterogeneity, and for many of them, the corresponding human homolog has been found.14, 15 These studies suggest the involvement of distinct molecular pathways in NTD pathogenesis, ranging from dysregulation of homocysteine/folate metabolism to defective cell proliferation or death, to disruption of cytoskeleton and migratory capabilities and to faulty pyrimidine synthesis.8, 9, 16, 17 Among these, special attention is being devoted to apoptosis. Apoptosis plays an important role in the morphogenesis and homeostasis of the developing central nervous system (CNS) by participating in folding, pinching off and fusion of neural walls, in neural precursor selection and in postmitotic competition of neurons for their cellular targets (reviewed in De Zio et al.,18Hidalgo and ffrench-Constant,19 Kuan et al.,20 and Nicotera21). Notably, the main phenotype, which is induced by disruption of apoptosis-related genes regards the developing nervous system (see Table 1). Autophagy, an important mechanism for degrading long-lived proteins and the only known cellular pathway for degrading organelles, has been originally linked to the occurrence of pathological conditions in the nervous system associated with alternative cell death morphotypes.43, 44, 45 Recently, autophagy has been recognized as a survival process46 and as a key player in vertebrate neurodevelopment;47 also, a complex interplay between autophagy and apoptosis has emerged (reviewed in Ferraro and Cecconi48 and Maiuri et al.49). Interestingly, comparative analysis of the phenotypes exhibited by mouse models’ defective in apoptosis or autophagy genes show several similitaries, although molecular analysis revealed that they are accounted for by distinct roles at different developmental stages. The data we discuss here suggest that, whereas apoptosis is the main mechanism by which mammals can prevent the survival of excess cells in the developing nervous system, autophagy regulation may have a crucial role in the ontogenesis of CNS through a mechanism involving the need for cell ‘renovation’ in differentiation and cell cycle control rather than for cell demise.
The Basis of Neural Development in Vertebrates
The development of vertebrates from the zygote follows a specific sequence of events: (i) division of the zygote into undifferentiated blastula cells, (ii) formation of three progenitor cell layers in a gastrula, (iii) differentiation and reorganization of the progenitor cells into tissues and organs, (iv) full growth of the organs and (v) birth.
In the course of development, cell differentiation begins with the emergence of the cells in the three primordial layers of the gastrula: the ectoderm (outer layer), the mesoderm (middle layer) and the endoderm (inner layer). Roughly speaking, the gastrula ectoderm develops into the skin, the sense organs, and the nervous system. The progenitor cells for all neurons and glial cells of CNS begin as a further differentiation of ectoderm cells into a layer known as the neural plate (see Smith and Schoenwolf5 and schematics in Figure 1). The neural plate folds and differentiates into neural crest cells and a neural tube. The neural plate in its unvaginate state has a series of cell linings along its periphery. These cell linings persist during the formation of the neural tube, thus forming two longitudinal structures on the dorsal side of the neural tube, called the neural crests. Neural plate formation is induced by chemical signals from the mesoderm. The neural crest contributes to many cell types, among which are the cells of the peripheral nervous system, whereas the neural tube becomes the CNS. The cells that constitute the neural tube go on to form neurons, astrocytes and oligodendrocytes. Once the neural tube is closed at both extremes, namely the rostral and caudal neuropores, the cavity that is formed inside the neural tube will eventually form the ventricles of the encephalon and the central canal of the spinal cord. The anterior portion of the neural tube is then divided into three distinct, interconnected rounded cavities, which distinguish the portion that will become the encephalon, whereas the middle and caudal portions remain as a tube and will become the spinal cord. The most anterior cavity is called the forebrain or prosencephalon, the second is the midbrain or mesencephalon and finally the most posterior one is the hindbrain or rhombencephalon. Later, in development, the CNS will go through a stage termed ‘five vesicles’, namely telencephalon, diencephalon, mesencephalon, metencephalon and myelencephalon. From the telencephalon originates the cerebral cortex, basal ganglia, hippocampal formation, amygdala and olfactory bulb, and from the diencephalon the thalamus and surrounding nuclei, hypothalamus, retina and optic nerve.
