Abstract
Pollen germination on the surface of compatible stigmatic tissues is an essential step for plant fertilization. Here we report that the Arabidopsis mutant bcl1 is male sterile as a result of the failure of pollen germination. We show that the bcl1 mutant allele cannot be transmitted by male gametophytes and no homozygous bcl1 mutants were obtained. Analysis of pollen developmental stages indicates that the bcl1 mutation affects pollen germination but not pollen maturation. Molecular analysis demonstrates that the failure of pollen germination was caused by the disruption of AtBECLIN 1. AtBECLIN 1 is expressed predominantly in mature pollen and encodes a protein with significant homology to Beclin1/Atg6/Vps30 required for the processes of autophagy and vacuolar protein sorting (VPS) in yeast. We also show that AtBECLIN 1 is required for normal plant development, and that genes related to autophagy, VPS and the glycosylphosphatidylinositol anchor system, were affected by the deficiency of AtBECLIN 1.
Similar content being viewed by others
Introduction
Eukaryotic cells have evolved two major sophisticated intracellular constituent recycling mechanisms to maintain homeostasis, i.e., the ubiquitin-mediated proteosome system that commits degradation of specific, short-lived proteins, and autophagy that takes care of the turnover of bulk cytoplasmic contents including less-specific, long-lived proteins and even organelles 1, 2. In autophagy, cytoplasmic constituents are sequestered in double-membraned vesicles (designated as autophagosomes) and delivered to the vacuole/lysosome for degradation. Molecular genetic analysis in yeast showed that the biosynthetic cytoplasm to vacuole targeting (Cvt) pathway was topologically and morphologically similar to the degradative autophagy pathway, and identified at least 27 autophagy-related (ATG) genes involved in these two pathways 3, 4, 5, 6.
The orthologs of most of these yeast Atg proteins have been found in animals and plants, indicating that the autophagic process is highly conserved in yeast and higher multi-cellular organisms. It was reported that autophagy plays essential roles in response to starvation stress. For example, nutrient starvation induces a high level of autophagy and mutations of some ATG genes affect the differentiation process of sporulation under nutrient limitation conditions in yeast. Autophagy was also shown to be involved in normal development. Disruption of the Atg6 ortholog in nematodes arrests larval development at early stages 7 and a loss-of-function mutation in Beclin 1/Atg6 is embryonic lethal in mice 8. In addition, autophagy is involved in human diseases including cancer, muscular disorders and neurodegenerative diseases 8, 9, 10. For instance, decreased Beclin 1 expression results in high frequency of breast, ovarian and prostate cancers 8, 11. Until recently, however, very little was known about autophagy in plants. In Arabidopsis, there are at least 36 ATG-like genes, some of which have been characterized by genetic analysis 6. Disruption of either AtAtg7, AtAtg9, AtAtg3, AtAtg4a/b, AtAtg5, AtAtg18a or AtVTI12 results in similar phenotypes, such as increased chlorosis, premature leaf senescence and hyposensitivity to limited nitrogen or carbon nutrition 12, 13, 14, 15, 16, 17. This suggests that these ATG-like genes are required for normal senescence and nutrient recycling. Recently, it was reported that NbBECLIN 1 was essential for restriction of the hypersensitive response (HR) and programmed cell death (PCD) during disease resistance 6, 18, 19. However, to date there is no evidence that ATG-like genes are involved in pollen germination.
In higher plants, pollen, i.e., the male gametophyte, develops in the anthers, and becomes mature pollen after the microgametogenesis phase and two consecutive mitoses. On a compatible stigma, the desiccated pollen hydrates for germination; the pollen tube protrudes from the pollen grain and rapidly grows through the style and enters into the ovule to achieve double fertilization 20, 21. Many mutations that interfere with the production of pollen have been obtained and described in many plant species. Most of these mutants show a sporophytic requirement and are nuclear recessive, and thus the mutant locus can be transmitted by pollen or embyo sacs to form homozygous progeny 22, 23. By contrast, mutations of genes essential for pollen function can only be transmitted by the embryo sac, and thus can only be recovered as heterozygotes 24. These mutations show non-Mendelian segregation and this segregation distortion has been used successfully to identify several pollen mutants, such as kinky, limpet pollen (now named ingressus) and halfman 25, 26, 27, 28, 29. Assiduous screening based on morphology or staining characteristics of pollen has successfully isolated additional male gametophytic mutants, such as sidecar pollen, gemini pollen, raring-to-go (rtg), gum and mud 30, 31, 32, 33, 34. In addition to mutant screens, analysis of T-DNA insertion lines, RNA interference lines or antisense lines corresponding to particular genes has also been adopted to determine the function of the genes expressed in pollen 35, 36, 37. For instance, the Arabidopsis gene NPG1 encoding a pollen-specific calmodulin-binding protein was found to be essential for pollen germination 38, whereas a plant Rho-like GTPase (Rop), a plasma membrane Ca2+ pump ACA9 and the homolog of yeast Vps52p are shown to play important roles in pollen tube growth 39, 40, 41. Mutations in ADL1C, encoding a dynamin-like protein required for plasma membrane functions 42, and in an apyrase gene, are shown to result in an inability of the homozygous pollen to germinate 43. Other examples include the glycosylphosphatidylinositol (GPI) biosynthetic pathway genes SETH1 and SETH2 and an inositol polyphosphate kinase gene AtIPK2α that are all required for pollen germination and tube growth 44, 45.
In this study, ∼8 000 independent Arabidopsis transformants from our T-DNA mutant collection 46 were screened by determining the segregation of the seedling DL-Phosphinothricin (PPT)-resistance phenotype conferred by the T-DNA. Here we describe the isolation, genetic transmission, and phenotypic characterization of the bcl1 mutant. Phenotypic analysis showed that the bcl1 mutant appears to produce structurally and morphologically normal pollen grains, but that bcl1 pollen fails to germinate. The pollen germination defects of bcl1 are caused by the disruption of the gene At3g61710, which encodes a protein homologous to Atg6/Vps30/Beclin 1 in yeast or mammals. We also observed that AtBECLIN 1-deficent plants displayed retarded growth, dwarfism and early senescence. These results suggest that AtBECLIN 1 plays an essential role in both pollen germination and normal plant development.
Materials and Methods
Plant material, growth conditions and mutant screening
The T-DNA insertion mutant collection was generated as described previously 46. The first selfing progeny of the T1 generation were grown on the ½MS medium containing 10 μg/mL of PPT for a week and segregation ratios for PPT-resistance were calculated to identify mutants with a gametophyte defect.
