Abstract
MYB proteins play important roles in eukaryotic organisms. In plants, the R1R2R3-type MYB proteins function in cell cycle control. However, whether the R2R3-type MYB protein is also involved in the cell division process remains unknown. Here, we report that an R2R3-type transcription factor gene, AtMYB59, is involved in the regulation of cell cycle progression and root growth. The AtMYB59 protein is localized in the nuclei of onion epidermal cells and has transactivation activity. Expression of AtMYB59 in yeast cells suppresses cell proliferation, and the transformants have more nuclei and higher aneuploid DNA content with longer cells. Mutation in the conserved domain of AtMYB59 abolishes its effects on yeast cell growth. In synchronized Arabidopsis cell suspensions, the AtMYB59 gene is specifically expressed in the S phase during cell cycle progression. Expression and promoter-GUS analysis reveals that the AtMYB59 gene is abundantly expressed in roots. Transgenic plants overexpressing AtMYB59 have shorter roots compared with wild-type plants (Arabidopsis accession Col-0), and around half of the mitotic cells in root tips are at metaphase. Conversely, the null mutant myb59-1 has longer roots and fewer mitotic cells at metaphase than Col, suggesting that AtMYB59 may inhibit root growth by extending the metaphase of mitotic cells. AtMYB59 regulates many downstream genes, including the CYCB1;1 gene, probably through binding to MYB-responsive elements. These results support a role for AtMYB59 in cell cycle regulation and plant root growth.
Similar content being viewed by others
Introduction
MYB proteins are characterized by a DNA-binding MYB domain located in the N-terminus. The MYB domain usually contains one, two or three conserved motifs of around 50 amino acids. These repeated motifs have a helix-helix-turn-helix conformation and may facilitate interaction with the target DNA. The first MYB gene identified in plants was C1 from maize, which is involved in anthocyanin biosynthesis 1. Since then, several MYB genes have been isolated 2, 3, 4, 5. Recently, Chen et al. 6 found 198 MYB genes from Arabidopsis and 183 from rice. Plant MYB proteins can be classified into three types: the R2R3-type MYB, with two repeats; the R1R2R3-type MYB, with three repeats; and a third type, usually containing a single repeat or an atypical repeat. Among the MYB proteins from Arabidopsis, 126 are R2R3-type and 5 are R1R2R3-type. The remaining 67 are MYB-related or atypical types 6.
MYB proteins play important roles in multiple aspects of plant growth and development 2, 3, 4, 5. Several MYB proteins may be involved in secondary metabolism 7, 8, 9. AtMYB0/GL1 is required for leaf trichome differentiation 10. AtMYB66/WER plays roles in epidermal cell patterning 11. MYB genes also have functions in lateral meristem initiation 12. All the above functions are known from studying R2R3-type MYB proteins. Plant R1R2R3-type MYB proteins have also been studied, and they may play roles in regulating cell cycle progression, similar to the roles played by the three vertebrate MYB proteins with three repeats 2, 13, 14. The unusual MYB protein AtMYBCDC5 may also be involved in cell cycle control 15. However, whether the R2R3-type MYB proteins are involved in cell cycle regulation is not known.
Cell cycle progression is important for cell proliferation and for plant growth and development. In general terms, the cell cycle is divided into four phases, including G1, S (DNA replication), G2 and M (mitosis). G1 and G2 are two phases during which the cell determines whether it will go into the following S phase or M phase. The mechanism for cell cycle control is highly conserved among different organisms, and the major regulators of the plant cell cycle have been identified, including cyclin-dependent kinases (CDKs), cyclins, CDK-inhibitory proteins, the WEE kinase and components of the RB/E2F/DP pathway 16, 17, 18. In Arabidopsis, two classes of CDKs and 49 cyclins have been identified. A-type cyclins are important for the S-G2-M phase control, and B-type cyclins may regulate the G2 to M transition. D-type cyclins may control the G1 to S transition 16. CDKs and cyclins can form kinase complexes, which phosphorylate various substrates, leading to the progression of the cell cycle. All the components of the cell cycle progression may be regulated at various levels and by different developmental or environmental factors 17, 18. Plant hormones, including auxin, cytokinin, brassinosteroids, gibberellin and abscisic acid, also affect the cell cycle and cell division 18.
Previously, we analyzed the ethylene receptor-regulated genes using a microarray approach, and identified one MYB gene 19. In the present study, we investigated the function of this R2R3-type transcription factor gene, AtMYB59. We found that this gene affects cell growth by influencing DNA replication and cell division in yeast cells and regulates root development by altering the progression of mitosis in the root tip cells. AtMYB59 may function through regulation of its downstream target genes.
Results
AtMYB59 gene expression, protein localization and transcriptional activation
AtMYB59 (At5g59780) was identified during a microarray analysis for ethylene receptor-regulated genes 19 and was investigated further. It encodes an R2R3-type MYB protein of 236 amino acids. The two repeats are located at the N-terminus from amino acids 9 to 59 and 62 to 110. The AtMYB59 gene is highly expressed in roots but is barely detectable in stems, leaves, flowers and young siliques (Figure 1A). To further examine the specific expression pattern of AtMYB59 in plants, the 1.6 kb promoter region of the gene was used to drive expression of the GUS gene in a plant expression vector, pCAMBIA1301. The transgenic plants exhibited GUS activity in hypocotyls and root systems, including root hairs, vascular tissue and the root tip meristem region (Figure 1B). GUS staining was also observed in some points along the leaf edge and the pedicel supporting the silique (Figure 1B, upper middle and upper right). These results indicate that AtMYB59 is abundantly expressed in roots. It may also be detected in other organs at different developmental stages.
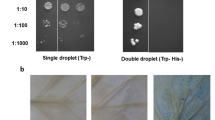
AtMYB59 gene expression, protein localization and transcriptional activation. (A) AtMYB59 expression in different organs of Arabidopsis plants. (B) GUS staining of transgenic plants harboring the GUS reporter gene driven by the 1.6 kb promoter of the AtMYB59 gene. (C) Subcellular localization of the AtMYB59-GFP protein in onion epidermal cells. The GFP plasmid was transformed as a control. The photographs were taken in the dark for green fluorescence (left), in bright light for morphology (middle) and in combination (right). (D) Transactivation activity of AtMYB59 in the yeast assay system. Transformants harboring pBD-AtMYB59, the positive control pGAL4 or the negative control pBD vector were streaked onto YPAD (upper left) or SD-HIS− (upper right) for growth examination. The LacZ marker gene was examined by X-gal assay (lower left). (E) Transactivation ability of AtMYB59 in Arabidopsis protoplasts. VP16 was used as a positive control. The GAL4 DNA-binding domain (GAL4DBD) was used as a negative control. The LUC gene was used as a reporter.