Schematic diagram of neural plate bending and neural tube formation in vertebrates. Neural folds form at the lateral extremes of the neural plate (a), elevate (b) and converge toward the dorsal midline (c) to form the neural tube (d). not, notochord; nf, neural folds; np, neural plate; nt, neural tube; nc, neural crest cells; se, surface ectoderm
Neural Tube Defects
Abnormalities in neural tube formation result in NTDs and both environmental and genetic factors can lead to NTDs in 1 in every 1000 births and cause 1 in 20 of every spontaneous abortion.7, 50 There are different types of NTDs, depending on the region of the CNS that is affected. The most known and studied types are called the open NTDs. Open NTDs occur when the brain and/or spinal cord are exposed at birth through a defect in the skull or vertebrae. Examples of open NTDs are the spina bifida (SB; also termed myelomeningocele), anencephaly and encephalocele.7, 50 Cranioschisis is the failure of proper fusion of the cephalic part of the neural tube, which leaves the brain and cranium open. The most extreme case results in anencephaly. Nonclosure of the posterior neuropore in the lumbosacral region leads to SB. SB may cause paralysis of the legs, anesthesia of the skin, urologic disturbances and defecatory dysfunctions.
Rare types of NTDs are called closed NTDs. Closed NTDs occur when the spinal defect is covered by skin. Common examples of closed NTDs are lipomyelomeningocele, lipomeningocele and tethered cord. SB occulta is potentially another form of an NTD in which there is a typically benign bony change in one or more vertebrae, but which does not involve the nerves within the spinal column.50
Our understanding of the causes of NTDs is incomplete, and the identification of preventive measures is very limited. Direct analysis of human embryos is not possible, owing to both practical and ethical considerations, and there are few families available that would be suitable for genetic linkage studies. Another approach for gaining insight into the causes of human NTDs is provided through studies on the mice. Over 70 mouse mutants have been identified that exhibit NTDs, and they are providing important information on the cellular and molecular basis of neural tube formation.10, 11, 12, 13
Functional Genomics Approaches to Unravel NTDs Ontogenesis
Using the gene targeting approach, mutants have been generated for a huge variety of genes and their function widely demonstrated. This methodology depends a priori on the isolation and molecular analysis of a given gene. As an example, the specific inactivation through gene targeting of particular genes, such as transcriptional factors or signaling molecules, has shed new light on the functional complexity of mammalian development. Various strategies have been used to isolate such genes, starting from the classic genetic studies in lower organisms, such as Drosophila melanogaster and Caenorhabditis elegans. The Otx genes, for example, which are required for proper brain and sensory organ development were initially isolated from the mouse genome on the basis of their sequence similarities to conserved sequence motifs within homologous genes of Drosophila.51 However, the vast majority of genes present in the mouse genome have not yet been structurally characterized, which precludes their disruption by homologous recombination. To identify and mutate new murine genes, an excellent experimental strategy is the gene trap. In our laboratory and in many others, several genes important for cell metabolism and embryonic development have been captured and functionally analyzed in this way.52 Both apoptosis and autophagy typically occur through ordered, programmed series of events whose biochemical machinery has been largely identified. By using combined gene targeting and gene trap approaches, it has been possible to verify the role of specific apoptotic or autophagic pathways in neurulation.
Mouse Models of Reduced or Abolished Cell Death in Neurodevelopment
Apoptosis is principally regulated by the Bcl2 family of proteins (proapoptotic and antiapoptotic molecules), the adaptor protein Apaf1 and the cysteine protease caspase family (for a recent review, see Maiuri et al.49). During the development of the nervous system, neurons share the same basic apoptosis program with all other cell types. However, at various developmental stages, to provide the specificity of regulation, the distinct neuronal populations express different combinations of Bcl2 and caspase family members.18, 53
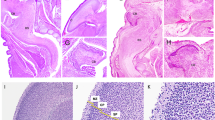
The analysis of various knockout mice has led to an increased knowledge of how the various apoptotic factors are involved in neuronal cell death (see Table 1). Apaf1 plays a fundamental role in the apoptotic machinery (Figure 2). It is activated upon cytochrome c release from mitochondria and interacts with dATP/ATP and procaspase 9, thereby forming the apoptosome, the core of the apoptotic program.48, 54 The apoptosome activates effector caspases, such as caspases 3, 6 and 7, which execute cell death. Apaf1 knockout and gene trap mice32, 33 show an abnormal phenotype, which involves many tissues and organs: severe craniofacial malformations, brain hyperplasia and alterations of the limb, ear and eye structures (see Figure 3). Targeted gene disruptions of caspases 3 and 9 in mice26, 27, 28, 29 lead to decreased neuronal apoptosis and neurodevelopmental abnormalities including ectopic and duplicated neuronal