Abstract
Myotonic dystrophy (DM) is caused by a (CTG)n expansion in the 3′-untranslated region of DMPK gene. Mutant transcripts are retained in nuclear RNA foci, which sequester RNA binding proteins thereby misregulating the alternative splicing. Controversy still surrounds the pathogenesis of the DM1 muscle distress, characterized by myotonia, weakness and wasting with distal muscle atrophy. Eight primary human cell lines from adult-onset (DM1) and congenital (cDM1) patients, (CTG)n range 90–1800, were successfully differentiated into aneural-immature and contracting-innervated-mature myotubes. Morphological, immunohistochemical, RT-PCR and western blotting analyses of several markers of myogenesis indicated that in vitro differentiation–maturation of DM1 myotubes was comparable to age-matched controls. In all pathological muscle cells, (CTG)n expansions were confirmed by long PCR and RNA fluorescence in situ hybridization. Moreover, the DM1 myotubes showed the splicing alteration of insulin receptor and muscleblind-like 1 (MBNL1) genes associated with the DM1 phenotype. Considerable myotube loss and atrophy of 15-day-differentiated DM1 myotubes indicated activated catabolic pathways, as confirmed by the presence of apoptotic (caspase-3 activation, cytochrome c release, chromatin fragmentation) and autophagic (P62/LC3) markers. Z-VAD treatment significantly reduced the decrease in myonuclei number and in average width in 15-day-differentiated DM1 myotubes. We thus propose that the muscle wasting typical in DM1 is due to impairment of muscle mass maintenance–regeneration, through premature apoptotic–autophagic activation, rather than altered myogenesis.
Similar content being viewed by others
Main
Myotonic dystrophy (DM) is a multi-systemic disorder caused by two different microsatellite expansions in non-coding regions. Together, these two mutations affect 1 out of 8000 individuals and represent the most common form of muscular dystrophy in adults. DM1 and DM2 have common symptoms such as myotonia, muscle weakness and early cataract development.1, 2 Although DM1 and DM2 initially affect different muscles (distal versus proximal), histological analysis of the muscular tissues shows common aspects such as central nucleation. The classic form of DM1 is characterized by muscle distress with myotonia, progressive muscle weakness and wasting. Atrophy has also been reported, occurring preferentially in type-1 fibers in DM1 and in type-2 in DM2.3 DM1 but not DM2 also presents a congenital form (cDM1), characterized by a high neonatal mortality and symptoms such as hypotonia, mental retardation and respiratory distress.4, 5
DM1 is associated with an unstable (CTG)n trinucleotide expansion located in the 3′-untranslated (3′-UTR) region of the DM protein kinase (DMPK) gene on chromosome 19q13.3. The mutant DMPK transcript, containing the expanded (CTG)n sequence, accumulates in discrete nuclear foci able to sequester various nuclear factors such as RNA-binding proteins or splicing regulators, causing different and highly variable downstream deleterious effects.2, 6
It has been reported that (CTG)n trinucleotide repeat length in muscle directly correlates with both frequency of severe cDM17 and rate of splicing impairment typical of DM.8
The pathology of skeletal muscle, including both myotonic and dystrophic features, has been reproduced in different mouse models by the following different strategies: random insertion of a genomic fragment carrying the expanded sequence in its human DM1 context9 or in human skeletal actin (HSA) gene,10 or insertion of a humanized DMPK locus containing a human expanded sequence.11 An inducible model for the pathology carrying 960 (CTG)n and showing severe muscle wasting was also recently created.12
Cell models are also available, such as C2C12 lines expressing the (CTG)n expansion13, 14, 15, 16, 17 or human DM1 fibroblasts converted into myoblasts by the induction of the myogenic factor MyoD.18, 19 Use of human DM1 primary myoblasts cultures has been rare, owing to limited availability, and has resulted in contradictory findings.20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31
Various experimental models have been used to speculate about the possible effects of the DM1 mutation on the myogenic process and thus find explanation for the severe skeletal muscle immaturity and wasting of DM muscle.16 The most commonly accepted theory is that (CTG)n expansion could affect differentiation of DM myoblasts by interfering with the signals leading to the withdrawal of cell cycle and the shift toward the differentiation program.26, 32, 33 However, it should be noted that the majority of the data on in vitro differentiation of DM muscle cells derive from studies on DM1 fibroblasts converted into skeletal muscle,18, 19 from primary muscle cell cultures obtained from cDM1 fetuses31 or from mouse muscle cells.14, 15, 17, 33, 34 No studies focusing on muscle differentiation of human primary DM1 myoblasts cultures obtained from adult muscle DM1 biopsies are available, underlining the need to work with a model as close as possible to human regenerating skeletal muscle. To address this point we chose as experimental model primary cultures from skeletal muscle biopsies of five healthy, six DM1 and two cDM1 patients with different degrees of pathology. Cultured cells at various times of differentiation were tested for their myogenic potential and for the main molecular markers associated with the DM1 pathology – number of foci and splicing alteration. Moreover, we investigated different catabolic pathways including the expression of the critical ubiquitin-ligases atrogin-1 and MuRF1, apoptosis and autophagy in well-differentiated myotubes. We provide evidence that apoptosis and autophagy are possible mechanisms leading to degenerative loss of muscle tissue and impairment in regeneration.
Results
Differentiation of DM1 myoblasts is normal
Morphological, molecular and immunological analysis were performed during the first 15 days of differentiation in primary skeletal muscle cell culture lines derived from eight DM1 patients (six adult-onset DM1, two cDM1; Table 1), with blood (CTG)n repeats within the DMPK gene/mRNA ranging from 90–1800 and from five age-matched controls. Muscle terminal differentiation was induced by a shift to a medium containing a low concentration of mitogens. Normal and DM1 cells grew at the same rate and went through the same number 2–7 of passages. As our culture procedure established a mixture of fibroblasts and myoblasts, in all the cell lines the percentage of desmin-positive myoblast population was calculated and found to exceed 80%.