The subcellular localization of the AtMYB59 protein was studied. Figure 1C shows that the AtMYB59-GFP protein is localized in the nuclei of onion epidermal cells, whereas the GFP control protein is observed in the cytoplasm. This result indicates that AtMYB59 is a nuclear protein.
The transcriptional activation ability of the AtMYB59 protein was investigated using a yeast assay system. The AtMYB59 coding region was fused to the GAL4 DNA-binding domain to generate pBD-AtMYB59. This plasmid, together with the positive control pGAL4 and the negative control pBD vector, was transformed into the yeast strain YRG-2, and the growth status of the transformants was examined. Figure 1D shows that all the transformants grew well on the YPAD normal medium. However, only the positive control pGAL4 and the pBD-AtMYB59 transformants were able to grow on the selection medium SD-HIS− (Figure 1D, upper right panel). In the X-gal assay, the yeast cells harboring pGAL4 and pBD-AtMYB59 turned blue (Figure 1D, lower left panel). These results indicate that AtMYB59 can activate the expression of the HIS3 and LacZ marker genes in the yeast cell, implying that AtMYB59 has transcriptional activation activity.
The transcriptional activation ability of AtMYB59 was also examined in an Arabidopsis protoplast system (Figure 1E). The effector AtMYB59 protein had moderate transactivation activity in activating the reporter LUC gene, whereas the positive control effector VP16 had very high activity. The negative control GAL4BD protein had no transcriptional activation ability. These results indicate that AtMYB59 has transcriptional activation activity in plant cells.
AtMYB59 affects cell division of yeast cells
Yeast is a model system for studying gene function, and MYB-related proteins have been found in yeast cells 20. We investigated the function of AtMYB59 using the yeast system. The AtMYB59 gene in the pESPM-AtMYB59 plasmid was driven by the NMT promoter and can be activated in the absence of thiamine (−VB1) (Figure 2A). Under inductive conditions (−VB1) on a plate, the proliferation of the transformants harboring the AtMYB59 gene was severely inhibited in comparison with the transformants harboring the control vector (Figure 2B, 2C). Under normal non-inductive conditions (+VB1), both the transformants containing the AtMYB59 gene and those containing the vector grew well. The yeast transformants under inductive conditions were also stained with 4,6-diamidino-2-phenylindole (DAPI) and observed under a fluorescence microscope. The results in Figure 2D show that, whereas the yeast cells containing the vector had a normal phenotype, the yeast cells containing the AtMYB59 gene exhibited a significant change in morphology. The latter cells were ∼threefold longer than the vector-transformed cells (Figure 2D, 2E, right panel), and more than 20% of the AtMYB59-transformed cells had double nuclei or abnormal nuclei (Figure 2E, left panel). AtMYB59 was further mutated at four highly conserved amino acid positions (S48G, R50S, R52S, W53G), and the mutated AtMYB59 showed no significant effects on yeast cell growth compared with the pESPM vector control (Figure 2F). All these results indicate that the AtMYB59 gene specifically suppresses cell proliferation by affecting cell division and cell growth.
AtMYB59 affects cell division and morphology of the yeast transformants. (A) AtMYB59 expression in pESPM-AtMYB59 transformants (AtMYB59) with (+VB1) or without VB1 (−VB1). The transformants (pESPM) harboring the pESPM vectors were used as a control. (B) Dilution analysis of the yeast transformants harboring pESPM-AtMYB59 or the pESPM vector with (+VB1) or without VB1 (−VB1) on EMM plates. (C) Growth of the yeast transformants harboring pESPM-AtMYB59 or the pESPM vector in EMM liquid culture with (+VB1) or without VB1 (−VB1). Optical density at 595 nm was measured. (D) Phenotypic comparison of the yeast transformants containing pESPM-AtMYB59 or the pESPM vector in the absence of VB1. The cells were stained with DAPI. (E) Comparison of cell length and proportions of the cells with more nuclei. The cells from (D) were examined. (F) Effects of the AtMYB59 mutant gene on yeast transformants in the absence of VB1.
Because the AtMYB59-transformed cells showed defects in cell nuclei distribution, we further measured the DNA content in these cells using flow cytometry. Figure 3A (upper panel) shows that normal vector-transformed cells had two peaks in DNA content during the cell cycle. The first peak represents cells at the G2 phase, whereas the second peak indicates cells that have divided but have not yet separated from each other. Therefore, the cells in the second peak have twice the DNA content of a single cell but remain at the G2 phase. These features did not change during the culture. However, the AtMYB59-transformants showed significant alterations in DNA content during cell division (Figure 3A, lower panel). Several peaks at different DNA contents were observed in addition to the normal peak, suggesting the presence of cells with aneuploid DNA content (Figure 3A, lower panel). During progression of the culture, more than 70% of the AtMYB59-transformed cells showed variation in DNA content (Figure 3B, left panel). An increasing number of cells undergoing apoptosis were also observed in the AtMYB59-containing yeast cells but not in control cells (Figure 3A, lower panel; 3B, right panel). These results further indicate that the expression of AtMYB59 in yeast cells disturbs DNA synthesis and/or chromosome separation and hence affects the cell division process.
Flow cytometric analysis of the yeast transformants harboring pESPM-AtMYB59. (A) Comparison of the flow cytometric profiles of the transformants harboring pESPM-AtMYB59 or the pESPM vector. Red peaks indicate normal G2 cells. Yellow peaks indicate aneuploid cells. Blue peaks indicate cells in apoptosis. (B) Proportions of cells with aneuploid DNA content and in apoptosis. The percentage was derived from (A).
AtMYB59 is specifically expressed in the S phase of the cell cycle
Because AtMYB59 affects the cell division process, we examined its gene expression during cell cycle progression using a suspension culture. Several cell cycle marker genes were also examined. Figure 4 shows that CYCD3;1 gene expression starts during the S phase and extends to the G2/M phase; CDKA;1 is constantly expressed during the cell cycle; CDKB1;1 is abundantly expressed in the S to G2 phases; and the Histone H4 gene is highly expressed in the G1 to S phases. The expression patterns of these marker genes are consistent with those previously reported 21. Compared with the marker genes, AtMYB59 was strongly expressed in the S or S to G2 phases (Figure 4). In the G2 to M phases and the G1 phase, the AtMYB59 gene was only weakly expressed. The higher expression of the AtMYB59 gene in the S phase suggests that AtMYB59 may play roles in this phase or the following phase.