structures, often culminating in a peculiarly convoluted brain. Owing to these abnormalities, caspase 9 and caspase 3 knockouts show embryonic lethality. This particular phenotype may be due to the disproportionate number of neural precursor cells (NPCs), which do not die and thus undergo differentiation, so providing supernumerary cells. It should be mentioned that this neural phenotype appears in a small percentage of caspase 3−/− embryos (∼10%) and in most caspase 9−/− embryos (∼97%), and strictly depends on the genetic background.55, 56 The similar phenotypes of null mutations of caspase 9 and caspase 3 embryos suggest that these two caspases might participate in the same apoptosis pathway during neurodevelopment. In fact, activated caspase 9 can cleave and activate caspase 3 in the mitochondrial pathway of cell death. However, despite the severe defects of apoptosis in the brain, the developmental apoptosis of other structures in the caspase 3 and caspase 9 null mutants is largely preserved. This means that these two proteases regulate district-specific apoptosis, indicating that individual caspases have a dominant and non-redundant role in apoptosis in a tissue-selective or stimulus-dependent manner. It should be said that the Apaf1 knockout does not precisely mimic caspase 3 and caspase 9 knockouts, which exhibit a predominantly neuronal phenotype. The number of developmental alterations observed in Apaf1-deficient embryos is higher than in other mutants and they are also more distributed all over the organism.32, 33, 55 However, as well as for caspase 3 and 9, the importance of Apaf1 during neurogenesis lies especially in the regulation of apoptosis of NPCs. Indeed, Apaf1-deficient embryos exhibit marked hyperplasia of the embryonic ventricular zone where NPCs are confined, this resulting in an evident exencephaly (see Figure 3). The NPCs, which should undergo apoptosis and are devoid of the apoptosome machinery proliferate generating both protruding forebrain masses and in some cases, such as in the hindbrain, differentiating immature neurons. The capability of Apaf1-deficient cells to proliferate seems to be guaranteed by an autophagy-dependent mechanism, that has been demonstrated to be based on the selective survival of a few intact mitochondria, which (a) retain cytochrome c, (b) will be spared from autophagosome degradation and (c) could sustain ATP generation by oxidative phosphorylation.57 The result of Apaf1 deficiency in the developing nervous system is thus the inhibition of proper morphogenesis of the neural tube, which often leads to the generation of SB and exencephaly, the most common NTDs.32
Autophagosome formation and apoptosome-mediated cell death in neurodevelopment. (a) Autophagosome nucleation is driven by phosphatidylinositol (PI) phosphorylation. This process is mediated by a lipid kinase signaling complex (Beclin 1, Vps15, Vps34). Ambra1 favors Vps34/Beclin 1 interaction, whereas ATG1/Ulk1 is dowsntream of mTOR (see text) and is involved in autophagy induction. UVRAG and Bif-1 have been described as additional Beclin 1 complex regulators. Beclin 1 proautophagic roles are inhibited by its binding to Bcl2 and/or Bcl-XL, which act in the crosstalk between autophagy and apoptosis. (b) Bcl2-like pro-apoptotic and antiapoptotic proteins regulate cytochrome c release from mitochondria. Cytosolic cytochrome c binds Apaf1 and induces the recruitment of the initiator caspase 9 (Casp9) on the active apoptosome (see text). The active apoptosome, in turn, activates caspase 3 (Casp3), which mediates cell destruction. The relative roles of the two pathways in neurodevelopment are indicated. Notably, they are both required for neural tube closure
Comparison of the neurodevelopmental knockout phenotypes of a proapoptotic gene (Apaf1) versus a proautophagic gene (Ambra1). The upper panels (a, b and c) show the head of an E12.0 mouse embryos from both homozygous genotypes and compared with the wild type. The embryos were microtome-sectioned through a transverse plan and the section fixed and haematoxylin/eosin stained (middle panels, d, e and f). Finally, cell death by apoptosis on adjacent sections was detected in the rostral spinal cord by the TUNEL method (g, h and i). The arrows point to a few autofluorescent blood red cells (unspecific signal). Autophagy inactivation leads to excessive apoptosis, at variance with apoptosome deficiency. However, the phenotypes of both knockout embryos are strikingly similar and result in dramatic NTDs (see arrows in b and c)
The proapoptotic member of Bcl2 family, Bax, also regulates neural apoptosis during CNS development (Figure 2).22, 23 Bax is highly expressed in both the embryonic and the adult nervous system and Bax mutant mice show reduced apoptosis of target-derived, neurotrophic factor-dependent neuronal subpopulations such as spinal cord motor neurons and sympathetic neurons in the peripheral nervous system. In contrast to Apaf1 and caspase knockouts, Bax−/− mice do not show embryonic lethality or any evident neuronal overgrowth, as Bax regulates the neuronal apoptosis affecting only the postmitotic neurons, which are developing their synaptic connections. It has been also observed at early developmental stages that Bax deficiency leads to an increased number of immature postmitotic neurons confined to the intermediate and marginal zones.22, 23 Thus, Bax modulates the neural apoptosis, which involve the young neurons rather than their undifferentiated precursor cells.