Primers for identification of T-DNA mutants
The flanking sequence of the T-DNA insertion site was identified by thermal asymmetric interlaced PCR (TAIL-PCR) 47. The arbitrary degenerate primers and specific primers used in TAIL-PCR were as described previously 46. The primers P1 (5′-GAG CAG GAA ACA ACA ACC AAC T-3′), P2 (5′-GGC CTT GGA GAA TTA GGA TTG G-3′), LS4 (5′- TTG GTA ATT ACT CTT TCT TTT CCT CC -3′) and LBb1 (5′-GCG TGG ACC GCT TGC TGC AAC T-3′) were designed for co-segregation analysis of bcl1 and SALK_109281.
Plant crosses
For genetic transmission analysis, heterozygous bcl1 was reciprocally crossed with wild-type. The seeds obtained were grown on ½MS medium plus 10 μg/mL PPT for a week and the number of PPT-resistant and PPT-sensitive seedlings were calculated for transmission efficiency 27.
To generate the +/bcl1 qrt1/qrt1 double mutant, the flowers of heterozygous bcl1 plants were emasculated before maturation and 2 d later pollinated with qrt1 pollen. F2 progeny were screened on ½MS medium with 10 μg/mL of PPT and pollen from PPT-resistant plants was examined under the microscope for the qrt1 phenotype.
Constructs and transformation
The open reading frame (ORF) of ATBECLIN 1 was amplified from Arabidopsis pollen cDNA with primers BCL-1 (5′- ATG AGG AAA GAG GAG ATT CCA G -3′) and BCL-2 (5′- GAT TCA GTC TAT CAC GAG TTT CTA AGT-3′), and BCL-1 and BCL-3 (5′-AGT TTT TTT ACA TGA AGG CTT ACT AGA GTC-3′) were used for amplifying the ATBECLIN 1 coding region without a stop codon. The ATBECLIN 1 promoter was amplified from Arabidopsis genomic DNA with primers BCP-1 (5′-CAT GTT CTG AGA CAA TTC TCT AGC-3′) and BCP-2 (5′-CTC CTC TTT CCT CAT CGC TCT C-3′). GFP was amplified from pIRES-GFP (CLONTECH) with primers GFP-1 (5′- ATG GTA GAT CTG ACT AGT AAA GG -3′) and GFP-2 (5′- AGC TGG TCA CCA ATT CAC ACG T -3′). The products were cloned into the EcoR V site of pBluescript SK+ (pBS) in both sense and anti-sense orientations. The pBS plasmid carrying sense ATBECLIN 1 (pBCL1) was digested by Xba I and Kpn I and cloned into the plant binary vector fragment pQG110 48 to generate p35SPBCL. pBCPBCLGFP was constructed by the ligation of pQG110/Hind III-Xba I, pBS plasmids carrying the antisense ATBECLIN 1 promoter/Hind III-Pst I, pBS plasmids carrying sense ATBECLIN 1 without the stop codon/Pst I-Sal I and pBS plasmids carrying antisense GFP/Sal I-Xba I. The p35SPBCL and pBCPBCLGFP plasmids were transformed into +/bcl1 and T0 seeds were screened on the ½MS medium supplemented with PPT and kanamycin.
For expression pattern analysis, a 782-bp promoter sequence of ATBECLIN 1 was amplified with primers BCP-1 and BCP-3 (5′-CGC TCT CCC AGT TTT TGT GG-3′) and cloned into pBS/EcoR V. The construct pBCPGUS was obtained by ligating EcoR I- Hind III pCAMBIA1381Xa with EcoR I- Hind III pBS with a sense insert of the ATBECLIN 1 promoter sequence. The pBCPGUS was transformed into wild-type Arabidopsis and T0 seeds were screened on the ½MS medium with hygromycin.
Phenotypic analysis by microscopy, cytochemistry and pollen germination
The inflorescences from wild-type and +/bcl1 heterozygous plants were collected and fixed in 4% (v/v) glutaraldehyde in 25 mM sodium phosphate buffer (pH6.8). After dehydration through a concentration-increasing ethanol series, the samples were embedded into Historesin (Leica, Wetzlar, Germany). Sectioning was performed using a Leica microtome and the sections mounted on slides. Sections of 5 μm were stained with 0.25% (w/v) toluidine blue O (Sigma) and observed under an Olympus BX51 microscope (Olympus, Tokyo, Japan). Digital images were captured with a SPOT camera (Diagnostic Instruments, Inc., Sterling Heights, MI, USA) and processed using Adobe Photoshop.
For scanning electron microscopy (SEM), mature flowers of +/bcl1 qrt1/qrt1 were fixed in pure methanol for 5 min and then washed with 100% ethanol before being mounted and coated as described 49. SEM analysis was carried out using a Hitachi S-450 scanning electron microscope (Hitachi, Japan).
For cytochemical analysis of pollen, solutions of 4,6-diamidino-2-phenylindole (DAPI) and rhodamine 123 (Sigma) were prepared and used as described previously 50, and Alexander stain was prepared also as described previously 51. Mature pollen were stained with DAPI, Alexander stain or rhodamine 123 respectively, and observed with epifluorescence or differential interference contrast (DIC) under an Olympus BX51 microscope (Olympus, Tokyo, Japan). More than 3 000 pollen tetrads from at least ten different mutant lines were examined. Pollen tubes in pistils were stained using aniline blue as described previously 35.
In vitro pollen germination was assayed either in liquid medium for pollen tetrads from +/bcl1 qrt1/qrt1 or qrt1/qrt1 mutants or on solid medium. The germination medium was used as described previously 52. For statistical analysis of the germination ratios, each assay was repeated at least three times, and pollen grains were collected from more than 10 different plants each time. Pollen grains were incubated at 25 ºC and 100% humidity on solid medium for 10 h. Pollen tetrads were germinated in 10 μL liquid medium, dropped on slides, and incubated at 25 °C and 100% humidity for 20 h.
Affymetrix microarray analysis
Wild-type and pBCPBCLGFP bcl1/bcl1 plants were grown for about 45 d under the same conditions as those described above. 30-40 homozygous bcl1 plants harboring pBCPBCLGFP or wild-type plants, without roots, were pooled and homogenized in liquid nitrogen. Total RNA was isolated independently from three aliquots of 100 mg tissue separated from homogenized tissues using TRIzol reagent (Invitrogen). RNA was purified using RNeasy mini kits (Qiagen). RNA from three independent purifications was pooled and used for microarray hybridization with Arabidopsis whole genomic microarrays (ATH1, Affymetrix) according to manufacturer's instructions. Data images were analyzed using Affymetrix Microarray Suit (version 5.0) software. After invariant normalization, log2 ratio analysis was carried out and genes showing a significant difference (P<0.05) between wild-type and pBCPBCLGFP bcl1/bcl1 plants were selected. Biochemical pathways were analyzed using the software KOBAS 53.