AtMYB59 expression during cell cycle progression of Arabidopsis suspension cultures. RT-PCR was performed to examine the expression of AtMYB59 and four other marker genes and the control gene Actin7. The same images as those in Zhang et al. 47 for the marker genes were used. The cell cycle phases (G1, S, G2 and M) are labeled on top of the panel.
AtMYB59 suppresses root growth and affects cell cycle progression
To study the AtMYB59 gene function in vivo, an expression vector harboring the 35S-driven AtMYB59 gene was transformed into Arabidopsis plants. Two homozygous lines with higher AtMYB59 expression, 3 and 11, were selected for further analysis (Figure 5A). Two T-DNA insertion mutants, myb59-1 and myb59-2, were also identified; they had insertion sites in the third exon and the 3′-untranslated region, respectively (Figure 5B). As shown in Figure 5B, myb59-1 is a knockout mutant, while myb59-2 is a knockdown mutant. Seedlings of the two overexpressing lines L3 and L11, myb59-1, myb59- 2, and Arabidopsis accession Col-0 (Col) were compared. Figure 5C and D show that the two AtMYB59-overexpressing lines had shorter roots and that myb59-1 had longer primary roots when compared with Col. However, myb59-2 did not show a significant difference in root growth. The aerial parts appeared not to have significant differences among these compared plants. The cross-sections of leaves from these plants were also examined, and no significant difference was found (Supplementary information, Figure S1B). These results indicate that AtMYB59 may inhibit root growth.
Overexpression of AtMYB59 affects root growth and cell division of root tip cells. (A) AtMYB59 expression in various AtMYB59 transgenic lines. (B) Identification of the AtMYB59 T-DNA insertion mutants myb59-1 and myb59-2. The left panel shows AtMYB59 expression in myb59-1 and myb59-2. The right panel shows the T-DNA insertion sites in the AtMYB59 genomic region. (C) Root growth of the AtMYB59-transgenic lines L3 and L11, Col, myb59-1 and myb59-2. (D) Comparison of root lengths of the two AtMYB59-transgenic lines, myb59-1, myb59-2, and Col plants. Seedlings (Three3-day-old) seedlings were transferred onto plates and grown vertically for 10 days. Bars indicate SD. (n = 20). Different letters above each column indicate significant differences (P < 0.01) between lines according to one-way ANOVA (SPSS 10.0). (E) Cell division phases in Arabidopsis root tip cells. Chromosome behavior was revealed by DAPI staining. Tubulin distribution was revealed by an FITC immunofluorescence assay. Five phases are shown. (F) Proportions of the mitotic cells at each phase in root tips of the AtMYB59 transgenic line L11, myb59-1, myb59-2, and Col plants. In total, around 100 mitotic cells were examined for each plant, and the mitotic phase of each cell was identified based on comparison with the phases in (E).
Short roots in the AtMYB59-overexpressing plants may result from altered cell size in the root. However, when cross-sections of the roots in the elongation zone were examined, no significant difference in root cell structure or size was observed among the AtMYB59-overexpressing plants, the mutants and Col (Supplementary information, Figure S1A). Confocal microscopic observation did not reveal significant differences in root cell length among these plants (data not shown).
Because the AtMYB59 gene is specifically expressed in the S phase of the cell cycle, affects yeast cell division and inhibits root growth (Figures 2, 3, 4, 5), we then examined whether the cell division phase was altered in root tips of the AtMYB59-overexpressing plants and of the mutants myb59-1 and myb59-2 in comparison with those of Col. Root tips were stained with DAPI and assayed for FITC immunofluorescence to reveal DNA and tubulin distribution, respectively. Four phases of the cells at mitosis can be found, i.e., prophase, metaphase, anaphase and telophase (Figure 5E). A cell at interphase is also shown for comparison (Figure 5E). The phases of the cells at mitosis were identified, and the percentage of cells at each phase was calculated. Figure 5F shows that, in Col root tips, the mitotic cells at each phase were in similar percentages (21%–27%). In the two T-DNA insertion mutants, myb59-1 had 14% of cells at metaphase and myb59-2 had 24% at metaphase. These percentages were lower than the 27% at metaphase in Col. However, in the AtMYB59-overexpressing line, a very high percentage of cells were at metaphase (54%) and a very low percentage of cells were at telophase (Figure 5F). These results indicate that AtMYB59 changes the behavior of cells at mitosis, possibly through disturbance of metaphase.
AtMYB59 regulates downstream genes and has DNA binding specificity
Because AtMYB59 affected root development and the cell division process, we then performed an Affymetrix genechip analysis using the AtMYB59-overexpressing line L11 and Col to identify the genes regulated by this transcription factor. Among the differentially expressed genes, CYCB1;1 and a NAM-like gene (At5g14000) were upregulated in the AtMYB59-overexpressing line L11 but were downregulated in the two myb59 mutants (Figure 6A). The CYCB1;1 gene mainly functions during the G2 to M transition of the cell cycle 22. NAM-like genes usually play roles in meristem formation and lateral root formation 19, 23. In addition, the AtMYB59 gene affected the expression of the cytokinin signal-transduction gene ARR16 (Figure 6A). These genes, together with another ∼150 genes identified from the genechip analysis (Supplementary information, Tables S1 and S2), may contribute to the function of the AtMYB59 gene in plant cell cycle progression and root growth. It should be noted that elimination of AtMYB59 expression in Atmyb59-1 only reduced – it did not abolish – the expression of the three genes, suggesting that AtMYB59 plays a partial role in regulation of these genes.
AtMYB59 regulates cell cycle-related genes and has DNA binding specificity. (A) Expression of the NAM-like gene (At5g14000), CYCB1;1 and ARR16 in the AtMYB59-transgenic line L11, myb59-1, myb59-2, and Col plants. RT-PCR was used for examination. (B) Potential MYB-binding elements in the promoter region of the AtMYB59 gene. MSA has the AACGG core sequence. MBSII: TAACTAAC. MRE: AACCaaa. MBS: CAACtg. Two ethylene-responsive elements (ERE: ATTTCAAA) are also indicated. (C) DNA binding specificity of the AtMYB59 in a yeast one-hybrid assay. The effector pAD-AtMYB59 and the reporter pHIS2, harboring different cis-elements (three tandem repeats) plus the HIS3 gene, were co-transformed into yeast cells. The growth of the transformants was examined on SD/-Trp-Leu-His or 3-AT plates. The cis-element in each transformant is labeled on the top or bottom of each panel. CK− indicates negative control cells harboring pAD-AtMYB59 and the pHIS2 vector without any cis-element. MSA: CCCAACGGT. MRE1: CCGGCAGTTAGGAT. MRE2: TATAACGGTTTTTT. MRE3: TCTAACCTACCA. MRE4: TCTCACCTACC. MYBR: TGGTTAG. Other element sequences are the same as those in (B).