Mouse Models of Reduced Autophagy in Neurodevelopment
Autophagy is a self-degradative process involved both in basal turnover of cellular components and in response to nutrient starvation or organelle damage in a wide range of eukaryotes.58 During autophagy, portions of the cytoplasm are sequestered by double-membraned vesicles called autophagosomes, and are degraded after fusion with lysosomes for subsequent recycling.59 Autophagy can be divided into three stages: initiation, execution and maturation.60 Autophagy initiation is mainly controlled by the Beclin 1-class III phosphatidylinositol 3-kinase (PI3K)/Vps34 complex,61 whose activity could be positively or negatively regulated by the interaction with different cofactors such as Ambra1, UVRAG, Bif-1 and Bcl2 (see Figure 2).38, 62, 63, 64 Autophagy initiation is stimulated by many different intracellular or extracellular stress stimuli.65 The classical autophagy inducer is represented by amino-acid deprivation, which leads to inhibition of mTOR, a protein kinase central to nutrient-sensing signal transduction, regulation of translation and cell cycle progression control.66 Autophagy can also be induced by an mTOR-independent route by lowering myo-inositol-1,4,5-triphosphate (IP3) levels.67 Autophagy execution involves autophagosomal vescicle formation and is mediated by two covalent-conjugation pathways: the covalent linkage of ATG5 and ATG12, and the covalent lipidation of ATG8/LC3 by phosphatidylethanolamine.68 These post-translational modifications allow, in both cases, the translocation of these proteins to the nascent autophagosomal membrane. Finally, during the maturation step, autophagosomes fuse with endosomal vesicles and acquire lysosome-associated membrane protein 1 (Lamp1) and Lamp2, becoming amphisomes, which, in turn, fuse with lysosomes that contain cathepsins and acid phosphatases necessary for protein degradation.69
In vertebrates, autophagy acts as a prosurvival or prodeath mechanism in many physiological and pathological conditions.70 A lot of evidence underlines the vital role that autophagy plays in the function of the nervous system.71 Morphological features of autophagy have been detected both during neural development and, in adult brain, in various neurodegenerative diseases.72 More recently, genetic inactivation in the mouse model of genes involved in autophagy have formally demonstrated their importance in the nervous system (see Table 1)47.
A fundamental role of Beclin 1 during embryogenesis is demonstrated by the fact that Beclin 1-deficient mice die very early during development.36, 73 Premature death of Beclin 1 mutant mice did not make it possible to assess the specific role of Beclin 1 during neurogenesis. However, recent in vitro studies indicate that Beclin 1 is required for retinoic acid-induced differentiation of neuroblastoma N2a cells.40 A definitive demonstration of the role of the autophagy-regulating genes during neural development is due to the identification of Ambra1 (activating molecule in Beclin 1-regulated autophagy), a WD40 containing protein, which interacts with Beclin 1 and positively regulates its ability to stimulate VPS34 kinase activity (Figure 2).38
At early stages during development, Ambra1 expression is restricted to the nervous system. At embryonic day (E)8.5, strong staining was detected throughout the neuroepithelium. At E11.5, a robust expression was observed in the ventralmost part of the spinal cord, the encephalic vesicles, the neural retina, the limbs and the dorsal root ganglia. At later developmental stages, the expression became abundant throughout the developing nervous system as well as in other tissues. The main phenotype of the Ambra1 loss-of-function homozygous embryos is an apparent overgrowth of the nervous system, culminating in an accentuated mid-hindbrain exencephaly often associated with SB (see Figure 3). Importantly, basal autophagy appears to be reduced in Ambra1 mutant embryos. This was assessed either by localization of GFP-LC3-positive dots within neuroepithelial cells or LC3 cleavage to form II. Moreover, impairment of autophagy was corroborated by the presence in the same cells of a strong ubiquitin staining in the mutant Ambra1 embryos, a hallmark of defective basal autophagy in vivo.