To compare the myogenic capacity of myoblasts from normal and DM1 patients we examined morphological aspects together with the expression of different markers of myogenesis at 4 and 10 days of differentiation (stages T4 and T10). Slow myosin heavy chain (MHC) and fetal myosin were detected by immunofluorescence (Figure 1b; Supplementary Figure 1); the levels of myogenin were estimated by a combination of RT-PCR and western assays (Figure 1c, d1 and d2); the level of muscle-specific creatine kinase (CK-M) was quantified by RT-PCR (Figure 1e). We found that in the T4- and T10-differentiated DM1 myotubes, all the tested parameters were similar to controls. A representative set of images in Figure 1a and b shows that the differentiation capacity was unaffected by the presence of the (CTG)n expansion. All the DM1 cell lines were then subjected to fusion assays. Again, the myogenic potential, expressed as number of syncytial nuclei per myotube, was similar to controls (Figure 1f). Moreover, to check the maturation capacity of DM1 muscle cells, primary DM1 myotubes were innervated with sections of rat embryo spinal cord (Supplementary Figure 1 insert B). This kind of procedure allows further muscle maturation to be achieved, thanks to the diffusion of axons from the gangliar roots.28, 35, 36 All the innervated DM1 myotubes acquired the ability to depolarize and contract as controls (Supplementary movie). Taken together, these data suggest that our DM1 human primary myoblasts did not present any impairment in the early steps of myogenesis, behaving as controls until 10 days of differentiation.
Normal myogenic potential of DM1 myoblasts. (a) Representative pictures of 10-day-differentiated myotubes from control and DM1 patients. DM1 lines generated multi-nucleated myotubes similar to controls. Expansion (CTG)n range in myotubes: DM1-E2: 130–280; DM1-E3: >1500, cDM1-E4: congenital form, 2500–4000. Scale bars 30 μm. (b) No differences of slow MHC expression were detected in fixed 10-day-differentiated myotubes from control and DM1 patients. Nuclear DMPK mRNA foci (arrows) were labeled by (CAG)10 probe fluorescence in situ hybridization. Sarcomeric organization is well defined in DM1 myotubes. Scale bar 25 μm. (c, d1, d2) In 4- to 10-day-differentiated myotubes (T4–T10) from five DM1 and three controls, the RNA (c) and the protein presence (d1, d2) of myogenin were similar. DM1-E21, DM1-E32, and cDM1-E41 patients were tested in myogenin blot. The values of myogenin mRNA and protein amount are given as arbitrary units (A.U.) of ratio with β2-microglobulin housekeeping gene or with β-actin, respectively. The data are expressed as mean±S.E. of three different experiments of RT-PCR (carried out in triplicate) and a single WB analysis. (e) Analogous expression levels of muscle-specific creatine kinase (CK-M) in T4–T10 myotubes from five DM1 and three control subjects. The values of CK-M mRNA are given in arbitrary units (A.U.) of ratio with β2-microglobulin housekeeping gene. The data are expressed as mean±S.E. of three different experiments of RT-PCR done in triplicate. (f) Similar average number of syncytial nuclei in T4–T10 of three DM1 and three control myotubes, expressed as mean±S.E. The analysis was performed on at least 100 myotubes for each line
Pathological hallmark in DM1 muscle cells
To exclude the possible selection of healthy differentiating cells, our cultured DM1 myotubes were characterized for the presence of the main molecular hallmarks of the DM1 disease: the presence of ribonuclear foci and the splicing misregulation of two genes representative of the DM1 spliceopathy, the insulin receptor (IR) and muscleblind-like 1 (MBNL1).
In all DM1 myotubes expanded DMPK transcripts, accumulating in the nucleus together with nuclear proteins with affinity for the expanded CTG sequence, generated aggregates easily detectable by fluorescence in situ hybridization (FISH),37, 38 confirming the presence of DMPK pathological transcripts in all the studied muscle cells (Figure 2a).
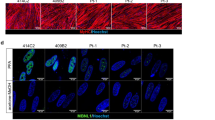
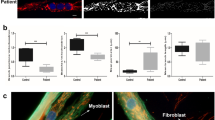
Pathological myotubes retain the hallmarks of myotonic dystrophy. (a) Representative images of nuclear foci detection in DM1 fixed myotubes, obtained by FISH with a TexasRed-labeled (CAG)10 probe. All DM1 cells showed the presence of nuclear foci. Scale bar 25 μm. Panels showing the RT-PCR splicing assay of IR (b1, b2) and MBNL1 (c1, c2) in T0–T4–T10 DM1-E21, -E32, and -E41 and three normal myotubes and in the corresponding parental muscle biopsies. Exons analyzed for each gene are shown. The diagrams represent the ratio of aberrant isoform to total transcript in DM1 and control samples given as median values
Aberrant isoforms of various genes, including IR-A and MBNL1 (MBNL1ex7), prevail in affected adult DM1 muscle in relation to the (CTG)n expansion.8 To assess the in trans effect of the CTG repetition on the splicing regulation in DM1 differentiating myoblasts, we analyzed the expression of IR and MBNL1 gene isoforms at different times of differentiation (stages T0, T4 and T10). RT-PCR amplification showed that in DM1 cells the ratios of the IR-A/TOT (Figure 2b1 and b2) and MBNLex7/TOT (Figure 2c1 and c2) increased in correlation with the (CTG)n length and were significantly higher compared to controls especially in the E3 (IR-A P<0.05; MBNL1 P<0.01) and E4 samples (IR-A P<0.01; MBNL1 P<0.001) (Figure 2b1, b2 and c1, c2).
Catabolic pathways in DM1 muscle cells
These results were accompanied by the observation that DM1 myotube population drastically decreased after 12–15 days of differentiation compared with controls (Figure 3a), and that average width of DM1 myotubes with (CTG)n expansion was statistically reduced by 30% compared with controls (P<0.01) (Figure 3c). Several studies showed that muscle atrophy is accompanied by a reduction in mean number of myonuclei per fiber. Consistently, we found a statistically reduced number of nuclei/myotube in DM1 lines, by 16% compared with controls (Figure 3b).39 Together, these results suggested the activation of catabolic pathways in our muscle cells of DM1 patients.