Because the CYCB1;1 gene was regulated by the R2R3-type AtMYB59, we further examined whether the promoter region of CYCB1;1 has any MYB protein-binding elements and whether AtMYB59 can bind these elements. The 1.6 kb promoter region of the CYCB1;1 gene was analyzed using the PlantCare program and based on previous studies 24. Eight cis-elements, including three MSA sequences (AACGG), one MBSII sequence (TAACTAAC), two MRE sequences (AACC) and two MBS sequences (CAAC), were identified as potential MYB protein-binding elements (Figure 6B). Two ethylene-responsive elements (ERE) were also identified (Figure 6B). A yeast one-hybrid assay was used to test whether AtMYB59 can bind to five of these elements (MSA (−156), CCCAACGGT; MBSII (−356), TAACTAAC; ERE (−393 and −770), ATTTCAAA; MRE (−411), AACCAAA; MBS (−850 and −992), CAACTG). Other elements (MYBR, MRE1, MRE2, MRE3 and MRE4) that have previously been identified for other MYB proteins were also examined for AtMYB59 binding. Each of the pHIS2 reporter plasmids harboring different elements plus the reporter gene HIS3, together with the effector plasmid pAD-AtMYB59, was transformed into yeast cells. Growth of the transformants on 3-AT indicates activation of the HIS3 gene and binding of AtMYB59 to the corresponding cis-elements. Figure 6C shows that all the transformants grew well on SD/-Trp-Leu-His. However, only the transformants harboring the MRE, ERE and MRE2 (TATAACGGTTTTTT)-controlled reporter gene grew well on 3-AT, suggesting that AtMYB59 binds to these three elements but not to the other elements examined (Figure 6C, middle and lower panels). ERE represents an ethylene-responsive element, and the binding of AtMYB59 to this sequence may indicate that AtMYB59 has multiple regulatory roles by binding to different cis-elements in the CYCB1;1 gene.
Discussion
In vertebrate cells, the MYB proteins regulate cell cycle progression at the G1 to S transition or function by binding telomeres 14, 25, 26. In tobacco plants, the R1R2R3-type MYB proteins NtmybA1, NtmybA2 and NtmybB play roles in cell cycle progression by binding to cis-elements and regulating cyclin and NACK1 genes 2, 13. However, the first two proteins appear to be expressed at the G2 to M phase and function as activators in tobacco BY2 cells, whereas the latter protein (NtmybB) may be constantly expressed during the cell cycle and functions as a repressor 13. The present R2R3-type AtMYB59 also plays roles in cell cycle progression in both yeast cells and plant cells. In yeast cells, AtMYB59 may suppress cell division by affecting DNA synthesis in the S phase, since a large proportion of yeast cells with aneuploid DNA content were observed. Uneven separation of the replicated chromosomes may also result in the aneuploid DNA content in yeast cells. Some AtMYB59-overexpressing cells had two nuclei and appeared to be three times longer than the control cells. These transformed cells may have had normal chromosome separation and nuclei formation, but the two daughter cells could not separate from each other, leading to longer cells. This phenomenon further suggests that the cell division process is altered in AtMYB59-transformed yeast cells, probably due to the inability to form a cell plate between the two daughter cells or due to other unknown mechanisms. It is also possible that the change in cell division in AtMYB59-transformed yeast cells is due to the interference of AtMYB59 with yeast CDC5, which shows similarity to the c-MYB protein and is involved in cell cycle progression 27.
In plants, AtMYB59 appears to be a functional activator of transcription. It is specifically expressed during the phases S to G2 of the cell cycle in cell suspensions. This timing of expression is slightly later than those of the R1R2R3-type MYB genes in animal cells but earlier than that of the two tobacco genes NtmybA1 and NtmybA2, suggesting that AtMYB59 has specific roles in cell cycle progression. In root tips of AtMYB59-transgenic plants, a high proportion of mitotic cells were at metaphase, and a low proportion of mitotic cells were at metaphase in myb59-1. However, the cell size and other characteristics related to cell division appeared not to be changed (data not shown). It is possible that S/G2 phase expression of the AtMYB59 gene may shorten the S/G2 phase and then cause a significant increase in the proportion of metaphase cells and the length of metaphase. AtMYB59 may affect cell division through regulation of the CYCB1;1 gene by direct binding to the DNA elements in its promoter region (Figure 6).
Other MYB genes have been found to be expressed at different phases of the cell cycle. Three MYB genes have peak expressions at the S phase, whereas five other MYB genes are mainly expressed at the M phase 28, 29. Two other MYB genes are expressed at the G1 and G2 phases 29. Detailed characterization of these genes should determine their roles in cell cycle progression. AtCDC5, an Arabidopsis homolog of yeast cdc5, plays roles in the cell cycle that are similar to those of yeast cdc5 15, 30. The AtCDC5 MYB domain has two repeats that are only distantly related to those of the R2R3-type MYB domain 5, 15. Another unusual MYB protein (At2g03470) has also been identified during characterization of the promoter region of the Arabidopsis CYCB1;1 gene 31. This protein can function as a transcription activator and bind to the AACGG element of the CYCB1;1 gene, suggesting a role in cell cycle regulation 31. These studies suggest that multiple MYB proteins, including R1R2R3-type, R2R3-type and unusual types, may function during the cell cycle progression of plant cells.
Root growth and development are initiated from the root apical meristem and are regulated by transcription factors 32. Mutations of SHR and SCR result in shorter roots. SHR may specify endodermal cell fate, whereas SCR mediates the division of the cortex/endodermal initial daughter cell 33, 34. Another MYB coiled-coil transcription factor gene, APL, may be necessary for phloem cell formation 35. The AtMYB59 gene studied herein is abundantly expressed in the vascular tissue of roots, consistent with the expression pattern reported by Li et al. 36. Transgenic Arabidopsis overexpressing AtMYB59 exhibited shorter roots, and myb59-1 had longer roots than Col, indicating that AtMYB59 may inhibit root growth and function as a negative regulator. Because the dividing root tip cells of the AtMYB59-overexpressing plants had a high proportion at metaphase and the proportion of cells at each phase may reflect the relative duration of each phase 21, the dividing cells in root tips of AtMYB59-transgenic plants may have an extended metaphase. This relative extension of the metaphase of root tip cells during mitotic cell division likely resulted in the slower root growth of the AtMYB59-transgenic plants. The disturbance in cell division may be caused by an alteration in the cyclin gene CYCB1;1 and in many other genes, since AtMYB59 regulates the expression of these genes (Figure 6A; Supplementary information, Table S1 and S2). Changes in gene expression and/or activity of the cell cycle components have been found to affect root growth and differentiation 22, 37. The inhibitory effect of AtMYB59 on root growth is consistent with a recent report by Petroni et al. 38, who found that another MYB protein (AtMYB11) delays root growth and other developmental processes by reducing the proliferation activity of meristematic cells 38.
