Abstract
Some plant and animal pathogens can manipulate their hosts to cause them to release odors that are attractive to the pathogens' arthropod vectors. However, the molecular mechanism underlying this process is largely unexplored, and the specific effectors the pathogens employ as well as the pathways within the hosts they target are currently unknown. Here we reveal that the aphid-borne cucumber mosaic virus (CMV) employs its 2b protein, a well-characterized viral suppressor of host RNA interference (VSR), to target the host's jasmonate (JA) hormone pathway, thus acting as a viral inducer of host attractiveness to insect vectors (VIA). 2b inhibits JA signaling by directly interacting with and repressing JA-induced degradation of host jasmonate ZIM-domain proteins, instead of using its VSR activity. Our findings identify a previously defined VSR protein as a VIA and uncover a molecular mechanism CMV uses to manipulate host's attractiveness to insect vectors by targeting host hormone signaling.
Similar content being viewed by others
Introduction
Many important diseases in plants and animals are caused by pathogens that are naturally transmitted by arthropod vectors such as mosquitoes and aphids. The emergence and success of these pathogens are shaped by their molecular interactions with both the host and the vector1,2,3. Intriguingly, infection with diverse arthropod-borne pathogens in plants and mammals induces changes in the host that make the host more attractive to the disease vector4,5,6. Recent studies have shown that pathogen-induced vector attraction can be odor-dependent7,8,9,10, suggesting, presumably, the presence of a specific mechanism by which pathogens manipulate the host's ability to emit odors that could attract disease vector.
Cucumber mosaic virus (CMV) is probably one of the most successful plant pathogens as it can infect more than 1 200 plant species including the model plant Arabidopsis thaliana11. CMV is transmitted by aphids in a non-persistent manner11, and infection of CMV can induce odor-dependent attractiveness to aphids in squash plants9. It is well known that the primary defense against CMV and other viruses in A. thaliana is directed by antiviral RNA interference (RNAi) guided by the virus-derived small interfering RNAs (siRNAs)12. As a common counter-defense strategy of plant and animal viruses, CMV encodes a viral suppressor of host RNAi (VSR), the 2b protein, which is essential for infection12. The 2b protein exerts its VSR activity predominantly by inhibiting the production of viral siRNAs as a double-stranded RNA (dsRNA)-binding protein, though it also inhibits the RNA-induced silencing complex activity by direct interaction with Argonaute proteins13,14,15,16.
Many defense responses to insect pests in A. thaliana are regulated by the phytohormone jasmonate (JA)17,18,19. Upon JA perception, the F-box protein COI1 recruits jasmonate ZIM-domain proteins (JAZs), the key repressors of JA signaling, for ubiquitination and degradation by the 26S proteasome pathway20,21,22,23,24,25,26. The JAZ family of proteins is widely distributed among higher plants and includes 12 members in A. thaliana23,24,25. JAZs normally bind and repress transcription factors (such as MYC2, MYC3, MYC4, MYB21 and MYB24), which are released upon JA-induced degradation of JAZs to activate JA-responsive genes essential for host defense and development27,28,29. Some JA responsive genes mediate production of various plant secondary metabolites including plant volatiles to modulate plant-insect interactions18,30,31,32. For example, MYC2 can directly activate the expression of both TPS10, which promotes the biosynthesis of plant terpenes that reduce plant attractiveness to whiteflies33, and CYP81D11, which encodes a putative p450 cytochrome whose overexpression in transgenic plants decreases the attractiveness to aphids34.
In addition to suppressing host's antiviral immunity for a successful infection, arthropod-borne viruses also induce host attractiveness to insect vectors for efficient viral transmission. However, both the viral effector that mediates the induction of host attractiveness and its target have not been defined. In this study, we show that protein 2b from aphid-borne virus CMV serves as a viral inducer of host attractiveness to insect vectors (VIA), and enhances Arabidopsis's attractiveness to CMV's aphid vectors. Mechanistically, 2b suppresses Arabidopsis's JA signaling pathway by directly interacting with JAZ proteins and preventing the degradation of these key repressors of JA signaling by 26S proteasome.