Reduction of DM1 myotube population after 15 days of differentiation. (a) Representative bright field images of control and cDM1-E41 myotube population after 15 days of differentiation. Scale bar 100 μm. (b) Average number of nuclei per myotube at 10–15 days of differentiation. The nuclei/myotubes values were obtained considering at least 100 myotubes/cell line and are expressed as mean±S.E. In 15-day-old DM1 myotubes, the syncytial nuclei number were significantly reduced compared with controls (***P<0.001). (c) Average myotube width (AMW), measured in at least 100 myotubes/cell line with commercial software, is expressed as mean±S.E. Pathological lines after 15 days of differentiation showed a significant decreased AMW (**P<0.01). Analisys was carried out on DM1-E21, DM1-E32 and cDM1-E41 patients and three controls
To characterize the signaling involved in DM1 myotubes loss, we investigated the presence of oxidative stress, the expression of atrophy-related genes and the activation of apoptotic and autophagic systems. Expression of Atrogin1 and Murf1, the two critical atrophy-related muscle-specific ubiquitin-ligases, did not differ from controls (data not shown). This finding suggests a minor contribution, at this time point, of ubiquitin-proteasome system to myotube loss. ROS production, tested in myotubes from 4 to 15 days of differentiation, did not show noticeable differences compared with controls at 4–10 days of differentiation (data not shown). However, 15-day-differentiated DM1-E2 and cDM1-E4 myotubes significantly produced more H2O2 than controls (Supplementary Figure 2). This finding suggests that dysfunctional mitochondria or conditions that generate ROS persist in DM1-E2 and E4 myotubes and can therefore contribute to myotube wasting.
We next monitored the presence of apoptosis by studying chromatin fragmentation by terminal deoxynucleotidyltransferase-mediated dUTP nick end labeling (TUNEL), release of cytochrome c from the mitochondria and caspase-3 cleavage (Figure 4). All these approaches confirmed a higher amount of apoptosis in DM1 myotubes when compared with controls. Especially, TUNEL assay revealed a constant elevated presence of apoptotic nuclei in all DM1 myotubes during the differentiation, in particular, T10-T15 DM1-E21, -E22 and cDM1-E41, -E42 samples had ∼10–15 times more apoptotic myonuclei than control samples (Figure 4a1 and a2). Consistently, cytochrome c release (Figure 4b1 and b2) and activation of caspase-3 (Figure 4c1 and c2) were clearly increased in myotubes from the same lines of DM1-E2 and E4, whereas DM1-E3 lines showed reduced apoptotic values, which were, however, higher than controls. Interestingly pAKT, a pro-survival factor, is increased by 50% only in DM1-E3 cells but not in DM1-E2 and cDM1-E4 lines (Figure 4d1 and d2).40
Apoptotic features in 15-day-differentiated DM1 myotubes. (a1) Representative images of TUNEL reaction in normal and cDM1-E42 myotubes. Scale bar 20 μm. (a2) Undifferentiated (T0), T4, T10 and T15 DM1-E21, -E22, -E32, -E33 and cDM1-E41, -E42 and three control myotubes were scored for apoptotic nuclei positive to TUNEL staining. The data are expressed as percentage of TUNEL-positive nuclei-apoptotic myotubes to total myotubes, counted in ca 150 myotubes/cell line. Frequency of apoptosis in pathological myotubes was found to be significantly increased compared with control for P<0.001 by non-parametric two-way ANOVA analysis. (b1) Representative images of cytochrome c release in cDM1-E42. (b2) Localization index of cytochrome c in fixed myotubes from the same control and DM1 patient lines. During apoptosis cytochrome c leaks from the mitochondria to cytosol. The colocalization of cytochrome c stain (green) and mitochondrial marker TOM 20 (red) was near one in normal myotubes and significantly decreased in DM1-E2, -E4 myotubes (P<0.05) but not in DM1-E3 lines. Values were obtained in 5–10 myotubes/cell line. (Scale bars 10 μm.) (c1) Western blot analysis for activated caspase-3 proved positive only in pathological DM1-E21, -E32 and -E41 myotubes, not in normal myotubes. (c2) The diagram represents the values of activated caspase-3 compared to a β-actin, obtained by integrated optical density quantification. (d1) Western blot analysis for pAKT and AKT in 15-day-differentiated DM1-E21, -E32 and -E41 and three control myotubes. (d2) pAKT amount given as ratio to total AKT
To establish the relevance of apoptotic process, 10-day DM1 myotubes were treated for 5 days with 50 μM Z-VAD, a pancaspase inhibitor. The treatment produced a visual decrease in atrophic DM1 myotubes (Figure 5a), a significant 23% increase in number of myonuclei (Figure 5b) and a 26% rise of average myotube width (AMW) (Figure 5c) in DM1-E2 and DM1-E4 myotubes compared to the untreated. Overall, these data provided evidence that apoptosis actively contributed to the observed catabolic pathways.
Z-VAD treatment in 10- to 15-day-differentiated DM1 myotubes. The pancaspase inhibitor Z-VAD caused a visual impression of increased trophicity in the DM1-treated myotubes. (a) Representative images of DM1-E21 and E42 myotubes with and without Z-VAD treatment. Scale bar 25 μm. A consistent, statistically significant increase was found in (b) myonuclei (*P<0.02); (c) AMW (**P<0.01) in DM1-treated myotubes compared with the untreated. The number of single lines studied is given in brackets
The alter ego of apoptosis is the autophagic system, which is the vehicle for delivering proteins and organelles to the lysosome.41 The modern view considers autophagy as a pro-survival system that keeps the cell clear of toxic proteins and damaged organelles. However, excessive autophagy contributes to protein breakdown and muscle atrophy in skeletal muscle42, 43 and to cell death in mononucleated cells. In view of this relationship with apoptosis and muscle loss, we studied the level of autophagy activation in control and DM1 myotubes. LC3-positive vesicles, a marker of activated autophagy,41 were significantly increased in DM1-E2, -E3 and -E4 myotubes at T15 compared with controls; the quantification is represented in the graph as number of LC3-positive vesicles/μm2 (Figure 6a1 and a2). Instead, P62 showed no substantial changes among the various groups (Figure 6b1 and b2). It is interesting to note that the DM1-E3 lines (which showed the lowest levels of apoptosis and no ROS production) present the highest levels of autophagy, giving rise to a similar apoptosis effect, that is degeneration of 15-day myotubes.