Abundant expression of the AtMYB59 gene in roots correlates with its roles in root growth regulation. AtMYB59 is also slightly expressed in aerial plant organs. However, no significant phenotypic change was observed in the aerial parts of the overexpressing lines or the mutant lines. The reason for this remains unknown. In the two mutants, the residual AtMYB59 expression differs, whereas the examined downstream genes seem to be expressed at similar levels. This phenomenon may be due to the fact that AtMYB59 only partially regulates the expression of these genes. However, phenotypic changes in myb59-1 are well correlated with loss of AtMYB59 expression, suggesting that other genes regulated by AtMYB59 are also involved in the process in addition to the tested genes.
Overall, we have demonstrated a role for the R2R3-type MYB protein AtMYB59 in cell division and root growth. Further study should reveal more about the function of AtMYB59 in plants.
Materials and Methods
Plant materials and treatments
Seeds of wild-type Arabidopsis (Arabidopsis thaliana, ecotype Columbia, Col-0), and transgenic plants or mutants were sowed on MS plate after sterilization, stratified at 4 °C for 3 days and germinated at 22 °C under continuous illumination. Seedlings (14-day-old) of these plants were harvested for RNA isolation. Wild-type plants (2-month-old) grown in soil were collected for organ-specific expression analysis.
For root growth comparison, 3-day-old seedlings were transferred to plates for vertical growth. The lengths of the primary roots from 20 plants were measured. The experiment was repeated independently three times, and the results were identical. The result from one experiment is presented.
RNA isolation, northern hybridization and RT-PCR analysis
Total RNA isolation and hybridization were carried out according to the description by Zhang et al. 39. AtMYB59 cDNA probe was synthesized using templates amplified with primers 5′-CCG GTT TAA GGA CAA ACC GGG-3′ and 5′-TCA GAA ATT CCA AAC GCA ATC-3′. After stripping the probes, the same blots were re-hybridized with 18S rDNA gene to examine the RNA loading.
For AtMYB59-regulated gene expression analysis, two independent sets of total RNAs were used for cDNA synthesis and then RT-PCR analysis. The results were consistent, and only one set of results was presented. Specific primers for each gene were as follows: for AtMYB59, 5′-AAC ATG GGA AAG TTC CTT GG-3′ and 5′-CCA TGT TCA AAA CTT AGA GG-3′; for NAM-like (At5g14000), 5′-GAT TTG CGT CGG CTA AGG TGT CT-3′ and 5′-ATC TCA ATT CTT CAT CAT CTC CC-3′; for CYCB1;1, 5′-ACT GTG CAA AGC TGT TGG CGT AT-3′ and 5′-TTC GTC CTT GGA GTA TTT CTT TCG T- 3′; for ARR16, 5′-TGG AGT ATT TGG GTT TGG GAG AT-3′ and 5′-CCA GGC ATA CAG TAA TCG GTG AT-3′; for actin7 gene, 5′-GGA CCT GAC TCA TCG TAC TC-3′ and 5′-TAC AGT GTC TGG ATC GGA GG-3′.
Construction of AtMYB59 expression vector and transformation
The coding sequence of AtMYB59 gene was obtained from Arabidopsis by RT-PCR using primers 5′-ATA GAA TTC ATG GAT TAC GAG GCA TCA AG-3′ and 5′-ACT GTC GAC TCA GAA ATT CCA AAC GCA ATC-3′. The PCR product was digested with BamHI and SacI, cloned into the plant expression vector pBI121 and confirmed by sequencing. AtMYB59 was driven by the CaMV 35S promoter. Plant transformation was performed using the vacuum infiltration method. The transgenic seeds were screened on MS plates containing 50 μg/ml kanamycin and the homozygous T3 seeds were analyzed further.
For promoter-GUS analysis, 1.6 kb promoter region of the AtMYB59 gene was amplified using specific primers 5′-GCA TGC GAC CAA ACC CAA CTA ATC AC-3′ and 5′-GGA TCC AGA GAG AGA GAG AGA GAG AG-3′, sequenced and cloned into the BamHI and XhoI sites of the plant expression vector pCambia1301 to drive the GUS reporter gene expression. This construct was transformed into Arabidopsis, and the transgenic plants were selected on hygromycin. GUS staining was performed to examine GUS activity.
Identification of T-DNA insertion mutants for the AtMYB59 gene
Two T-DNA insertion lines (N460129 for myb59-1 from NASC, European Arabidopsis Stock Center, and Salk_044751 for myb59-2 from ABRC at Ohio State University) for the AtMYB59 gene were requested and the homozygous plants were identified by PCR. The primers used for screening homozygous mutants myb59-1 were F: 5′-GAA ATG ACC TGG TTT TGG TTG TAG-3′ and R: 5′-ATG TTG GAG AAG CCA GAG GAG G-3′ for the gene, and pCA161F: 5′-CCG GAC ATG AAG CCA TTT ACA AT-3′ from the T-DNA region. The primers used for the myb59-2 mutant were LP: 5′-TGG GAA CTG ATC TTT TAT CAT GGA-3′ and RP: 5′-GGC TCA AGA GAA GAA GCG ACC-3′ from the gene, and the primers LBb1: 5′-GCG TGG ACC GCT TGC TGC AAC T-3′ from the T-DNA region. These two mutants were further identified by RT-PCR.
Subcellular localization of AtMYB59-GFP in onion epidermal cells
The AtMYB59 coding region was amplified with primers 5′-ATA GGA TCC ATG GAT TAC GAG GCA TCA AG-3′ and 5′-ATC GTC GAC GAA ATT CCA AAC GCA ATC C-3′, sequenced and fused to the 5′- end of GFP in transient expression vector, resulting in the pUC-AtMYB59-GFP construct. The AtMYB59-GFP was driven by the CaMV 35S promoter. The pUC-GFP vector, controlled by the 35S promoter, was used as a control. The two constructs were introduced into onion epidermal cells with a Biolistic Particle Delivery System (BioRad). The fluorescence of GFP in onion epidermal cells was visualized under a Confocal microscope (Olympus FV500).