Results
VSR 2b controls the viral induction of host attractiveness to aphid vectors
We performed two feeding preference assays to determine if the CMV infection induces odor-dependent attraction to aphid vectors in Arabidopsis. In the circular-dish bioassay, leaves from wild-type Arabidopsis (Col-0) either systemically infected with a subgroup I strain of CMV (SD-CMV) or mock inoculated were arranged next to each other in a circle. Apterous aphids (Myzus persicae) were set free from the center of the dish and their crawling traces were recorded with a digital camera (Supplementary information, Figure S1A and S1B). We found that significantly higher percentages of aphids approached the CMV-infected leaves compared with leaves from the mock-inoculated plants (Figure 1A), indicating that CMV infection increases the attractiveness of Arabidopsis to aphid vectors.
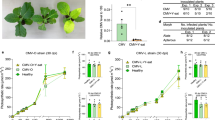
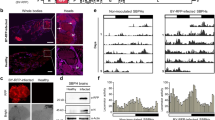
VSR 2b controls viral induction of host attractiveness to aphid vectors. (A) Leaves from CMV-infected plants exhibited higher attractiveness to aphids in circular-dish bioassay. Four-week-old soil-cultivated WT Arabidopsis plants (Col-0) were infected with SD-CMV (CMV) or mock inoculated with buffer (Mock). Upper non-inoculated leaves were used for aphid bioassay 14 days after inoculation. Data are mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. (B) Plants infected by CMV or transgenic for 2b exhibited higher attractiveness to aphids in Y-tube olfactometer bioassay. Six-week-old soil-cultivated Arabidopsis plants transgenic for 2b (2b) and WT were used in the assay. The number of aphids that made a choice is 65 in the comparison of CMV and Mock, and 57 in the comparison of Arabidopsis plants transgenic for 2b (2b) and the wildtype (WT). Data are shown as percentages. **P< 0.01, χ2-test. (C) Plants transgenic for 2b exhibited higher attractiveness to aphids in circular-dish bioassay. Leaves of WT Arabidopsis plants or Arabidopsis plants transgenic for 2b (2b) grown in MS media for 3 weeks were used. Data shown are mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. (D) Fny-CMVΔ2b caused similar symptom as Fny-CMV in dcl2/4 plants. Four-week-old dcl2/4 seedlings were inoculated with Fny-CMV, Fny-CMVΔ2b or buffer (Mock). Photos were taken 14 days post inoculation. Plant tissues that exhibited obvious disease symptoms were indicated within red circles. (E) Fny-CMV but not Fny-CMVΔ2b induced aphid attraction in dcl2/4 plants. The number of aphids that made a choice is 60 in both the comparison of Fny-CMV and Mock, or Fny-CMVΔ2b and Mock. Data are shown as percentages. **P< 0.01, χ2-test.
In the Y-tube olfactometer bioassay, two airstreams carrying volatiles emitted from soil-grown Arabidopsis plants either systemically infected with SD-CMV or mock inoculated flowed through the two arms of the Y-tube. Individual apterous aphids were released from the base of Y-tube stem and allowed to make the choice to crawl toward either arm of the Y-tube while the plants were invisible to the aphids (Supplementary information, Figure S1C). CMV-infected plants also attracted significantly more aphids than mock-inoculated plants (Figure 1B) in this assay. Thus CMV infection in Arabidopsis can induce odor-dependent attraction to aphid vectors.
Previous studies showed that 2b suppresses host antiviral RNAi with its VSR activity to ensure CMV infection12. To examine whether 2b has a role in viral induction of odor-dependent host attractiveness to aphid vectors, we compared feeding preferences of aphids between wild-type Arabidopsis plants and transgenic plants stably expressing 2b. Similar to CMV-infected plants, transgenic plants expressing 2b were significantly more attractive to aphids than wild-type plants (Figure 1B and 1C; Supplementary information, Figure S2). These data demonstrate that expression of 2b alone in Arabidopsis is sufficient to induce odor-dependent aphid attraction.