Real-time RT-PCR analysis
Roots, stems, leaves, flowers and siliques were collected from about 45-day-old Arabidopsis (Col-0) plants. Mature pollen was collected by washing the opening flowers with water, filtering through a 6-micron mesh, and centrifugation. After treatment with RNase-free DNase (TaKaRa, Japan), 5 μg of total RNA was reverse-transcribed in a reaction of 20 μL using Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA). The first strand cDNA was used as the template for real-time PCR amplification. Specific primer pairs used in real-time PCR were designed as shown in Supplementary information Table S1. Real-time PCR amplification was performed on a MJ Research thermocycler using the DyNAmo SYBR Green qPCR kit (Finnzymes) 54. The expression levels of different genes were normalized to the constitutive expression level of UBQ11. Each real-time PCR experiment with different primer pairs and/or different templates was replicated three times. The relative gene expression levels were calculated by the 2−ΔΔt method 55.
Results
The bcl1 mutant lacks functional pollen
We screened about 8 000 T2 lines from our T-DNA collection 46 for mutants defective in gametophyte development by analyzing the segregation of the T-DNA insertion that confers DL-Phosphinothricin (PPT)-resistance. One mutant line, later named bcl1, exhibited a segregation ratio of 1:1 (PPT-resistant:PPT-sensitive) for its progeny (Table 1), suggesting that the T-DNA is inserted in a gene essential for the formation of viable gametophytes. To determine which gametophyte, male or female, was affected in the mutant, reciprocal crosses between wild-type and the heterozygote mutant were conducted. As shown in Table 1, 95.6% of the mutated allele was passed to the next generation via the female gametes but none via the male gametes, indicating that the gene disrupted by the T-DNA insertion is specifically required for pollen development or viability.
AtBECLIN 1 encodes a protein with homology to Beclin 1/Atg6/Vps30
To analyze the molecular nature of bcl1, we identified the flanking sequences of the T-DNA insertion site by TAIL-PCR 47. The T-DNA was inserted in the second intron of the gene At3g61710 (Figure 1A) and Southern blots indicated that there was only a single T-DNA insertion in bcl1 (data not shown). To examine whether the T-DNA insertion co-segregated with the phenotype, the genotype of the T3 progeny was analyzed with primers that amplify an 837-bp fragment for the wild-type allele and a 600-bp fragment for the T-DNA insertion allele (Figure 1A). All of the T3 PPT-resistant plants tested were heterozygous (Figure 1B). The T4 progeny of these heterozygous plants were also found to display a 1:1 ratio (PPT-resistant:PPT-sensitive). These results indicate that the bcl1 mutant phenotype co-segregates with the T-DNA insertion in the gene At3g61710. We obtained an additional allele of bcl1, SALK_109281, that has a T-DNA insertion in the second intron of At3g61710, 54-bp away from the insertion site in bcl1 (Figure 1A). Like bcl1, the selfing progeny of SALK_109281 heterozygous plants exhibited a distorted segregation ratio of approximately 1:1 (415:422) for kanamycin-resistance (Table 1), and no homozygous progeny were obtained (data not shown). These results suggest that disruption of At3g61710 is correlated with a pollen-defective phenotype.
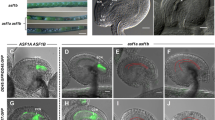
Identification of bcl1 and SALK_109281 mutants. (A) Schematic representation of T-DNA insertion sites in bcl1 and SALK_109281. Exons of AtBECLIN 1 are represented by blue boxes, introns by dark red boxes. The T-DNA insertion site in bcl1 mutant is indicated by a green “X” and that in SALK_109281 by a red diamond. The numbers 0, 803, 857 and 2 777 indicate the position of the “A” of the “ATG” start codon, the T-DNA insertion sites in bcl1 and SALK_109281, and the “G” of the “TAG” stop codon in the BCL1 gene, respectively. The bar and NPTII genes are represented by the purple and green boxes with arrows. The arrow direction indicates the transcriptional orientation of the genes. The red arrow tetrad represents the four 35S enhancers from pSKI015. The line arrows represent the primers used in co-segregation analysis. LB, T-DNA left border; bar, Basta resistance gene; 4 enhancer, CaMV 35S enhancer tetrad; RB, T-DNA right border; NPTII, kanamycin resistance gene. (B) Cosegregation analysis of the T-DNA insertion and the phenotype of distorted segregation of PPT resistance. P1 and P2 amplify an 837-bp fragment from wild-type while P1 and LS4 amplify a 600-bp fragment from the T-DNA insertion allele in the bcl1 mutant. Marker, λ EcoR I/Hind III marker. (C) Real-time quantitative PCR analysis of AtBECLIN 1 expression with the mature pollen cDNA from wild-type and +/bcl1. (D) Complementation of bcl1 phenotypes by transformation of AtBECLIN 1 cDNA driven by the CaMV 35S promoter or its own promoter. Upper part shows the schematic representation of constructs used in the transformation experiments. LB, T-DNA left border; polyA, CaMV 35S polyA; KanR, kanamycin resistance gene NPTII; 35S-P, CaMV 35S promoter; BCL1-P, promoter of the AtBECLIN 1 gene; BCL1, ORF of the AtBECLIN 1 gene without the stop codon; GFP, ORF of green fluorescent protein gene; Ter, NOS terminator; RB, T-DNA right border. Lower part shows genotyping of transgenic plants using PCR with primers P1, P2 and LS4. Homozygous bcl1 lines could be obtained in the progeny of p35SPBCL and pBCPBCLGFP transgenic plants. (E) Aniline blue staining indicates that the pollen of homozygous bcl1 lines harboring p35SPBCL can germinate in vivo, suggesting that AtBECLIN 1 can complement the pollen germination phenotype of bcl1. (F) The setting of seeds was unaffected in p35SPBCL bcl1/bcl1 plants.
To further investigate whether the pollen-defective phenotype is caused by the T-DNA insertion in At3g61710, we examined the expression level of At3g61710 in wild-type and heterozygous plants by real-time quantitative PCR. This experiment showed that the transcription level of At3g61710 was reduced by about half in heterozygous plants (Figure 1C). We then transformed heterozygous bcl1 plants with the coding region of At3g61710 driven by either a CaMV 35S promoter (construct designated as p35SPBCL) or the At3g61710 promoter (construct designated as pBCPBCLGFP) (Figure 1D). We obtained viable T2 transgenic plants in the homozygous bcl1 background, with both the p35SPBCL and pBCPBCLGFP constructs (Figure 1D), in which the pollen-defective phenotype had been rescued (Figure 1E) and seed fertility was normal (Figure 1F). These results indicate that the T-DNA insertion in At3g61710 is responsible for the pollen defects in the bcl1 mutant.