Autophagy and proliferative response in differentiated DM1 myotubes. The clusterization of LC3 was measured as autophagic marker in T15 myotubes from three controls and six DM1 by immunofluorescence. (a1) Representative images of LC3-positive vesicles in control and DM1-E32 myotubes. Scale bar 25 μm. (a2) The diagram represents mean±S.E. of the data obtained by scoring 10 images/cell line as described in ‘Materials and Methods’ and indicates a significant increase in clustered LC3 in DM1-E21, -E22, -E32, -E33 and cDM1-E41, -E42 myotubes compared with controls (**P<0.01). (b1, b2) The protein level of P62 was revealed by western blot analysis and quantified by densitometry in three controls and DM1-E22, -E33 and cDM1-E42 The bar graph represents the P62/β-actin ratio
Discussion
Myogenic potency in DM1
This study clearly shows that primary muscular satellite cells from DM1 patients with either congenital or adult form have normal fusion, differentiation and maturation capacity. When we studied the myogenic capacity we found that the number of syncytial nuclei, the expression level of myogenic markers (CK-M, myogenin and MHC) and the contractile capability after innervation were similar to controls and unaffected by the number of (CTG)n repeats. We excluded an in vitro selection toward low (CTG)n repeats, as (CTG)n expansion was confirmed at RNA level by the presence of ribonuclear foci, the pathological hallmark of DM1, which was always found in undifferentiated (data not shown), differentiated and innervated DM1 muscle cells. In vivo, the retention into ribonuclear foci of the expanded DMPK transcript is associated with the misregulation of alternative splicing. This in trans effect on the alternative splicing of many RNAs, which does not result in the production of mutant protein but leads to expression of spliced products inappropriate for a particular tissue, is the major molecular defect identified in DM1.1, 6 We provide evidence that splicing unbalance is present in undifferentiated and differentiated DM1 muscle cells. Moreover, similarly to what is observed in vivo,8 also in vitro there is a correlation between the extend of the (CTG)n repeat size and the degree of abnormal splicing. Differentiation capacity of primary human DM1 muscle cells is controversial. Normally-differentiated human myotubes were reported in calcium homeostasis studies from a large cohort of 15 adult DM1 patients20, 21, 22 and recently in 5 DM2 patients.23 Moreover, in vitro successfully innervated muscle fibers from 17 adult DM1 patients were used for studies of electrophysiological properties of DM1 mature myotubes28 and developmental regulation of DMPK.29, 30 In contrast, other reports claim an impaired myogenesis26, 31 or a delayed maturation of primary myotubes from three cDM1 and few adult DM1-DM2 patients.24, 27 In addition, a large number of publications describe myogenic alterations in mouse muscle models13, 14, 15, 16, 17 or in primary DM1 fibroblasts converted in myoblasts by MyoD transfection.18, 19
The delayed muscle development has been proposed as being determined by dysfunction of the signaling pathway that includes MyoD, Myf5, MRF4 factors responsible for the commitment of muscle stem cells to the myogenic lineage, and myogenin and MFR4, factors involved in the expression of the terminal muscle phenotype.
It was found that the levels of the commitment factor MyoD were reduced both in C2C12 cells expressing mutant DMPK 3′-UTR RNA and in DM1 patient myoblasts.15, 26 However, this reduction of MyoD protein levels was not accompanied by a decrease in its homologue Myf5.
Moreover, Timchenko et al.26 proposed that myoblasts from one DM1 and one DM2 patients failed to undergo differentiation because of an impairment of the p21/CDK4/Rb/E2F pathway usually essential for the withdrawal from cell cycle. A recent publication proposed32 that the ectopic expression of cyclin D3 corrects differentiation in DM1 myoblasts by increasing the CUGBP1–eIF2 interaction necessary for activation of the myogenic program and the correct expression of myogenic markers such as myogenin.
The discordance between our results showing a normal DM1 primary myoblast differentiation and the reported alteration in the myogenic process of DM1 myoblasts might be accounted for by the different models used (i.e. myoblasts from human adult versus fibroblasts converted into myoblasts and mouse model) or by a difference in the morphological and clinical severity of the disease (i.e. myoblasts from human cDM1 patients that survived after the neonatal period versus myoblasts from aborted cDM1 fetuses) or by a difference in the age of the satellite cells, which may also influence myogenesis44 because of the recently reported implication of the p16 premature senescence of DM1 myoblasts.45, 46
Catabolic pathways and a novel pathogenetic mechanism
During differentiation, myoblasts undergo sequential events, ending with fusion into syncytial cells, a cell type more resistant to sublethal damage than proliferating myoblasts, which go on to generate muscle fibers. However, in many degenerative and metabolic diseases, muscle fibers die with no marked inflammatory response, this could probably be due to apoptosis.47 Therefore, the study of myotubes in vitro is a good model for predicting if apoptosis is active at least in developing myofibers. Apoptosis in primary myotubes was evident in merosin, dystrophin and dystrophin-associated protein-deficient cell lines.48, 49 Recently, Ullrich congenital dystrophy, caused by collagen-VI mutations, presented an increased occurrence of spontaneous apoptosis.50, 51
At 15 days of culture, we observed a statistically significant decrease in the number of myonuclei in DM1 patients and concomitant evidences of atrophy. It has been reported that ubiquitin-proteasome, autophagy-lysosome and caspase-3/9 activation contribute, albeit to a different extent, to muscle loss. Indeed, we found an increase in apoptosis and autophagy even though no induction of atrophy-related genes, atrogin-1 and MuRF1, was observed in any of the 15-day-differentiated DM1 myotubes; however, it could not be excluded that these ubiquitin-ligases might contribute to myotube atrophy between 10 and 15 days. It is interesting to note that the reduced incidence of apoptosis in DM1-E3 cells matches with a concomitant significant increase of autophagy and of AKT activation. This trend can be explained in terms of the well-known inverse relationship found between the pro-cell death system, apoptosis and the pro-survival system autophagy.41 In our DM1 myotubes, apoptosis occurred after differentiation and not in undifferentiated myoblasts. The role of apoptosis was confirmed by Z-VAD treatment, which significantly recovered the number of myonuclei and AMW, so identifying apoptosis as a potential candidate for therapeutic approaches (Figure 5).