Ambra1 mutant mice also showed an increased proliferation at E8.5, followed by excessive apoptosis from E9.0 onwards. In vitro experiments support the view that autophagy inhibition is directly linked to uncontrolled proliferation.38 Indeed, modulation of Ambra1 expression by means of overexpression or RNA interference leads to altered rates of cell proliferation in a Beclin 1-dependent manner. These results are in line with the well established role of other components of the class III PI3K complex, such as Beclin 1, UVRAG and Bif-1, in the regulation of proliferation and tumor growth.38, 62, 64, 73 Taken together, these data suggest that autophagy initiation is devoted to the control of both autophagosome formation and cell proliferation. Whether and how these two processes are mechanistically linked, or are independently regulated by these proteins, is an important issue that remains to be elucidated.
The protein serine/threonine kinases ATG1 (also called unc-51.1 or ULK1), a protein whose role in autophagy initiation has been well established in yeast,41 has been reported as one of the earliest genes in neuronal differentiation and is required for granule cell axon formation.42 The protein is localized to both axonal shafts and growth cones of extending axons and is essential for neurite extension/parallel fiber formation in cerebellar granule neurons, in vitro. It has not yet been proved whether detective autophagy is responsible for the observed defects.
Mice deficient for ATG5 specifically in neural cells develop progressive deficits in motor function and die a few months after birth.35 In ATG5−/− neurons, autophagy inhibition leads to an abnormal accumulation of ubiquitinated proteins, which accumulate and over time form aggregates and inclusions. Although no major morphological defects were reported during neural development, an increase in ubiquitinated proteins was detected during embryogenesis starting from E15.5. Thus, basal autophagy is important for preventing the accumulation of abnormal proteins during nervous system development, which can disrupt neural function and ultimately lead to neurodegeneration in adults.
Lamp-1/Lamp-2 double-deficient mouse embryos die in utero at stage E16.5, exhibiting predominantly craniofacial abnormalities.39 Besides cartilage and epithelial malformation, this phenotype includes the foreshortening of the forebrain. Although lysosome impairment alongside autophagy maturation could account for the observed defects, the abnormal presence of cytoplasmic autophagic vacuoles in the neuroepithelium of double knockout mice suggests that autophagy inhibition could be a consequence of Lamp-1/Lamp-2 inactivation.
Taken together, these data indicate that, at variance with the Ambra1 mutant, most autophagy-related phenotypes in the CNS include morphological defects associated with the late steps of the nervous system development, that is, axonal growth impairment or abnormalities of differentiating neurons, rather than common NTDs. This evidence parallels the data regarding apoptosis gene targeting, which showed that NTDs mainly result from functional alteration of molecules involved in the control of NPC survival.
Conclusions
An important conclusion could be drawn by direct comparison of two of the above-described phenotypes. As evident from Figure 3, the NTD phenotypes exhibited from Apaf1 (a proapoptotic factor) and Ambra1 (a proautophagic factor) gene trap mutant embryos are quite similar. However, a striking difference is visible when detecting apoptosis by the TUNEL method, with Apaf1 mutant sections being negative and Ambra1 mutant sections being strongly positive. In both cases, the consequence of the phenotype is a hyperplasic neural tube, generated by excessive cell proliferation. In the autophagic mutants, NPC proliferation is directly accelerated and cell death attempts inefficiently to compensate for the increase in cell number. In the apoptotic mutants, NPCs selection does not occur properly, leading to the accumulation of supernumerary precursors that continue to grow without control. Therefore, in the developing nervous system, specific regulators of apoptosis or autophagy are involved in cell growth control of NPCs in a direct or indirect manner and their inactivation results, in the end, to the lack of neural tube closure (Figure 2). Understanding the precise roles of autophagy and apoptosis and unraveling their complex interplay in neurodevelopment could make it possible to hypothesize their pharmacological modulation in vivo. This might have an important role in the prevention and treatment of human NTDs.
Abbreviations
- Ambra1:
-
activating molecule in Beclin 1-regulated autophagy
- ATG:
-
autophagy-related gene
- CNS:
-
central nervous system
- E:
-
embryonic day
- IP3:
-
myo-inositol-1,4,5-triphosphate
- Lamp:
-
lysosome-associated membrane protein
- NPC:
-
neuronal precursor cell
- NTDs:
-
neural tube defects
- PI3K:
-
phosphatidyl Inositol 3-kinase
- SB:
-
spina bifida
- TUNEL:
-
terminal uridine deoxynucleotidyl transferase dUTP nick end labeling
References
Eskes TK . From birth to conception. Open or closed. Eur J Obstet Gynecol Reprod Biol 1998; 78: 169–177.
Gibson KM, Bottiglieri T . Genetic predisposition to neural tube defects? Pediatr Res 2000; 48: 135.