At3g61710 encodes a protein that is 69% identical to Nicotiana benthamiana NbBECLIN 1 18, 31% identical to a human tumor suppressor Hsbeclin 1 11, and 26% identical to a yeast autophagy-related protein Atg6/Vps30 56, 57. At3g61710 has been reported to be able to complement the autophagy-defective phenotype in Δatg6/vps30 yeast 18, and therefore is the ortholog of yeast Atg6/Vps30 in Arabidopsis.
Pollen germination rather than early pollen development was abolished in bcl1
In order to reveal the cause of the pollen defect in bcl1, we first examined pollen development through all stages in +/bcl1 heterozygous plants. It appeared that pollen development in +/bcl1, including meiosis of the microspore mother cell, tetrad formation, microspore release, and mitosis, was not affected (Figure 2A and 2B)58. These results suggest that the microgametogenesis phase of pollen ontogeny is not affected in the bcl1 mutant. To further confirm this conclusion, we crossed +/bcl1 heterozygous plants with the qrt1/qrt1 mutant, and obtained +/bcl1 qrt1/qrt1 plants for tetrad analysis. The qrt1/qrt1 mutant will produce fully functional pollen grains in the form of tetrads because the gene coding for a pectin methylesterase has been mutated and it has been used previously for identifying mutants with pollen defects 59, 60, 61. The anthers of +/bcl1 qrt1/qrt1 plants should produce tetrads with two wild-type ATBECLIN 1 and two mutated genes. SEM analysis showed that all four pollen grains in the tetrads from +/bcl1 qrt1/qrt1 plants are similar in appearance including the size and the shape (Figure 2C). In addition, neither DAPI (4′,6-diamidino-2-phenylindole) (Figure 2D) nor Alexander (Figure 2E) staining reveals any differences 51, and staining with rhodamine 123 showed no differences in the mitochondrial distribution and viability (Figure 2F) 50. These results clearly indicate that the bcl1 mutation does not affect meiosis and other steps of pollen development.
Phenotypic analysis of pollen from the bcl1 T-DNA insertion mutant. (A) Sections of anthers at different developmental stages from the bcl1 heterozygous mutant. Stage 6, stage 7, stage 9 and stage 11 were shown (from left to right) 58. (B) Sections of anthers of different developmental stages from wild-type plants. The stages shown were the same as those in (A). The pollen development of +/bcl1 plants, including meiosis of the microspore mother cell, tetrad formation, microspore release, and mitosis, is not different from that of wild-type. (C) SEM analysis of a pollen tetrad from +/bcl1 qrt1/qrt1 plants. (D) DAPI staining of a pollen tetrad from +/bcl1 qrt1/qrt1 plants. (E) Alexander's staining of a pollen tetrad from +/bcl1 qrt1/qrt1 plants. (F) RDM123 staining of a pollen tetrad from +/bcl1 qrt1/qrt1 plants. (G) and (H) In vitro germination of pollen from wild-type, +/bcl1, SALK_109281 and pBCPBCLGFP bcl1/bcl1 plants. (I) and (J) In vitro germination of pollen tetrads from qrt1/qrt1 and +/bcl1 qrt1/qrt1 plants. Tetrads from the qrt1/qrt1 mutant can generate one to four pollen tubes, whereas those from +/bcl1 qrt1/qrt1 plants can only produce one or two pollen tubes.
We then examined the germination ratio of mature pollen from wild-type, +/bcl1 and SALK_109281 heterozygous plants, and pBCPBCLGFP bcl1/bcl1 plants in vitro. In contrast to wild-type pollen for which the germination rate ranged from 61% to 75%, only 33% to 37% of pollen grains from +/bcl1 and SALK_109281 heterozygous plants germinated (Figure 2G and 2H). To further confirm this, we examined the germination of the tetrads of the qrt1/qrt1 mutant and +/bcl1 qrt1/qrt1 double mutant in vitro. As shown in Figure 2I and 2J, out of the hundreds of tetrads we analyzed, we found that each tetrad from the qrt1/qrt1 mutant could generate one to four pollen tubes, whereas tetrads from +/bcl1 qrt1/qrt1 plants could only produce one or two pollen tubes, suggesting that pollen germination is indeed affected in the bcl1 mutants.
ATBECLIN 1 is predominantly expressed in pollen
To examine the expression pattern of the AtBECLIN 1 gene, a 782-bp promoter fragment of AtBECLIN 1 was amplified to drive the expression of β-glucuronidase (GUS), and this construct was transformed into Arabidopsis. The results showed that, although some activity was detected in the germinating seeds and hypocotyls (Figure 3A and 3B), strong GUS activity was detected in anthers and mature pollen (Figure 3C and 3D). Real-time quantitative RT-PCR analysis further confirmed that the expression level of AtBECLIN 1 is highest in mature pollen among the tissues examined (Figure 3E), suggesting that AtBECLIN 1 is expressed in mature pollen preferentially.
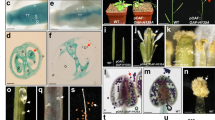
The expression pattern of AtBECLIN 1. (A) to (D) GUS staining of plants transformed with a construct in which the GUS gene is driven by the AtBECLIN 1 promoter. (E) Expression analysis of AtBECLIN 1 in mature pollen and other organs by real time quantitative PCR. AtBECLIN 1 is expressed predominantly in mature pollen.
AtBECLIN 1 is essential for plant development
Although the heterozygous bcl1 and SALK_109281 mutants grew normally, some pBCPBCLGFP transgenic plants in the bcl1/bcl1 background, i.e., pBCPBCLGFP bcl1/bcl1, displayed a range of phenotypic abnormalities including short roots, early leaf senescence, small leaves, dwarfism, fewer flowers and low fertility (Figure 4A–4D). Real-time quantitative RT-PCR analysis revealed that the expression level of AtBECLIN 1 was three-fold lower in these pBCPBCLGFP bcl1/bcl1 lines compared to wild-type (Figure 4E). These data indicate that, in addition to delaying leaf senescence as seen for NbBECLIN 1 in tobacco 18, AtBECLIN 1 is also important for other development processes such as leaf development and fertility (Figure 4A–4D).