A novel pathogenetic mechanism is emerging from our findings. It is important to underline that activation of autophagy matches with the (CTG)n expansion, autophagy being highly activated in samples with high numbers of (CTG)n (Figure 6a). This correlation is consistent with the hypothesis that autophagy is induced to clear toxic proteins and organelles to maintain cell viability. Thus, autophagy may act as a pro-survival system, which can also reduce muscle mass as a result of to its proteolytic activity. Increasing the number of (CTG)n triplets can affect the expression of critical factors for protein-folding processes, response to protein unfolding/misfolding and many other cellular components including proteins involved in Ca2+ homeostasis. Thus autophagy failure or exhaustion can lead to accumulation of toxic proteins, which can interfere with organelle (mitochondria) function and with cellular signaling leading to myofiber degeneration. It should be noted that we found an increase in H2O2 production in 15-day-differentiated DM1-E2 and cDM1-E4 myotubes, which showed higher level of apoptosis and lower level of autophagy. As one of the main sources of ROS is dysfunctional mitochondria, the presence of H2O2 suggests a failure in the removal system for altered mitochondria. The persistence of abnormal mitochondria would induce the release of pro-apoptotic factors. Therefore, a failure or overload of autophagy induces accumulation of death-signaling components, which triggers apoptosis and myofiber degeneration. This hypothesis is in line with the established fact that autophagy and apoptosis are mutually exclusive and not synergistic.
In conclusion, this is the first study focused on the differentiation of myoblasts from patients with adult classical DM1 and cDM1 that survived at the perinatal crisis, and allows us to formulate a new hypothesis to explain the progressive DM1 muscular pathogenesis. We propose that apoptosis–autophagy is the key event, probably coupled to oxidative stress and misregulation of calcium homeostasis, which may also be linked to premature senescence of satellite cells.
Materials and Methods
Clinical data
After informed consent, biopsies of vastus lateralis were obtained from five healthy donors (two females and three males, age range 2–55 years), and from eight unrelated DM1 patients (three E2<450 (CTG), three E3>1000 (CTG) and two congenital E4). DM1 patients were diagnosed at the Department of Neurology, University of Padua, Italy (Table 1). The diagnosis of DM was based on clinical, electromyographic (high-frequency repetitive discharges), ophthalmologic and cardiac investigations. The degree of muscle impairment was assessed using muscular disability grading (Muscular Disability Rating Scale, MDRS), based on a five-point scale as described earlier:52 grade 1, no clinical impairment; grade 2, minimal signs of clinical impairment; grade 3, distal weakness; grade 4, mild or moderate proximal weakness; grade 5, severe proximal weakness (confined to wheelchair for short or long distances). In addition to MDRS rating of muscle impairment, we also assessed cognitive impairment, cataract, cardiac involvement, endocrine dysfunctions and motor impairment (Table 1).
Cell cultures
Primary myoblasts were obtained as described earlier.35 Cells were cultured with Ham's F14 medium (Euroclone, Milan, Italy) plus 20% FBS (Gibco, Invitrogen, Milan, Italy) and 10 μg/ml insulin (Sigma, Milan, Italy). When a 70% confluence was reached, differentiation was triggered by lowering FBS to 2%. Samples were collected at 0, 4, 10 and 15 days of differentiation using techniques specific to the various different analyses. Z-VAD-FMK (50 μM) (ALEXIS Biochemicals, Modena, Italy) treatment was performed for 5 days (from 10 to 15 days of differentiation), with daily change of medium.
Functional innervation was obtained by co-culturing differentiating myoblasts with 13-day-old Sprangue-Dawley rat embryo spinal cord maintaining dorsal root ganglia, as described earlier.35
Morphological analysis and immunofluorescence
Bright-field images of myotubes were collected using a Zeiss IM35 microscope equipped with a standard camera. Differentiation was quantified considering the average number of nuclei per myotube in × 100 images of at least 100 myotubes for each cell line. Myotubes average width at 10 and 15 days of differentiation was measured using ImageJ software. For each parameter, at least 100 myotubes per class were considered. Similar parameters were measured in a separate set of experiments with and without Z-VAD treatment: 100 and 50 myotubes from untreated–treated controls and DM1, respectively, were considered.
Immunofluorescence was performed in fixed myotubes, permeabilized with 0.2% Triton X-100 and incubated for 30 min with 0.5% BSA and 10% horse serum in PBS. Primary specific antibodies were diluted in PBS plus 2% BSA and incubated for 1 h at room temperature (slow MHC, gift from Professor Schiaffino) or overnight at 4°C (LC3, Cell Signaling, Danvers, CO, USA). Secondary Alexa 488 or Cy3-conjugated antibodies (Invitrogen-Molecular Probes, Milan, Italy) were incubated for 1 h. Samples, mounted in Vectashield mounting medium with DAPI (4′-6-diamidino-2-phenylindole) (Vector Laboratories, Burlingame, CA, USA), were observed with an Olympus BX60 fluorescence microscope ( × 20 magnification). Quantitative analysis of at least 30 LC3 stained myotubes per sample was performed using commercial software by creating specific regions of interest corresponding to single myotubes, and counting the amount of LC3-positive vesicles. Data were expressed as number of LC3-positive vesicles/μm2.
RNA fluorescence in situ hybridization
Myoblasts and 4-, 10-, 15-day myotubes grown on coverslips were fixed in 4% paraformaldehyde, 10% acetic acid in PBS for 15 min at 4°C and permeabilized in 0.2% Triton X-100 in PBS for 5 min at room temperature. RNA fluorescence in situ hybridization was performed as described.8
TUNEL and cytochrome c release detection
Apoptosis detection in 0-, 4-, 10- and 15-day myotubes was performed using TUNEL and cytochrome c release assays. In the cells, fixed and processed for TUNEL analysis (TUNEL System, Promega, Milan, Italy), visualization of all nuclei was performed with Vectashield mounting medium with 0,1 μg/ml DAPI. At least 50 myotubes per cell line were considered. Cytochrome c/TOM20 colocalization was performed on 15-day-differentiated myotubes using a balanced mix of anti-cytochrome c (BD Pharmigen, San Diego, CA, USA) and anti-TOM20 (Santa Cruz, DBA, Milan, Italy) antibodies. At least 10 myotubes per cell line were analyzed using a Leica TCSP5 confocal microscope and the correlation parameter R was calculated using ImageJ software.