Colas JF, Schoenwolf GC . Towards a cellular and molecular understanding of neurulation. Dev Dyn 2001; 221: 117–145.
Rogner UC, Spyropoulos DD, Le Novere N, Changeux JP, Avner P . Control of neurulation by the nucleosome assembly protein-1-like 2. Nat Genet 2000; 25: 431–435.
Smith JL, Schoenwolf GC . Neurulation: coming to closure. Trends Neurosci 1997; 20: 510–517.
Tanabe Y, Jessell TM . Diversity and pattern in the developing spinal cord. Science 1996; 274: 1115–1123.
Copp AJ, Brook FA, Estibeiro JP, Shum AS, Cockroft DL . The embryonic development of mammalian neural tube defects. Prog Neurobiol 1990; 35: 363–403.
Corcoran J . What are the molecular mechanisms of neural tube defects? Bioessays 1998; 20: 6–8.
Fleming A, Copp AJ . Embryonic folate metabolism and mouse neural tube defects. Science 1998; 280: 2107–2109.
Juriloff DM, Gunn TM, Harris MJ, Mah DG, Wu MK, Dewell SL . Multifactorial genetics of exencephaly in SELH/Bc mice. Teratology 2001; 64: 189–200.
Juriloff DM, Harris MJ . Mouse models for neural tube closure defects. Hum Mol Genet 2000; 9: 993–1000.
Murdoch JN, Rachel RA, Shah S, Beermann F, Stanier P, Mason CA et al. Circletail, a new mouse mutant with severe neural tube defects: chromosomal localization and interaction with the loop-tail mutation. Genomics 2001; 78: 55–63.
van Straaten HW, Copp AJ . Curly tail: a 50-year history of the mouse spina bifida model. Anat Embryol (Berl) 2001; 203: 225–237.
Harris MJ, Juriloff DM . Genetic landmarks for defects in mouse neural tube closure. Teratology 1997; 56: 177–187.
Stegmann K, Boecker J, Richter B, Capra V, Finnell RH, Ngo ET et al. A screen for mutations in human homologues of mice exencephaly genes Tfap2alpha and Msx2 in patients with neural tube defects. Teratology 2001; 63: 167–175.
Harris MJ, Juriloff DM . Mini-review: toward understanding mechanisms of genetic neural tube defects in mice. Teratology 1999; 60: 292–305.
Herrera E, Samper E, Blasco MA . Telomere shortening in mTR−/− embryos is associated with failure to close the neural tube. EMBO J 1999; 18: 1172–1181.
De Zio D, Giunta L, Corvaro M, Ferraro E, Cecconi F . Expanding roles of programmed cell death in mammalian neurodevelopment. Semin Cell Dev Biol 2005; 16: 281–294.
Hidalgo A, ffrench-Constant C . The control of cell number during central nervous system development in flies and mice. Mech Dev 2003; 120: 1311–1325.
Kuan CY, Roth KA, Flavell RA, Rakic P . Mechanisms of programmed cell death in the developing brain. Trends Neurosci 2000; 23: 291–297.
Nicotera P . Development and death of neurons: sealed by a common fate? Cell Death Differ 2002; 9: 1277–1278.
Deckwerth TL, Elliott JL, Knudson CM, Johnson Jr EM, Snider WD, Korsmeyer SJ . BAX is required for neuronal death after trophic factor deprivation and during development. Neuron 1996; 17: 401–411.
White FA, Keller-Peck CR, Knudson CM, Korsmeyer SJ, Snider WD . Widespread elimination of naturally occurring neuronal death in Bax-deficient mice. J Neurosci 1998; 18: 1428–1439.
Shindler KS, Latham CB, Roth KA . Bax deficiency prevents the increased cell death of immature neurons in bcl-x-deficient mice. J Neurosci 1997; 17: 3112–3119.
Lindsten T, Golden JA, Zong WX, Minarcik J, Harris MH, Thompson CB . The proapoptotic activities of Bax and Bak limit the size of the neural stem cell pool. J Neurosci 2003; 23: 11112–11119.
Haydar TF, Kuan CY, Flavell RA, Rakic P . The role of cell death in regulating the size and shape of the mammalian forebrain. Cereb Cortex 1999; 9: 621–626.
Kuida K, Haydar TF, Kuan CY, Gu Y, Taya C, Karasuyama H et al. Reduced apoptosis and cytochrome c-mediated caspase activation in mice lacking caspase 9. Cell 1998; 94: 325–337.
Kuida K, Zheng TS, Na S, Kuan C, Yang D, Karasuyama H et al. Decreased apoptosis in the brain and premature lethality in CPP32-deficient mice. Nature 1996; 384: 368–372.