Characterization of an AtBECLIN 1-deficient Arabidopsis mutant. (A) 7-day-old wild-type and pBCPBCLGFP bcl1/bcl1 (AtBECLIN 1-deficient) plants. The AtBECLIN 1-deficient plants display shorter roots. (B) 50-day-old wild-type and AtBECLIN 1-deficient plants. The AtBECLIN 1-deficient plants display earlier leaf senescence, smaller leaves, dwarfism, and fewer flowers. (C) 30-day-old wild-type and AtBECLIN 1-deficient plants. (D) Rosette leaves from 30-day-old wild-type (upper) and AtBECLIN 1-deficient plants (bottom). AtBECLIN 1-deficient plants produced less leaves and these leaves showed earlier senescence. (E) Expression of the AtBECLIN 1 gene was determined using real-time quantitative PCR. Expression of AtBECLIN 1 in pBCPBCLGFP bcl1/bcl1 plants was about three-fold lower than in wild-type. Bars=5 mm in (B) and 1 mm in all other panels.
Alteration of gene expression caused by the deficiency of AtBECLIN 1
In order to investigate the molecular mechanism behind the phenotype caused by loss of AtBECLIN 1 function, we performed microarray analysis on pBCPBCLGFP bcl1/bcl1 plants which displayed severe growth and developmental defects including early leaf senescence and low fertility (Figure 4) using the Arabidopsis whole genome Affymetrix microarray chip (see methods). Real-time RT-PCR analysis showed that the expression of pathogenesis-related genes such as PR1, EDS1, EDS5 and SEN1 was greatly increased, confirming the fidelity of the microarray data (Supplementary information, Table S2, Figure 5A, 5B and 5E) 18. The microarray data showed that 2 184 genes were up-regulated and 886 genes were down-regulated in the pBCPBCLGFP bcl1/bcl1 plants, based on a two-fold threshold, compared to wild-type plants (Supplementary information, Table S3 and S4).
Real time quantitative PCR confirmation of gene expression changes identified by microarray analysis between wild-type and pBCPBCLGFP bcl1/bcl1 plants. (A) and (B) Selected pathogenesis-related genes were found to be up-regulated in homozygous bcl1 lines harboring pBCPBCLGFP. (C) and (D) Expression of AtVps34, AtVps15 and some ATG-like genes is up-regulated in pBCPBCLGFP bcl1/bcl1 plants. (E) Expression of SEN1 is up-regulated in pBCPBCLGFP bcl1/bcl1 plants. This result is consistent with the early leaf senescence phenotype in AtBECLIN 1-deficient plants. (F) SETH1 and GPI anchor protein genes are down-regulated in pBCPBCLGFP bcl1/bcl1 plants.
Because AtBECLIN 1 is the ortholog of Atg6/Vps30 which is essential for autophagy and vacuolar protein sorting (VPS), we first focused on analyzing those genes involved in membrane trafficking. As shown in Table 2, fourteen VPS-related genes including AtVps34 and AtVps15 (Figure 5C) and eight exocytosis-related genes including SEC8 were up-regulated in the AtBECLIN 1-deficient lines compared to wild-type (Table 2, Figure 5D), suggesting that membrane trafficking including VPS and exocytosis is affected. Taking into consideration the fact that disruption of a VPS-related gene Vps52 41 or an exocytosis-related gene SEC8 62 leads to pollen germination and pollen tube growth defects, the altered expression levels of VPS and/or exocytosis-related genes might contribute to the pollen germination defect in bcl1 mutant. Interestingly, all but one of the eleven autophagy-related genes were also found to be up-regulated when AtBECLIN 1 expression was low (Table 2). Genes involved in phosphatidylinositol (PI) metabolism and signaling were also significantly disturbed in pBCPBCLGFP bcl1/bcl1 plants. As shown in Table 3, sixty-six of these genes were up-regulated. We further examined the expression of genes related to the glycosylphosphatidylinositol (GPI) anchor system, which was previously reported to be essential for pollen germination 44. SETH1, one of the genes encoding a GPI anchor biosynthetic enzyme, was found to be down-regulated (Table 4, Figure 5F). Furthermore, out of the 47 GPI anchor protein (GAP) genes expressed in mature pollen 44, sixteen including AtAGP22, FLA12, GPDL and COBL8 were down-regulated whereas five genes were up-regulated (Table 4, Figure 5F). Taken together, these results suggest that, in addition to autophagy and VPS-related genes being affected by bcl1, the alteration in both PI metabolism and signaling and the GPI anchor system may also contribute to the failure of pollen germination and abnormal plant development in bcl1 mutants.
Discussion
Autophagy is an important intracellular process that recycles the cytoplasmic constituents including organelles either for growth under hostile conditions or for normal development 2, 3, 9. Although well demonstrated in yeasts and animals, knowledge about autophagy in plants is limited 3. In this paper, we have identified an Arabidopsis insertion mutant in which the AtBECLIN 1/AtAtg6 gene, the ortholog of the yeast autophagic gene Atg6, was mutated. The failure of bcl1 pollen to germinate and the retarded growth of AtBECLIN 1-deficient mutants suggested that this gene is essential for pollen germination and normal plant development.
In yeast, Atg6 plays important roles in recruiting autophagic proteins including Atg18, Atg20, Atg24 and Atg27 to the pre-autophagosome structure (PAS) 4. Loss of function of the BECLIN 1/Atg6 ortholog results in embryo lethality in mice and nematodes 8, 63. BECLIN 1/Atg6 orthologs are also required for normal dauer morphogenesis and life-span extension in nematodes 7. Liu et al. (2005) reported that NbBECLIN 1-silenced tobacco, generated using the VIGS method, exhibits leaf senescence and HR PCD outside the infection site during TMV inoculation. In this paper, we demonstrated that AtBECLIN 1 is essential for pollen germination (Figure 2G and 2H). This is a new finding compared to other reports about the functions of ATG-like genes in Arabidopsis. T-DNA insertion in the AtAtg7, AtAtg9, VTI12 and AtAtg5 genes led in all cases to accelerated leaf senescence under nutrient-limiting conditions, but pollen germination was not affected 12, 13, 14, 17. The double mutant atg4a4b-1 and the RNAi-silenced AtATG18a Arabidopsis plants displayed similar phenotypes 15, 16. NbBECLIN 1-silenced tobacco plants also exhibit earlier leaf senescence 18. None of these mutants were found to have pollen defects, except that atatg9-1 mutant plants had reduced seed productivity under nitrogen-deficient conditions 13. These results imply that autophagy may not contribute to the pollen germination defects in +/bcl1 plants. In yeast, Atg6, together with Vps34 and Vps15, forms two PI 3-kinase complexes with either Atg14 or Vps38 and these two complexes play important roles in autophagy or VPS, respectively 64, 65, 66, 67, 68, 69. It was previously reported that VPS and exocytosis were involved in pollen germination and tube growth 41, 62. Our microarray data showed that many VPS- and exocytosis-related genes were up-regulated in AtBECLIN 1-deficient plants (Table 2). Therefore, the pollen germination defect in bcl1 could possibly be attributed to the changes in VPS and exocytosis caused by loss-of-function in AtBECLIN 1/AtAtg6 and/or the corresponding PI 3-kinase complex.