RNA analysis
Total RNA was isolated from myoblasts and 4- and 10-days myotubes using Trizol reagent (Sigma). In all, 1 μg of total RNA was reverse-transcribed to cDNA using the SuperScript III First-Strand Synthesis System for RT-PCR (Invitrogen).
The expression levels of CK-M (muscle-specific creatine kinase) and MYOG (myogenin) genes, compared with the expression of the housekeeping gene β2-microglobulin, were measured by Sybrgreen RT-PCR with ABI PRISM7000 sequence detection system. The following specific primers were used: CK-M, F, 5′-CAAGGAACTCTTTGACCCCA-3′, R, 5′-CCACAGAGAGCTTCTCCACC-3′, myogenin, F, 5′-AAGAGAAGCACCCTGCTCAA-3′, R, 5′-CAGATGATCCCCTGGGTTG-3′, β2-microglobulin, F 5′-ATGAGTATGCCTGCCGTGTGA-3′, R, 5′-GGCATCTTCAAACCTCCATG-3′.
Atrogin (Hs00369709_m1) and Murf1 (Hs00822397_m1) gene expression was determined by a multiplex TaqMan QRT-PCR reaction using a VIC-labeled human β-actin (Hs99999903_m1)-specific probe as housekeeping. Genes of interest were all FAM labeled. The PCR splicing assays for IR and MBNL1 genes were performed as described earlier.8 Total PCR products, obtained within the linear range of amplification, were electrophoresed on 3.5% agarose gel. Quantitative analysis of the amplified products was performed using ethidium bromide staining. The integrated optical density of each band and the fraction of fetal transcript versus total transcript were quantified by densitometry using commercial software.
ROS production assay
The rate of H2O2 production in living cells was determined using the oxidation in the extracellular medium of 20 mM fluorogenic indicator Amplex red in the presence of 1 unit/ml horseradish peroxidase (POD) and was expressed as A.U./min/mg prot.
In all, 30 000 myoblasts were seeded in triplicate in 12-well culture plates and differentiated. The assay was conducted with 4-, 10- and 15-day myotubes. Fluorescence was recorded on a microplate reader (Ascent Fluoroscan FL2.5, Labsystem, Milan, Italy) (Ex: 530 nm; Em: 585 nm), in presence of 10 mM apocynin (40-hydroxy-30-methoxyacetophenone) to inhibit H2O2 production by plasma membrane NADPH-oxidase.
Western blotting
Myotubes were collected after 10 and 15 days of differentiation. For muscle samples, 10 μm cryosections were used. Total protein extracts were prepared as described earlier53 and electrophoresed in 7.5–17.5% T30C4 SDS-PAGE gels or 4–12% NuPAGE precast gels (Invitrogen). Proteins were blotted into nitrocellulose membrane or 0.45 μm PVDF (Invitrogen) and probed with specific antibodies against myogenin (Hybridoma bank, Iowa City, IA, USA), P-AKT (Ser473) (Cell Signaling), total AKT (Cell Signaling), cleaved caspase 3 (Cell Signaling), P62 (Progen, Heidelberg, Germany), LC3 (NanoTools). After incubation with specific secondary HRP-conjugated antibodies, recognized bands were visualized by chemiluminescence (GE HealthCare, Milan, Italy). Integrated optical density of each band was calculated with commercial software and normalized compared with actin amounts.
Statistical analysis
Statistical analysis was performed only where three or more experimental values were available. Quantitative data were presented as means±S.E. In the case of normal distribution of values, confirmed by Shapiro's test, statistical comparisons were performed using the Student's t-test. With non-Gaussian distributions, non-parametric Kruskal–Wallis and Wilcoxon tests were applied. TUNEL data were analyzed with the non-parametric two-way ANOVA test. In every analysis values of P<0.05 were considered significant.
Abbreviations
- AMW:
-
average myotube width
- ANOVA:
-
analysis of variance
- cDM1:
-
congenital myotonic dystrophy type 1
- CK-M:
-
muscle-specific creatine kinase
- DAPI:
-
4′-6-diamidino-2-phenylindole
- DM:
-
myotonic dystrophy
- DMPK:
-
myotonic dystrophy protein kinase
- FBS:
-
fetal bovine serum
- FISH:
-
fluorescence in situ hybridization
- IR-A/B:
-
insulin receptor-A/B
- MBNL1:
-
muscleblind-like 1
- MDRS:
-
Muscle Disability Rating Scale
- MHC:
-
myosin heavy chain
- PCR:
-
polymerase chain reaction
- POD:
-
peroxidase
- ROI:
-
region of interest
- ROS:
-
reactive oxygen species
- TUNEL:
-
terminal deoxynucleotidyl transferase dUTP nick end labeling
- Z-VAD-FMK:
-
Z-Val-Ala-Asp-fluoromethylketone, pancaspase inhibitor
References
Machuca-Tzili L, Brook D, Hilton-Jones D . Clinical and molecular aspects of the myotonic dystrophies: a review. Muscle Nerve 2005; 32: 1–18.
Cho DH, Tapscott SJ . Myotonic dystrophy: emerging mechanisms for DM1 and DM2. Biochim Biophys Acta 2007; 1772: 195–204.
Vihola A, Bassez G, Meola G, Zhang S, Haapasalo H, Paetau A et al. Histopathological differences of myotonic dystrophy type 1 (DM1) and PROMM/DM2. Neurology 2003; 60: 1854–1857.
Harper PS . Myotonic Dystrophy. WB Saunders: London, 2001.
Brook JD, McCurrach ME, Harley HG, Buckler AJ, Church D, Aburatani H et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 1992; 68: 799–808.
Wheeler TM, Thornton CA . Myotonic dystrophy: RNA-mediated muscle disease. Curr Opin Neurol 2007; 20: 572–576.
Tsilfidis C, MacKenzie AE, Mettler G, Barcelo J, Korneluk RG . Correlation between CTG trinucleotide repeat length and frequency of severe congenital myotonic dystrophy. Nat Genet 1992; 1: 192–195.