Hakem R, Hakem A, Duncan GS, Henderson JT, Woo M, Soengas MS et al. Differential requirement for caspase 9 in apoptotic pathways in vivo. Cell 1998; 94: 339–352.
Roth KA, Kuan C, Haydar TF, D’Sa-Eipper C, Shindler KS, Zheng TS et al. Epistatic and independent functions of caspase-3 and Bcl-X(L) in developmental programmed cell death. Proc Natl Acad Sci USA 2000; 97: 466–471.
Zaidi AU, McDonough JS, Klocke BJ, Latham CB, Korsmeyer SJ, Flavell RA et al. Chloroquine-induced neuronal cell death is p53 and Bcl-2 family-dependent but caspase-independent. J Neuropathol Exp Neurol 2001; 60: 937–945.
Cecconi F, Alvarez-Bolado G, Meyer BI, Roth KA, Gruss P . Apaf1 (CED-4 homolog) regulates programmed cell death in mammalian development. Cell 1998; 94: 727–737.
Yoshida H, Kong YY, Yoshida R, Elia AJ, Hakem A, Hakem R et al. Apaf1 is required for mitochondrial pathways of apoptosis and brain development. Cell 1998; 94: 739–750.
Yoshida H, Okada Y, Kinoshita N, Hara H, Sasaki M, Sawa H et al. Differential requirement for Apaf1 and Bcl-X(L) in the regulation of programmed cell death during development. Cell Death Differ 2002; 9: 1273–1276.
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006; 441: 885–889.
Yue Z, Jin S, Yang C, Levine AJ, Heintz N . Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA 2003; 100: 15077–15082.
Qu X, Zou Z, Sun Q, Luby-Phelps K, Cheng P, Hogan RN et al. Autophagy gene-dependent clearance of apoptotic cells during embryonic development. Cell 2007; 128: 931–946.
Fimia GM, Stoykova A, Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R et al. Ambra1 regulates autophagy and development of the nervous system. Nature 2007; 447: 1121–1125.
Eskelinen EL, Schmidt CK, Neu S, Willenborg M, Fuertes G, Salvador N et al. Disturbed cholesterol traffic but normal proteolytic function in LAMP-1/LAMP-2 double-deficient fibroblasts. Mol Biol Cell 2004; 15: 3132–3145.
Zeng M, Zhou JN . Roles of autophagy and mTOR signaling in neuronal differentiation of mouse neuroblastoma cells. Cell Signal 2008; 20: 659–665.
Suzuki K, Ohsumi Y . Molecular machinery of autophagosome formation in yeast, Saccharomyces cerevisiae. FEBS Lett 2007; 581: 2156–2161.
Tomoda T, Bhatt RS, Kuroyanagi H, Shirasawa T, Hatten ME . A mouse serine/threonine kinase homologous to C. elegans UNC51 functions in parallel fiber formation of cerebellar granule neurons. Neuron 1999; 24: 833–846.
Clarke PG . Developmental cell death: morphological diversity and multiple mechanisms. Anat Embryol (Berl) 1990; 181: 195–213.
Kegel KB, Kim M, Sapp E, McIntyre C, Castano JG, Aronin N et al. Huntingtin expression stimulates endosomal-lysosomal activity, endosome tubulation, and autophagy. J Neurosci 2000; 20: 7268–7278.
Nixon RA, Wegiel J, Kumar A, Yu WH, Peterhoff C, Cataldo A et al. Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol 2005; 64: 113–122.
Levine B, Yuan J . Autophagy in cell death: an innocent convict? J Clin Invest 2005; 115: 2679–2688.
Cecconi F, Di Bartolomeo S, Nardacci R, Fuoco C, Corazzari M, Giunta L et al. A novel role for autophagy in neurodevelopment. Autophagy 2007; 3: 506–508.
Ferraro E, Cecconi F . Autophagic and apoptotic response to stress signals in mammalian cells. Arch Biochem Biophys 2007; 462: 210–219.
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G . Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat Rev Mol Cell Biol 2007; 8: 741–752.
Botto LD, Moore CA, Khoury MJ, Erickson JD . Neural-tube defects. N Engl J Med 1999; 341: 1509–1519.
Simeone A, Puelles E, Acampora D . The Otx family. Curr Opin Genet Dev 2002; 12: 409–415.
Stanford WL, Cohn JB, Cordes SP . Gene-trap mutagenesis: past, present and beyond. Nat Rev Genet 2001; 2: 756–768.