Several ATG genes have been identified in plants in recent years, but the molecular mechanisms underpinning the phenotypes of loss of function mutants of these ATG genes have not been elucidated 12, 13, 14, 15, 16, 17. In this paper we intend to draw a map of transcriptional changes resulting from the deficiency of AtBECLIN 1 through Affimetrix chip analysis. The results showed that the expression of almost three thousand genes was altered. Dozens of pathogenesis-related genes were strongly up-regulated and this result is consistent with the fact that NbBECLIN 1 affects tobacco disease responses. However, it is unknown how AtBECLIN 1 regulates so many genes. Maybe some pathways such as the altered phosphatidylinositol metabolism and signaling pathway play an important role in the regulation process and result in pleiotropic phenotypes in the AtBECLIN 1-deficient plants. The previous studies showed that PI(4,5)P2 could conjugate with Rac GTPase in plant cells 70, 71. The plant Rho-like GTPase (Rop) and Rac-like GTPase were reported to be important for pollen tube growth and pollen germination 38, 39, 71, 72, 73, 74. Phosphoinositides such as PI(4,5)P2, InsP3, and inositol 1,3,4,5-tetrakisphosphate were also reported to be vital for pollen germination and pollen tube growth. Loss-of-function for either an inositol polyphosphate kinase gene AtIPK2α in Arabidopsis or PLC1 in Petunia inflate would result in defect in pollen germination or pollen tube growth, respectively 44, 45. In our microarray data, AtBECLIN 1-deficiency resulted in up-regulation of many genes encoding proteins involved in PI metabolism and signaling, e.g. PI 3-kinase, PI 4-kinase, PI-phosphate 5-kinase and PLC (Table 3), and down-regulation of two Rac-like GTP-binding protein genes RAC2 (AT5G45970) and RAC7 (AT4G28950) (Supplemental Table 4). These results suggest that many PI-related pathways have been affected by AtBECLIN 1 deficiency, which may all result in pollen germination defect. Some genes related to cellular trafficking were also affected in pBCPBCLGFP bcl1/bcl1 plants. During pollen germination and pollen tube growth, cellular trafficking is critical for cell wall deposition and cell shape remodeling 75, 76, 77. For example, inactivation of some syntaxins, which function in endomembrane transport vesicle fusion, resulted in inviable pollen 78. Vacuole biogenesis, VPS, and exocytosis are also reported to be essential for pollen germination and tube growth 41, 62, 79. It is thus possible that the alteration of the cellular trafficking system may contribute to the failure of pollen germination in bcl1 mutants.
The expression of AtBECLIN 1 is ubiquitous in roots, stems, leaves, flowers and siliques in addition to pollen and is already detectable in germinating seeds. This expression pattern implies that AtBECLIN 1 may play other important roles. The +/bcl1 mutant caused pollen malfunction, and thus the complete knock-out bcl1 homozygous mutant was not obtainable. The heterozygous bcl1 mutant is similar to wild-type at all stages of development and growth. Thus though we have identified a function for AtBECLIN 1 in pollen development, we can not reveal any other functions through the analysis of heterozygous bcl1 mutant plants. Fortunately, we found pBCPBCLGFP, bcl1/bcl1 plants had a lower level of expression of AtBECLIN 1 than heterozygous bcl1 mutants and displayed a pleiotropic phenotype including retarded growth rate, short roots, small leaves, early leaf senescence, dwarfism and low fertility under normal growth conditions. These results indicate that unlike other identified ATG genes, AtBENCLIN1/AtAtg6 plays more essential roles in both sporophytes and gametophytes of plants. This is consistent with previous studies showing that the AtBENCLIN1 ortholog is essential for the normal development of mice and nematodes 8, 63.
(Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Thompson AR, Vierstra RD . Autophagic recycling: lessons from yeast help define the process in plants. Curr Opin Plant Biol 2005; 8:165–173.
Nair U, Klionsky DJ . Molecular mechanisms and regulation of specific and nonspecific autophagy pathways in yeast. J Biol Chem 2005; 280:41785–41788.
Levine B, Klionsky DJ . Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004; 6:463–477.
Klionsky DJ . The molecular machinery of autophagy: unanswered questions. J Cell Sci 2005; 118:7–18.
Yorimitsu T, Klionsky DJ . Autophagy: molecular machinery for self-eating. Cell Death Differ 2005; 12 Suppl 2:1542–1552.
Seay M, Patel S, Dinesh-Kumar SP . Autophagy and plant innate immunity. Cell Microbiol 2006; 8:899–906.
Melendez A, Talloczy Z, Seaman M, et al. Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 2003; 301:1387–1391.
Yue Z, Jin S, Yang C, Levine AJ, Heintz N . Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA 2003; 100:15077–15082.
Kelekar A . Autophagy. Ann N Y Acad Sci 2006; 1066:259–271.
Komatsu M, Waguri S, Chiba T, et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 2006; 441:880–884.
Liang XH, Jackson S, Seaman M, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999; 402:672–676.
Doelling JH, Walker JM, Friedman EM, Thompson AR, Vierstra RD . The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence in Arabidopsis thaliana. J Biol Chem 2002; 277:33105–33114.
Hanaoka H, Noda T, Shirano Y, et al. Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an Arabidopsis autophagy gene. Plant Physiol 2002; 129:1181–1193.
Surpin M, Zheng H, Morita MT, et al. The VTI family of SNARE proteins is necessary for plant viability and mediates different protein transport pathways. Plant Cell 2003; 15:2885–2899.
Yoshimoto K, Hanaoka H, Sato S, et al. Processing of ATG8s, ubiquitin-like proteins, and their deconjugation by ATG4s are essential for plant autophagy. Plant Cell 2004; 16:2967–2983.
Xiong Y, Contento AL, Bassham DC . AtATG18a is required for the formation of autophagosomes during nutrient stress and senescence in Arabidopsis thaliana. Plant J 2005; 42:535–546.
Thompson AR, Doelling JH, Suttangkakul A, Vierstra RD . Autophagic nutrient recycling in Arabidopsis directed by the ATG8 and ATG12 conjugation pathways. Plant Physiol 2005; 138:2097–2110.
Liu Y, Schiff M, Czymmek K, et al. Autophagy regulates programmed cell death during the plant innate immune response. Cell 2005; 121:567–577.
Patel S, Caplan J, Dinesh-Kumar S . Autophagy in the control of programmed cell death. Curr Opin Plant Biol 2006; 9:391–396.