Botta A, Rinaldi F, Catalli C, Vergani L, Bonifazi E, Romeo V et al. The CTG repeat expansion size correlates with the splicing defects observed in muscles from myotonic dystrophy type 1 patients. J Med Genet 2008; 45: 639–646.
Seznec H, Agbulut O, Sergeant N, Savouret C, Ghestem A, Tabti N et al. Mice transgenic for the human myotonic dystrophy region with expanded CTG repeats display muscular and brain abnormalities. Hum Mol Genet 2001; 10: 2717–2726.
Mankodi A, Logigian E, Callahan L, McClain C, White R, Henderson D et al. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science 2000; 289: 1769–1773.
van den Broek WJ, Nelen MR, Wansink DG, Coerwinkel MM, te RH, Groenen PJ et al. Somatic expansion behaviour of the (CTG)n repeat in myotonic dystrophy knock-in mice is differentially affected by Msh3 and Msh6 mismatch-repair proteins. Hum Mol Genet 2002; 11: 191–198.
Orengo JP, Chambon P, Metzger D, Mosier DR, Snipes GJ, Cooper TA . Expanded CTG repeats within the DMPK 3′ UTR causes severe skeletal muscle wasting in an inducible mouse model for myotonic dystrophy. Proc Natl Acad Sci USA 2008; 105: 2646–2651.
Usuki F, Ishiura S . Expanded CTG repeats in myotonin protein kinase increase susceptibility to oxidative stress. Neuroreport 1998; 9: 2291–2296.
Usuki F, Ishiura S, Saitoh N, Sasagawa N, Sorimachi H, Kuzume H et al. Expanded CTG repeats in myotonin protein kinase suppresses myogenic differentiation. Neuroreport 1997; 8: 3749–3753.
Amack JD, Mahadevan MS . The myotonic dystrophy expanded CUG repeat tract is necessary but not sufficient to disrupt C2C12 myoblast differentiation. Hum Mol Genet 2001; 10: 1879–1887.
Amack JD, Mahadevan MS . Myogenic defects in myotonic dystrophy. Dev Biol 2004; 265: 294–301.
Amack JD, Paguio AP, Mahadevan MS . Cis and trans effects of the myotonic dystrophy (DM) mutation in a cell culture model. Hum Mol Genet 1999; 8: 1975–1984.
Kuyumcu-Martinez NM, Wang GS, Cooper TA . Increased steady-state levels of CUGBP1 in myotonic dystrophy 1 are due to PKC-mediated hyperphosphorylation. Mol Cell 2007; 28: 68–78.
Savkur RS, Philips AV, Cooper TA . Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat Genet 2001; 29: 40–47.
Jacobs AE, Benders AA, Oosterhof A, Veerkamp JH, van MP, Wevers RA et al. The calcium homeostasis and the membrane potential of cultured muscle cells from patients with myotonic dystrophy. Biochim Biophys Acta 1990; 1096: 14–19.
Benders AA, Timmermans JA, Oosterhof A, Ter Laak HJ, van Kuppevelt TH, Wevers RA et al. Deficiency of Na+/K(+)-ATPase and sarcoplasmic reticulum Ca(2+)-ATPase in skeletal muscle and cultured muscle cells of myotonic dystrophy patients. Biochem J 1993; 293: 269–274.
Benders AA, Wevers RA, Veerkamp JH . Ion transport in human skeletal muscle cells: disturbances in myotonic dystrophy and Brody's disease. Acta Physiol Scand 1996; 156: 355–367.
Cardani R, Baldassa S, Botta A, Rinaldi F, Novelli G, Mancinelli E et al. Ribonuclear inclusions and MBNL1 nuclear sequestration do not affect myoblast differentiation but alter gene splicing in myotonic dystrophy type 2. Neuromuscul Disord 2009; 19: 335–343.
Furling D, Lemieux D, Taneja K, Puymirat J . Decreased levels of myotonic dystrophy protein kinase (DMPK) and delayed differentiation in human myotonic dystrophy myoblasts. Neuromuscul Disord 2001; 11: 728–735.
Furling D, Lam LT, Agbulut O, Butler-Browne GS, Morris GE . Changes in myotonic dystrophy protein kinase levels and muscle development in congenital myotonic dystrophy. Am J Pathol 2003; 162: 1001–1009.
Timchenko NA, Iakova P, Cai ZJ, Smith JR, Timchenko LT . Molecular basis for impaired muscle differentiation in myotonic dystrophy. Mol Cell Biol 2001; 21: 6927–6938.
Buj-Bello A, Furling D, Tronchere H, Laporte J, Lerouge T, Butler-Browne GS et al. Muscle-specific alternative splicing of myotubularin-related 1 gene is impaired in DM1 muscle cells. Hum Mol Genet 2002; 11: 2297–2307.
Kobayashi T, Askanas V, Saito K, Engel WK, Ishikawa K . Abnormalities of aneural and innervated cultured muscle fibers from patients with myotonic atrophy (dystrophy). Arch Neurol 1990; 47: 893–896.
Shimokawa M, Ishiura S, Kameda N, Yamamoto M, Sasagawa N, Saitoh N et al. Novel isoform of myotonin protein kinase: gene product of myotonic dystrophy is localized in the sarcoplasmic reticulum of skeletal muscle. Am J Pathol 1997; 150: 1285–1295.
Kameda N, Ueda H, Ohno S, Shimokawa M, Usuki F, Ishiura S et al. Developmental regulation of myotonic dystrophy protein kinase in human muscle cells in vitro. Neuroscience 1998; 85: 311–322.
Furling D, Coiffier L, Mouly V, Barbet JP, St Guily JL, Taneja K et al. Defective satellite cells in congenital myotonic dystrophy. Hum Mol Genet 2001; 10: 2079–2087.
Salisbury E, Sakai K, Schoser B, Huichalaf C, Schneider-Gold C, Nguyen H et al. Ectopic expression of cyclin D3 corrects differentiation of DM1 myoblasts through activation of RNA CUG-binding protein, CUGBP1. Exp Cell Res 2008; 314: 2266–2278.
Timchenko NA, Patel R, Iakova P, Cai ZJ, Quan L, Timchenko LT . Overexpression of CUG triplet repeat-binding protein, CUGBP1, in mice inhibits myogenesis. J Biol Chem 2004; 279: 13129–13139.