Nijhawan D, Honarpour N, Wang X . Apoptosis in neural development and disease. Annu Rev Neurosci 2000; 23: 73–87.
Riedl SJ, Salvesen GS . The apoptosome: signalling platform of cell death. Nat Rev Mol Cell Biol 2007; 8: 405–413.
Lakhani SA, Masud A, Kuida K, Porter Jr GA, Booth CJ, Mehal WZ et al. Caspases 3 and 7: key mediators of mitochondrial events of apoptosis. Science 2006; 311: 847–851.
Zheng TS, Hunot S, Kuida K, Momoi T, Srinivasan A, Nicholson DW et al. Deficiency in caspase-9 or caspase-3 induces compensatory caspase activation. Nat Med 2000; 6: 1241–1247.
Colell A, Ricci JE, Tait S, Milasta S, Maurer U, Bouchier-Hayes L et al. GAPDH and autophagy preserve survival after apoptotic cytochrome c release in the absence of caspase activation. Cell 2007; 129: 983–997.
Mizushima N . Autophagy: process and function. Genes Dev 2007; 21: 2861–2873.
Xie Z, Klionsky DJ . Autophagosome formation: core machinery and adaptations. Nat Cell Biol 2007; 9: 1102–1109.
Kirkegaard K, Taylor MP, Jackson WT . Cellular autophagy: surrender, avoidance and subversion by microorganisms. Nat Rev Microbiol 2004; 2: 301–314.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402: 672–676.
Liang C, Feng P, Ku B, Dotan I, Canaani D, Oh BH et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat Cell Biol 2006; 8: 688–699.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005; 122: 927–939.
Takahashi Y, Coppola D, Matsushita N, Cualing HD, Sun M, Sato Y et al. Bif-1 interacts with Beclin 1 through UVRAG and regulates autophagy and tumorigenesis. Nat Cell Biol 2007; 9: 1142–1151.
Jin S, White E . Role of autophagy in cancer: management of metabolic stress. Autophagy 2007; 3: 28–31.
Lum JJ, DeBerardinis RJ, Thompson CB . Autophagy in metazoans: cell survival in the land of plenty. Nat Rev Mol Cell Biol 2005; 6: 439–448.
Rubinsztein DC, Gestwicki JE, Murphy LO, Klionsky DJ . Potential therapeutic applications of autophagy. Nat Rev Drug Discov 2007; 6: 304–312.
Ohsumi Y . Molecular dissection of autophagy: two ubiquitin-like systems. Nat Rev Mol Cell Biol 2001; 2: 211–216.
Eskelinen EL, Tanaka Y, Saftig P . At the acidic edge: emerging functions for lysosomal membrane proteins. Trends Cell Biol 2003; 13: 137–145.
Levine B, Kroemer G . Autophagy in the pathogenesis of disease. Cell 2008; 132: 27–42.
Yuan J, Lipinski M, Degterev A . Diversity in the mechanisms of neuronal cell death. Neuron 2003; 40: 401–413.
Rubinsztein DC . The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 2006; 443: 780–786.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest 2003; 112: 1809–1820.
Acknowledgements
This work was partially supported by grants from Fondazione Telethon, Compagnia di San Paolo, AIRC, The EU 7th Framework Programme, the Italian Ministry of University and Research (MUR) and the Italian Ministry of Health; FC is an Associate Telethon Scientist. We thank Martin W Bennet for the valuable editorial work and Marcello Giorgi for the drawings in Figure 1.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by G Melino
Rights and permissions
About this article
Cite this article
Cecconi, F., Piacentini, M. & Fimia, G. The involvement of cell death and survival in neural tube defects: a distinct role for apoptosis and autophagy?. Cell Death Differ 15, 1170–1177 (2008). https://doi.org/10.1038/cdd.2008.64
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2008.64
Keywords
This article is cited by
-
Characterization of the effects of heat stress on autophagy induction in the pig oocyte
Reproductive Biology and Endocrinology (2021)
-
Magnitude and determinants of neural tube defect in Africa: a systematic review and meta-analysis
BMC Pregnancy and Childbirth (2021)
-
Nuclear factor I-C disrupts cellular homeostasis between autophagy and apoptosis via miR-200b-Ambra1 in neural tube defects
Cell Death & Disease (2021)
-
Association between rare variants in specific functional pathways and human neural tube defects multiple subphenotypes
Neural Development (2020)
-
Rare mutations in apoptosis related genes APAF1, CASP9, and CASP3 contribute to human neural tube defects
Cell Death & Disease (2018)