Leyser O, Day S . Mechanisms in plant development. Oxford, Blackwell, 2003.
Faure J E DC . Fertilization in flowering plants. New approaches for an old story. Plant Physiol 2001; 125:102–104.
Yadegari R, Drews GN . Female gametophyte development. Plant Cell 2004; 16 Suppl:S133–141.
McCormick S . Control of male gametophyte development. Plant Cell 2004; 16 Suppl:S142–153.
Johnson MA, von Besser K, Zhou Q, et al. Arabidopsis hapless mutations define essential gametophytic functions. Genetics 2004; 168:971–982.
Feldmann KA, Coury DA, Christianson ML . Exceptional segregation of a selectable marker (KanR) in Arabidopsis identifies genes important for gametophytic growth and development. Genetics 1997; 147:1411–1422.
Bonhomme S, Horlow C, Vezon D, et al. T-DNA mediated disruption of essential gametophytic genes in Arabidopsis is unexpectedly rare and cannot be inferred from segregation distortion alone. Mol Gen Genet 1998; 260:444–452.
Howden R, Park SK, Moore JM, et al. Selection of T-DNA-tagged male and female gametophytic mutants by segregation distortion in Arabidopsis. Genetics 1998; 149:621–631.
Grini PE, Schnittger A, Schwarz H, et al. Isolation of ethyl methanesulfonate-induced gametophytic mutants in Arabidopsis thaliana by a segregation distortion assay using the multimarker chromosome 1. Genetics 1999; 151:849–863.
Oh SA, Park SK, Jang I, et al. halfman, an Arabidopsis male gametophytic mutant associated with a 150 kb chromosomal deletion adjacent to an introduced Ds transposable element. Sex Plant Reprod 2003; 16:99–102.
Chen YCS, McCormick S . Sidecar pollen, an Arabidopsis thaliana male gametophytic mutant with aberrant cell divisions during pollen development. Development 1996; 122:3243–3253.
Park SK, Howden R, Twell D . The Arabidopsis thaliana gametophytic mutation gemini pollen1 disrupts microspore polarity, division asymmetry and pollen cell fate. Development 1998; 125:3789–3799.
Johnson SA, McCormick S . Pollen germinates precociously in the anthers of raring-to-go, an Arabidopsis gametophytic mutant. Plant Physiol 2001; 126:685–695.
Lalanne E, Twell D . Genetic control of male germ unit organization in Arabidopsis. Plant Physiol 2002; 129:865–875.
Twell D, Park SK, Hawkins TJ, et al. MOR1/GEM1 has an essential role in the plant-specific cytokinetic phragmoplast. Nat Cell Biol 2002; 4:711–714.
Muschietti J, Dircks L, Vancanneyt G, McCormick S . LAT52 protein is essential for tomato pollen development: pollen expressing antisense LAT52 RNA hydrates and germinates abnormally and cannot achieve fertilization. Plant J 1994; 6:321–338.
Gupta R, Ting JT, Sokolov LN, Johnson SA, Luan S . A tumor suppressor homolog, AtPTEN 1, is essential for pollen development in Arabidopsis. Plant Cell 2002; 14:2495–2507.
Alonso JM, Stepanova AN, Leisse TJ, et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 2003; 301:653–657.
Golovkin M, Reddy AS . A calmodulin-binding protein from Arabidopsis has an essential role in pollen germination. Proc Natl Acad Sci USA 2003; 100:10558–10563.
Arthur KM, Vejlupkova Z, Meeley RB, Fowler JE . Maize ROP2 GTPase provides a competitive advantage to the male gametophyte. Genetics 2003; 165:2137–2151.
Schiott M, Romanowsky SM, Baekgaard L, et al. A plant plasma membrane Ca2+ pump is required for normal pollen tube growth and fertilization. Proc Nat Acad Sci USA 2004; 101:9502–9507.
Lobstein E, Guyon A, Ferault M, et al. The putative Arabidopsis homolog of yeast vps52p is required for pollen tube elongation, localizes to Golgi, and might be involved in vesicle trafficking. Plant Physiol 2004; 135:1480–1490.
Kang BH, Rancour DM, Bednarek SY . The dynamin-like protein ADL1C is essential for plasma membrane maintenance during pollen maturation. Plant J 2003; 35:1–15.
Steinebrunner I, Wu J, Sun Y, Corbett A, Roux SJ . Disruption of apyrases inhibits pollen germination in Arabidopsis. Plant Physiol 2003; 131:1638–1647.
Lalanne E, Honys D, Johnson A, et al. SETH1 and SETH2, two components of the glycosylphosphatidylinositol anchor biosynthetic pathway, are required for pollen germination and tube growth in Arabidopsis. Plant Cell 2004; 16:229–240.
Xu J, Brearley CA, Lin WH, et al. A role of Arabidopsis inositol polyphosphate kinase, AtIPK2alpha, in pollen germination and root growth. Plant Physiol 2005; 137:94–103.
Qin GJ, Kang DM, Dong YY, et al. Obtaining and analysis of flanking sequences from T-DNA transformants of Arabidopsis. Plant Sci 2003; 165:941–949.
Liu YG, Huang N . Efficient amplification of insert end sequences from bacterial artificial chromosome clones by thermal asymmetric interlaced PCR. Plant Mol Biol Rep 1998; 16:175–181.
Qin G, Gu H, Zhao Y, et al. An indole-3-acetic acid carboxyl methyltransferase regulates Arabidopsis leaf development. Plant Cell 2005; 17:2693–2704.
Weigel D, Glazebrook J . Arabidopsis: a laboratory manual. Cold Spring Harbor, NewYork: Cold Spring Harbor Laboratory Press, 2002.
Regan SM, Moffatt BA . Cytochemical analysis of pollen development in wild-type Arabidopsis and a male-sterile mutant. Plant Cell 1990; 2:877–889.
Alexander MP . Differential staining of aborted and nonaborted pollen. Stain Technol 1969; 44:117–122.
Fan LM, Wang YF, Wang H, Wu WH . In vitro Arabidopsis pollen germination and characterization of the inward potassium currents in Arabidopsis pollen grain protoplasts. J Exp Bot 2001; 52:1603–1614.
Wu J, Mao X, Cai T, Luo J, Wei L . KOBAS server: a web-based platform for automated annotation and pathway identification. Nucl Acids Res 2006; 34:W720–724.
Guo L, Wang ZY, Lin H, et al. Expression and functional analysis of the rice plasma-membrane intrinsic protein gene family. Cell Res 2006; 16:277–286.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001; 25:402–408.
