Abstract
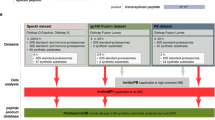
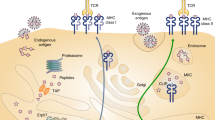
In the processing and presentation of antigenic peptides bound by the major histocompatibility complex (MHC) class I molecule, the ubiquitin-proteasome system of the eukaryotic cells plays an important role in proteolysis and degradation. The ubiquitinated protein substrate is delivered into the 26S proteasome to be digested and degraded. The proteasome degrading substrate is actually protein-protein interactions. Some researches of predicting proteasome cleave site rarely gave the information of the proteasome interacting with its substrate, and so the accuracy and reliability of these proteasome cleavage predictive methods still need to be improved. This paper used support vector machine method (SVM) to predict the proteasomal cleavage sites, and the predictive accuracy of the model was 82.8%. We showed analytically that the cleavage specificities of the cleavage sites “ | ” and its adjacent positions, and gave the information about the proteasome interacting with its substrate from our research results. It demonstrates that the proteasome cleaving to target protein is selective, but not random.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, T., Liu, W., Song, Z. et al. Computational Prediction of the Specificities of Proteasome Interaction with Antigen Protein. Cell Mol Immunol 6, 135–142 (2009). https://doi.org/10.1038/cmi.2009.19
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/cmi.2009.19