Sabourin LA, Tamai K, Narang MA, Korneluk RG . Overexpression of 3′-untranslated region of the myotonic dystrophy kinase cDNA inhibits myoblast differentiation in vitro. J Biol Chem 1997; 272: 29626–29635.
Martinuzzi A, Vergani L, Carrozzo R, Fanin M, Bartoloni L, Angelini C et al. Expression of muscle-type phosphorylase in innervated and aneural cultured muscle of patients with myophosphorylase deficiency. J Clin Invest 1993; 92: 1774–1780.
Martinuzzi A, Askanas V, Kobayashi T, Engel WK, Di MS . Expression of muscle-gene-specific isozymes of phosphorylase and creatine kinase in innervated cultured human muscle. J Cell Biol 1986; 103: 1423–1429.
Taneja KL, McCurrach M, Schalling M, Housman D, Singer RH . Foci of trinucleotide repeat transcripts in nuclei of myotonic dystrophy cells and tissues. J Cell Biol 1995; 128: 995–1002.
Davis BM, McCurrach ME, Taneja KL, Singer RH, Housman DE . Expansion of a CUG trinucleotide repeat in the 3′ untranslated region of myotonic dystrophy protein kinase transcripts results in nuclear retention of transcripts. Proc Natl Acad Sci USA 1997; 94: 7388–7393.
Adams V, Gielen S, Hambrecht R, Schuler G . Apoptosis in skeletal muscle. Front Biosci 2001; 6: D1–D11.
Sarbassov DD, Ali SM, Sabatini DM . Growing roles for the mTOR pathway. Curr Opin Cell Biol 2005; 17: 596–603.
Mizushima N, Levine B, Cuervo AM, Klionsky DJ . Autophagy fights disease through cellular self-digestion. Nature 2008; 451: 1069–1075.
Mammucari C, Milan G, Romanello V, Masiero E, Rudolf R, Del PP et al. FoxO3 controls autophagy in skeletal muscle in vivo. Cell Metab 2007; 6: 458–471.
Zhao J, Brault JJ, Schild A, Cao P, Sandri M, Schiaffino S et al. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab 2007; 6: 472–483.
Cornelison DD . Context matters: in vivo and in vitro influences on muscle satellite cell activity. J Cell Biochem 2008; 105: 663–669.
Bigot A, Klein AF, Gasnier E, Jacquemin V, Ravassard P, Butler-Browne G et al. Large CTG repeats trigger p16-dependent premature senescence in myotonic dystrophy type 1 muscle precursor cells. Am J Pathol 2009; 174: 1435–1442.
Thornell LE, Lindstom M, Renault V, Klein A, Mouly V, Ansved T et al. Satellite cell dysfunction contributes to the progressive muscle atrophy in myotonic dystrophy type 1. Neuropathol Appl Neurobiol 2009; 35: 603–613.
Sandri M, Carraro U . Apoptosis of skeletal muscles during development and disease. Int J Biochem Cell Biol 1999; 31: 1373–1390.
Biral D, Jakubiec-Puka A, Ciechomska I, Sandri M, Rossini K, Carraro U et al. Loss of dystrophin and some dystrophin-associated proteins with concomitant signs of apoptosis in rat leg muscle overworked in extension. Acta Neuropathol 2000; 100: 618–626.
Sandri M, El Meslemani AH, Sandri C, Schjerling P, Vissing K, Andersen JL et al. Caspase 3 expression correlates with skeletal muscle apoptosis in Duchenne and facioscapulo human muscular dystrophy. A potential target for pharmacological treatment? J Neuropathol Exp Neurol 2001; 60: 302–312.
Angelin A, Tiepolo T, Sabatelli P, Grumati P, Bergamin N, Golfieri C et al. Mitochondrial dysfunction in the pathogenesis of Ullrich congenital muscular dystrophy and prospective therapy with cyclosporins. Proc Natl Acad Sci USA 2007; 104: 991–996.
Irwin WA, Bergamin N, Sabatelli P, Reggiani C, Megighian A, Merlini L et al. Mitochondrial dysfunction and apoptosis in myopathic mice with collagen VI deficiency. Nat Genet 2003; 35: 367–371.
Mathieu J, De BM, Prevost C, Boily C . Myotonic dystrophy: clinical assessment of muscular disability in an isolated population with presumed homogeneous mutation. Neurology 1992; 42: 203–208.
Sandri M, Sandri C, Gilbert A, Skurk C, Calabria E, Picard A et al. Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 2004; 117: 399–412.
Acknowledgements
This work was supported by grants from ‘Progetti Ricerca di Interesse Nazionale-Ministero Istruzione-Università-Ricerca grant number 2005064759’ to L Vergani; ‘Progetti di Eccellenza’ Fondazione Cariparo, 2008–09 to C Angelini; ‘Association Française contre les Myopathies, grant number 13360’ to C Angelini; and ‘Telethon grant number GGP07250’ to G Novelli. AM was supported by the University of Padua. Muscle samples were provided by Telethon Biobank no. GTF05003.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by RA Knight
Supplementary Information accompanies the paper on Cell Death and Differentiation website
Rights and permissions
About this article
Cite this article
Loro, E., Rinaldi, F., Malena, A. et al. Normal myogenesis and increased apoptosis in myotonic dystrophy type-1 muscle cells. Cell Death Differ 17, 1315–1324 (2010). https://doi.org/10.1038/cdd.2010.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2010.33
Keywords
This article is cited by
-
Quantitative magnetic resonance imaging assessment of muscle composition in myotonic dystrophy mice
Scientific Reports (2023)
-
Defined d-hexapeptides bind CUG repeats and rescue phenotypes of myotonic dystrophy myotubes in a Drosophila model of the disease
Scientific Reports (2021)
-
miR-322/-503 rescues myoblast defects in myotonic dystrophy type 1 cell model by targeting CUG repeats
Cell Death & Disease (2020)
-
MBNL1 reverses the proliferation defect of skeletal muscle satellite cells in myotonic dystrophy type 1 by inhibiting autophagy via the mTOR pathway
Cell Death & Disease (2020)
-
Abnormal nuclear aggregation and myotube degeneration in myotonic dystrophy type 1
Neurological Sciences (2019)