Seaman MN, Marcusson EG, Cereghino JL, Emr SD . Endosome to Golgi retrieval of the vacuolar protein sorting receptor, Vps10p, requires the function of the VPS29, VPS30, and VPS35 gene products. J Cell Biol 1997; 137:79–92.
Kametaka S, Okano T, Ohsumi M, Ohsumi Y . Apg14p and Apg6/Vps30p form a protein complex essential for autophagy in the yeast, Saccharomyces cerevisiae. J Biol Chem 1998; 273:22284–22291.
Sanders PM, Bui AQ, Weterings K, et al. Anther developmental defects in Arabidopsis thaliana male-sterile mutants. Sex Plant Reprod 1999; 11:297–322.
Preuss D, Rhee SY, Davis RW . Tetrad analysis possible in Arabidopsis with mutation of the QUARTET (QRT) genes. Science 1994; 264:1458–1460.
Copenhaver GP, Keith KC, Preuss D . Tetrad analysis in higher plants. A budding technology. Plant Physiol 2000; 124:7–16.
Johnson-Brousseau SA, McCormick S . A compendium of methods useful for characterizing Arabidopsis pollen mutants and gametophytically-expressed genes. Plant J 2004; 39:761–775.
Cole RA, Synek L, Zarsky V, Fowler JE . SEC8, a subunit of the putative Arabidopsis exocyst complex, facilitates pollen germination and competitive pollen tube growth. Plant Physiol 2005; 138:2005–2018.
Takacs-Vellai K, Vellai T, Puoti A, et al. Inactivation of the autophagy gene bec-1 triggers apoptotic cell death in C. elegans. Current Biol 2005; 15:1513–1517.
Stack JH, Dewald DB, Takegawa K, Emr SD . Vesicle-mediated protein-transport-regulatory interactions between the Vps15 protein-kinase and the Vps34 Ptdins 3-kinase essential for protein sorting to the vacuole in yeast. J Cell Biol 1995; 129:321–334.
Kihara A, Noda T, Ishihara N, Ohsumi Y . Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J Cell Biol 2001; 152:519–530.
Budovskaya YV, Hama H, DeWald DB, Herman PK . The C terminus of the Vps34p phosphoinositide 3-kinase is necessary and sufficient for the interaction with the Vps15p protein kinase. J Biol Chem 2002; 277:287–294.
Volinia S, Dhand R, Vanhaesebroeck B, et al. Human phosphatidylinositol 3-kinase complex related to the yeast Vps34P-Vps15P protein sorting system. EMBO J 1995; 14:3339–3348.
Kihara A, Kabeya Y, Ohsumi Y, Yoshimori T . Beclin-phosphatidylinositol 3-kinase complex functions at the trans-Golgi network. EMBO Rep 2001; 2:330–335.
Obara K, Sekito T, Ohsumi Y . Assortment of phosphatidylinositol 3-kinase complexes–Atg14p directs association of complex I to the pre-autophagosomal structure in Saccharomyces cerevisiae. Mol Biol Cell 2006; 17:1527–1539.
Franklin-Tong VE, Drobak BK, Allan AC, Watkins P, Trewavas AJ . Growth of pollen tubes of Papaver rhoeas is regulated by a slow-moving calcium wave propagated by inositol 1,4,5-trisphosphate. Plant Cell 1996; 8:1305–1321.
Kost B, Lemichez E, Spielhofer P, et al. Rac homologues and compartmentalized phosphatidylinositol 4, 5-bisphosphate act in a common pathway to regulate polar pollen tube growth. J Cell Biol 1999; 145:317–330.
Zheng ZL, Yang Z . The Rrop GTPase switch turns on polar growth in pollen. Trends Plant Sci 2000; 5:298–303.
Gu Y, Vernoud V, Fu Y, Yang Z . ROP GTPase regulation of pollen tube growth through the dynamics of tip-localized F-actin. J Exp Bot 2003; 54:93–101.
Hwang JU, Gu Y, Lee YJ, Yang Z . Oscillatory ROP GTPase activation leads the oscillatory polarized growth of pollen tubes. Mol Biol Cell 2005; 16:5385–5399.
Parton RM, Fischer-Parton S, Trewavas AJ, Watahiki MK . Pollen tubes exhibit regular periodic membrane trafficking events in the absence of apical extension. J Cell Sci 2003; 116:2707–2719.
Parton RM, Fischer-Parton S, Watahiki MK, Trewavas AJ . Dynamics of the apical vesicle accumulation and the rate of growth are related in individual pollen tubes. J Cell Sci 2001; 114:2685–2695.
Hepler PK, Vidali L, Cheung AY . Polarized cell growth in higher plants. Annu Rev Cell Dev Biol 2001; 17:159–187.
Sanderfoot AA, Pilgrim M, Adam L, Raikhel NV . Disruption of individual members of Arabidopsis syntaxin gene families indicates each has essential functions. Plant Cell 2001; 13:659–666.
Hicks GR, Rojo E, Hong S, Carter DG, Raikhel NV . Geminating pollen has tubular vacuoles, displays highly dynamic vacuole biogenesis, and requires VACUOLESS1 for proper function. Plant Physiol 2004; 134:1227–1239.
Acknowledgements
We thank Dr Matthew Terry (University of Southampton, UK) and Dr Yunde Zhao (UCSD, USA) for editing the manuscript, Dr Weicai Yang and Mr Li Yuan (CAS Institute of Genetics and Developmental Biology, China) for helpful suggestions, assistance in tissue sectioning and providing the qrt1 mutant. The work was supported by the National Natural Science Foundation of China (Grant No. 30470172 and 30221120261) and the National Special Projects for R&D of Transgenic Plants (J99-A-001).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Qin, G., Ma, Z., Zhang, L. et al. Arabidopsis AtBECLIN 1/AtAtg6/AtVps30 is essential for pollen germination and plant development. Cell Res 17, 249–263 (2007). https://doi.org/10.1038/cr.2007.7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2007.7
Keywords
This article is cited by
-
Gold nanoparticles (AuNPs) can rapidly deliver artificial microRNA (AmiRNA)-ATG6 to silence ATG6 expression in Arabidopsis
Plant Cell Reports (2023)
-
Significance of brassinosteroids and their derivatives in the development and protection of plants under abiotic stress
Biologia (2021)
-
Autophagy-mediated Nutrient Recycling and Regulation in Plants: A Molecular View
Journal of Plant Biology (2019)
-
Genome-wide analysis of autophagy-related genes (ARGs) in grapevine and plant tolerance to copper stress
Planta (2018)
-
Genome-wide analysis of autophagy-related genes in banana highlights MaATG8s in cell death and autophagy in immune response to Fusarium wilt
Plant Cell Reports (2017)