Transcriptional activation activity of the AtMYB59 protein
The coding sequence of AtMYB59 was obtained by PCR using the primers 5′-ATA GAA TTC ATG GAT TAC GAG GCA TCA AG-3′ and 5′-ACT GTC GAC TCA GAA ATT CCA AAC GCA ATC-3′. The PCR products were cloned into the vector containing the GAL4 DNA-binding domain to obtain pBD-AtMYB59. The pBD-AtMYB59, the positive control pGAL4, and the negative control pBD vector were all transformed into the yeast strain YRG2. The transformed strains were confirmed by PCR, and then streaked on YPAD or SD/HIS− plates. The transcription activation activity of the protein was evaluated according to their growth status 40.
The transcriptional activation ability was also examined in Arabidopsis protoplast system according to our previous report 41. A plasmid containing 5X UAS and 35S promoter upstream of a reporter gene encoding a firefly luciferase (LUC) was used as a reporter. The GAL4 DNA-binding domain (BD)-coding sequence was fused to the transcription factor gene AtMYB59 and inserted into the pRT107 to generate effector plasmids pRT-BD-AtMYB59. The fusion genes were under the control of 35S promoter. The BD sequence was also fused to VP16 gene to generate positive control effector plasmid. The pRT107 containing the BD sequence was used as negative control. The reporter plasmids and the effector plasmids were transfected through PEG-mediated transfection. A pPTRL plasmid that contained a CaMV 35S promoter and a luciferase gene from Renilla was used as an internal control 42. Luciferase assays were performed with the Promega Dual-luciferase reporter assay system and the GloMaxTM20/20 Luminometer.
DNA binding specificity of the AtMYB59 in yeast one-hybrid assay
Three copies of each potential cis-acting DNA element in tandem repeat, with cohesive ends, were synthesized (Sengon of Shanghai, China), annealed and ligated into the SacI and MluI sites of the reporter plasmid pHIS2. The putative cis-elements in the promoter region of the Arabidopsis CYCB1;1 gene were predicted using the PlantCARE program. The nucleotide sequences of the two strands of these potential elements or other known MYB-binding elements were as follows: for ERE, 5′-CATTTc aaaATTTcaaaATTTcaaaA-3′ and 5′-CGCGTtttgAAATtttgAAATtttgAAATGAGCT-3′; for MBS, 5′-CCAACtgCAACtgCAACtgA-3′ and 5′-CGCGTcaGTTGcaGTTGcaGTTGGAGCT-3′; for MRE, 5′-CAACCaaaAACCaaaAACCaaaA-3′ and 5′-CGCGTtttGGTTtttGGTTtttGGTTGAGCT-3′; for MSA, 5′-CcccAACGgtcccAACGgtcccAACGgtA-3′ and 5′-CGCGTacCGTTgggacCGTTgggacCGTTgggGAGCT-3′; for MBS II 24, 5′-CTAACTAACTAACTAACTAACTAACA-3′ and 5′-CGCGTGTTAGTTAGTTAGTTAGTTAGTTAGAGCT-3′; for MYBR 43, 5′-CTGGTTAGTGGTTAGTGGTTAGA-3′ and 5′-CGCGTCTAACCACTAACCACTAACCAGAGCT-3′; for MRE1, 5′-CCCGGCAGTTAGGATCCGGCAGTTAGGATCCGGCAGTTAGGATA-3′ and 5′-CGCGTATCCTAACTGCCGGATCCTAACTGCCGGATCCTAACTGCCGGGAGCT-3′; for MRE2 44, 5′-CTATAACGGTTTTTTTATAACGGTTTTTTTATAACGGTTTTTTA-3′ and 5′-CGCGTAAAAAACCGTTATAAAAAAACCGTTATAAAAAAACCGTTATAGAGCT-3′; for MRE3 45, 5′-CTCTAACCTACCATCTAACCTACCA TCTAACCTACCAA-3′ and 5′-CGCGTTGGTAGGTTAGATGGTAGGTTAGA TGGTAGGTTAGAGAGCT-3′; for MRE4 46, 5′-CTCTCACCTACCTCTCACCTACC TCTCACCTACCA-3′ and 5′-CGCGTGGTAGGTGAGAGGTAGGTGAGA GGTAGGTGAGAGAGCT-3′; the underlined sequences represent the cohesive ends after annealing. These DNA elements were located upstream the nutritional reporter gene HIS3. The sequence of the inserts was confirmed by sequencing. The AtMYB59 gene was cloned into the plasmid pAD-GAL4–2.1 and the effector plasmid pAD-AtMYB59 was made. The effector and the reporter plasmids were transformed into yeast cells and the transformants were examined for their growth on SD/-Trp-Leu-His plate or this plate plus 3-AT based on the instruction of the BD Matchmaker™ Library Construction & Screening Kits. The pAD-AtMYB59 construct and the pHIS2 vector were co-transformed into yeast cells (Y187) as a negative control.
Affymetrix Genechip analysis
Two biological replicates were used for the analysis. Total RNAs from 12-day-old seedlings of the AtMYB59-transgenic line L11 and the wild-type Col plants were used for cDNA probe synthesis and Affymetrix Genechip hybridization analysis. Genechips (Affymetrix ATH1-121501) were used and the experiments were performed by the Shanghai Biochip Co Ltd. The genes with at least twofold difference in expression between the AtMYB59-overexpressing plants and the Col are listed in Supplementary information, Table S1 for upregulated genes and in Supplementary information, Table S2 for downregulated genes. Several genes were further examined by RT-PCR method.
Yeast transformation, growth and observation
The coding region of the AtMYB59 was inserted in expression vector pESPM and the pESPM-AtMYB59 was constructed. This recombinant plasmid and the pESPM vector were transformed into the fission yeast Leu strain SPQ-01 by lithium acetate-mediated method. Cells were selected on EMM agar plates with thiamine at 30 °C. The transformant was cultured in liquid EMM medium to mid-exponential phase with thiamine at 30 °C. The cells were washed three times with the EMM only to remove the thiamine and to de-repress the NMT1 promoter that drives the AtMYB59 gene, and then cultured at 30 °C for 22 h. The growth of the yeast cells was examined by measuring the optical density of the suspensions or observed under a light microscope. The transformed mid-exponential yeast cells were streaked onto the plates with or without thiamine, with different dilutions, and incubated at 30 °C. The grown of the yeast cells was observed after 60 h. The yeast cells harboring the pESPM vector were used as a control.
Expression of the AtMYB59 gene in the transformed yeast cells was examined by RT-PCR using primers 5′-CCG CTC GAG ATG AAA CTT GTG CAA GAA G-3′ and 5′-CGG GAT CCC TAA AGG CGA CCA CTA CCA TG-3′. Yeast actin gene was examined using primers 5′-CAG CCA CTT TCT CAA ATC AG-3′ and 5′-GCC AAG GGT CAC TAC AC-3′.
AtMYB59 mutation was generated in the peptide RTGKSCRLRWVNYL at positions S48G, R50S, R52S and W53G. Two primers, SalI-MYB59: 5′-GTC GAC ATG AAA CTT GTG CAA GA-3′ and mutmyb59-1: 5′-AAC CCC ACT TAA ACT GCA ACC CTT-3′, were used to amplify the 5′-part of the mutant gene. Two primers, BamHI-MYB59: 5′-GGA TCC CTA AAG GCG ACC ACT A-3′ and mutmyb59-2: 5′-AAG GGT TGC AGT TTA AGT GGG GT-3′, were used to amplify the 3′-part of the mutant gene. The two fragments harboring the mutation sites were used for PCR amplification with the SalI-MYB59 and BamHI-MYB59 primers to obtain the full-length of the mutant gene. The mutant gene was further cloned into pESPM vector to test the mutation effects in yeast transformants.
The yeast cells were stained with DAPI (1 mg/ml) to reveal the DNA and examined under an Olympus fluorescence microscope.
Flow cytometry analysis
The yeast cells harboring the pESPM-AtMYB59 or the pESPM vector were grown at 30 °C for 12 h in EMM medium containing thiamine. The cultures were washed six times with the EMM medium without thiamine to de-repress the NMT1 promoter, and diluted to less than 5 × 106 cells/ml in EMM, and then incubated at 30 °C. Samples were taken at 0, 3, 6 and 9 h and analyzed with a FACS Caliber cytometer (B-D Corporation, USA).
Staining and observation of cell division phase in Arabidopsis root tip cells
Whole seedlings grown on MS plates for 4 days were fixed in PEM buffer (50 mM Pipes, 5 mM MgCl2, 5 mM EGTA, pH 6.9) containing 4% paraformaldehyde for 1 h. After washing with PEM buffer, the samples were digested with 1% cellulase plus 0.1% pectase for 5 min. The root tips were cut and cells were spread out by pressing cover glasses gently. The materials were extracted with PEM buffer containing 0.1% Triton X-100 for 30 min. After washing, the materials were further incubated with PBSA containing 0.1% BSA for 10 min, and then incubated with mouse anti-tubulin antibody (Sigma T4026) (1:1 000 dilution) at 4 °C overnight. The materials were incubated with horse anti-mouse antibody (1:500 dilution) labeled by FITC at 37 °C for 2 h, and then stained with DAPI. The cells were observed under an Olympus fluorescence microscope, and the cell division phase was identified and counted.
Synchronization of cell suspensions and RT-PCR analysis of marker genes
Arabidopsis cell suspension was synchronized following the description by Zhang et al. 47. RT-PCR analysis and primers of the cell cycle marker genes were based on the descriptions, and the experiments were performed at the same time and the same images were used 47. For AtMYB59, the 5′- and 3′-coding sequences were used as primers for amplification.
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Paz-Ares J, Ghosal D, Wienand U, Peterson PA, Saedler H . The regulatory C1 locus of Zea mays encodes a protein with homology to myb proto-oncogene products and with structural similarities to transcriptional activators. EMBO J 1987; 6:3553–3558.
Ito M . Conservation and diversification of three-repeat Myb transcription factors in plants. J Plant Res 2005; 118:61–69.
Jin H, Martin C . Multifunctionality and diversity within the plant Myb-gene family. Plant Mol Biol 1999; 41:577–585.
Kranz HD, Denekamp M, Greco R, et al. Towards functional characterization of the members of the R2R3-MYB gene family from Arabidopsis thaliana. Plant J 1998 ; 16:263–276.
Stracke R, Werber M, Weisshaar B . The R2R3-Myb gene family in Arabidopsis thaliana. Curr Opin Plant Biol 2001; 4:447–456.
Chen Y, Yang X, He K, et al. The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 2006; 60:107–124.
Borevitz JO, Xia YJ, Blount J, Dixon RA, Lamb C . Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 2000; 12:2383–2393.
Goicoechea M, Lacombe E, Legay S, et al. EgMYB2, a new transcriptional activator from Encalyptus xylem, regulates secondary cell wall formation and lignin biosynthesis. Plant J 2005; 43:553–567.
Nesi N, Jond C, Debeaujon I, Caboche M, Lepiniec L . The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 2001; 13:2099–2114.
Oppenheimer DG, Herman PL, Sivakumaran S, Esch J, Marks MD . A myb gene required for leaf trichome differentiation in Arabidopsis is expressed in stipules. Cell 1991; 67:483–493.
Ryu KH, Kang YH, Park Y, Hwang I, Schiefelbein J, Lee MM . The WEREWOLF MYB protein directly regulates CAPRICE transcription during cell fate specification in the Arabidopsis root epidermis. Development 2005; 132:4765–4775.
Muller D, Schmitz G, Theres K . Blind homologous R2R3 Myb genes control the pattern of lateral meristem initiation in Arabidopsis. Plant Cell 2006; 18:586–597.
Ito M, Araki S, Matsunaga S, et al. G2/M-phase-specific transcription during the plant cell cycle is mediated by c-MYB-like transcription factors. Plant Cell 2001; 13:1891–1905.
Joaquin M, Watson RJ . Cell cycle regulation by the B-Myb transcription factor. Cell Mol Life Sci 2003; 60:2389–2401.
Hirayama T, Shinozaki K . A cdc5+ homolog of a higher plant, Arabidopsis thaliana. Proc Natl Acad Sci USA 1996; 93:13371–13376.
Inze D . Green light for the cell cycle. EMBO J 2005; 24:657–662.
Jakoby M, Schnittger A . Cell cycle and differentiation. Curr Opin Plant Biol 2004; 7:661–669.
Stals H, Inze D . When plant cells decide to divide. Trends Plant Sci 2001; 6:359–364.
He XJ, Mu RL, Cao WH, Zhang ZG, Zhang JS, Chen SY . AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant J 2005; 44:903–916.
Ju QD, Morrow BE, Warner JR . REB1, a yeast DNA-binding protein with many targets, is essential for growth and bears some resemblance to the oncogene myb. Mol Cell Biol 1990; 10:5226–5234.
Menges M, Samland AK, Planchais S, Murray JAH . The D-type cyclin CYCD3;1 is limiting for the G1-to-S-phase transition in Arabidopsis. Plant Cell 2006; 18:893–906.
De Jager SM, Maughan S, Dewitte W, Scofield S, Murray JAH . The developmental context of cell-cycle control in plants. Semin Cell Dev Biol 2005; 16:385–396.
Olsen AN, Ernst HA, Leggio LL, Skriver K . NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 2005; 10:79–87.
Romero I, Fuertes A, Benito BJ, Malpica JM, Leyva A, Paz-Ares J . More than 80 R2R3-MYB regulatory genes in the genome of Arabidopsis thaliana. Plant J 1998; 14:273–284.
Marhamati DJ, Bellas RE, Arsura M, Kypreos KE, Sonenshein GE . A-myb is expressed in bovine vascular smooth muscle cells during the late G1-to-S phase transition and cooperates with c-myc to mediate progression to S phase. Mol Cell Biol 1997; 17:2448–2457.
Broccoli D, Smogorzewska A, Chong L, de lange T . Human telomeres contain two distinct Myb-related proteins, TRF1 and TRF2. Nat Genet 1997; 17:231–235.
Ohi R, McCollum D, Hirani B, et al. The Schizosaccharomyces pombe cdc5+ gene encodes an essential protein with homology to c-Myb. EMBO J 1994; 13:471–483.
Menges M, Hennig L, Gruissem W, Murray JA . Cell-cycle-regulated gene expression in Arabidopsis. J Biol Chem 2002; 277:41987–42002.
Menges M, Hennig L, Gruissem W, Murray JAH . Genome-wide gene expression in an Arabidopsis cell suspension. Plant Mol Biol 2003; 53:423–442.
Lin Z, Yin K, Zhu D, Chen Z, Gu H, Qu LJ . AtCDC5 regulates the G2 to M transition of the cell cycle and is critical for the function of Arabidopsis shoot apical meristem. Cell Res 2007; 17:815–828.
Planchais S, Perennes C, Glab N, Mironov V, Inze D, Bergounioux C . Characterization of cis-acting element involved in cell cycle phase-independent activation of Arath;CycB1;1 transcription and identification of putative regulatory proteins. Plant Mol Biol 2002; 50:111–127.
Lee JY, Colinas J, Wang JY, Mace D, Ohler U, Benfey PN . Transcriptional and posttranscriptional regulation of transcription factor expression in Arabidopsis roots. Proc Natl Acad Sci USA 2006; 103:6055–6060.
Di Laurenzio L, Wysocka-Diller J, Malamy JE, et al. The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 1996; 86:423–433.
Helariutta Y, Fukaki H, Wysocka-Diller J, et al. The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 2000; 101:555–567.
Bonke M, Thitamadee S, Mahonen AP, Hauser MT, Helariutta Y . APL regulates vascular tissue identity in Arabidopsis. Nature 2003; 426:181–186.
Li J, Yang X, Wang Y, et al. Two groups of MYB transcription factors share a motif which enhances trans-activation activity. Biochem Biophys Res Commun 2006; 341:1155–1163.
Doerner P, Jorgensen JE, You R, Steppuhn J, Lamb C . Control of root growth and development by cyclin expression. Nature 1996; 380:520–523.
Petroni K, Falasca G, Calvenzani V, et al. The AtMYB11 gene from Arabidopsis is expressed in meristematic cells and modulates growth in planta and organogenesis in vitro. J Exp Bot 2008; 59:1201–1213.
Zhang JS, Zhou JM, Zhang C, Chen SY . Differential gene expression in a salt-tolerance rice mutant and its parental variety. Sci China 1996; 39:310–319.
Wang YJ, Zhang ZG, He XJ, et al. A rice transcription factor OsbHLH1 is involved in cold stress response. Theor Appl Genet 2003; 107:1402–1409.
Liao Y, Zou HF, Wei W, et al. Soybean GmbZIP44, GmbZIP62 and GmbZIP78 genes function as negative regulator of ABA signaling and confer salt and freezing tolerance in transgenic Arabidopsis. Planta 2008; 228:225–240.
Ohta M, Ohme-Takagi M, Shinshi H . Three ethylene-responsive transcription factors in tobacco with distinct transactivation functions. Plant J 2000; 22:29–38.
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K . Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 2003; 15:63–78.
Martin C, Paz-Ares J . MYB transcription factors in plants. Trends Genet 1997; 13:67–73.
Hartmann U, Valentine WJ, Christie JM, Hays J, Jenkins GI, Weisshaar B . Identification of UV/blue light-response elements in the Arabidopsis thaliana chalcone synthase promoter using a homlogous protoplast transient expression system. Plant Mol Biol 1998; 36:741–754.
Sugimoto K, Takeda S, Hirochika H . MYB-related transcription factor NtMYB2 induced by wounding and elicitors is a regulator of the tobacco retrotransposon Tto1 and defense-related genes. Plant Cell 2000; 12:2511–2528.
Zhang WK, Shen YG, He XJ, et al. Characterization of a novel cell cycle-related gene from Arabidopsis. J Exp Bot 2005; 56:807–816.
Acknowledgements
We are grateful to The Arabidopsis Biological Resource Center (ABRC) at Ohio State University and the European Arabidopsis Stock Center (NASC) for providing seeds of Arabidopsis AtMYB59 mutants. This work was supported by the National Key Basic Research Projects (2006CB100102) and the National Hightech Project of China (2006AA10Z113, 2007AA021402).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information, Figure S1
Cross-sections of roots and leaves from AtMYB59-overexpressing line L11, mutant myb59-1 and Col-0 plants. (PDF 67 kb)
Supplementary information, Table S1
Genes upregulated in AtMYB59-overexpressing Arabidopsis plant (PDF 14 kb)
Supplementary information, Table S2
Genes downregulated in AtMYB59-overexpressing Arabidopsis plant. (PDF 12 kb)
Rights and permissions
About this article
Cite this article
Mu, RL., Cao, YR., Liu, YF. et al. An R2R3-type transcription factor gene AtMYB59 regulates root growth and cell cycle progression in Arabidopsis. Cell Res 19, 1291–1304 (2009). https://doi.org/10.1038/cr.2009.83
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.83
Keywords
This article is cited by
-
Involvement of MYB family transcription factors, WsMYBL1 and WsMYBL2, in withanolide biosynthesis in Withania somnifera, a medicinal plant
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
Biological Function and Stress Response Mechanism of MYB Transcription Factor Family Genes
Journal of Plant Growth Regulation (2023)
-
Two Medicago truncatula growth-promoting rhizobacteria capable of limiting in vitro growth of the Fusarium soil-borne pathogens modulate defense genes expression
Planta (2023)
-
MIR159 regulates multiple aspects of stamen and carpel development and requires dissection and delimitation of differential downstream regulatory network for manipulating fertility traits
Physiology and Molecular Biology of Plants (2023)
-
Overexpression of AtMYB115 gene in transgenic rice exhibits increased early regeneration and results in dwarf phenotype
Journal of Plant Biochemistry and Biotechnology (2023)