To verify if 2b protein is required to control viral induction of host attractiveness to aphid vectors, we examined aphid attraction induced by Fny-CMV (another subgroup I strain of CMV) and the 2b deletion mutant, Fny-CMVΔ2b35. As shown in Figure 1D, Fny-CMVΔ2b was able to successfully infect Arabidopsis double mutant dcl2/dcl4(dcl2/4), which is deficient in host antiviral RNAi and, therefore, is susceptible to mutant CMV virus without VSR activity15,36,37, causing similar symptoms as Fny-CMV (Figure 1D). Consistent with what was observed in SD-CMV (Figure 1A and 1B), Fny-CMV infection significantly increased aphid attraction in dcl2/4 plants compared with mock-inoculated dcl2/4 plants in the Y-tube bioassay (Figure 1E). However, no significant difference in aphid attraction was observed between Fny-CMVΔ2b-infected and mock-inoculated dcl2/4 plants (Figure 1E). Both mutant virus Fny-CMVΔ2b and wild-type virus Fny-CMV caused similar viral disease symptom in dcl2/4 plants (Figure 1D); however, enhanced aphid attraction was detected in the dcl2/4 plants only after infection with Fny-CMV, but not with Fny-CMVΔ2b (Figure 1E), suggesting that the 2b protein, but not other viral components, is responsible for viral induction of host attractiveness to aphids.
Taken together, these data demonstrate that 2b is the viral effector responsible for CMV-induced odor-dependent aphid attraction, and suggest that 2b serves as a VIA in addition to its well-defined role as a VSR.
2b physically interacts with host JAZ proteins in the JA pathway
2b was previously found to alter the normal expression of many Arabidopsis genes regulated by abscisic acid38, salicylic acid39 or JA40. In particular, VSR 2b was found to inhibit normal responses of over 90% of Arabidopsis genes regulated by JA40. We thus determined whether 2b interacts with Arabidopsis proteins required in the early steps of JA signaling. Our in vitro pull-down results showed that 2b specifically interacted with JAZ1, but not COI1 or MYC2 (Figure 2A and Supplementary information, Figure S3). Moreover, JAZ1 co-immunoprecipitated with 2b in the extract of Arabidopsis plants expressing both flag-JAZ1 and myc-2b transgenes (Figure 2B). Notably, 2b interacted also with Arabidopsis JAZ3, JAZ6 and JAZ10 (Figure 2C), suggesting that the viral effector protein 2b can interact with multiple JAZ family proteins.
2b interacts with Arabidopsis JAZ proteins. (A) 2b interacts with Arabidopsis JAZ1, but not with COI1 or MYC2. Precipitation of MBP-JAZ1, MBP-MYC2 and flag-COI1 with equal amount of immobilized GST-2b in the pull-down assay was detected by immunoblots using anti-MBP or anti-flag antibody. The input abundances of MBP-JAZ1, MBP-MYC2 and flag-COI1 were detected by Coomassie blue staining (Input). (B) 2b interacts with JAZ1 in co-IP assay. Total proteins extracted from Arabidopsis seedlings expressing flag-JAZ1, myc-2b or both flag-JAZ1 and myc-2b(flag-JAZ1/myc-2b) were incubated with anti-flag affinity resin. Flag-JAZ1 and myc-2b proteins before (Input) and after immunoprecipitation (co-IP) were detected by immunoblots using anti-flag or anti-myc antibody. The smaller-sized bands in the anti-flag antibody-detected co-IP samples were from partially degraded flag-JAZ1 protein. (C) 2b interacts with various Arabidopsis JAZ proteins including JAZ3, JAZ6 and JAZ10. Precipitation of MBP-JAZs and MBP with equal amount of immobilized GST-2b was detected by immunoblots using anti-MBP antibody. Input abundances of MBP-JAZs and MBP were detected by Coomassie blue staining (Input). (D) 2b interacts with JAZ1 in luciferase complementation imaging (LCI) assay. Luciferase activity was detected 50 h after co-expression of indicated construct pairs in leaves of N. benthamiana. (E) 2b interacts with JAZ1 and JAZ10 in bimolecular fluorescence complementation (BiFC) assay. YFP signal was detected 50 h after the co-expression of indicated construct pairs in leaves of N. benthamiana.
We used two independent approaches to determine if interaction of 2b with JAZ1 occurs in vivo. Luciferase (LUC) complementation imaging (LCI) assay showed that co-expression of JAZ1 fused with N-terminal half of LUC and 2b fused with C-terminal half of LUC in Nicotiana benthamiana leaves produced intense luminescence (Figure 2D). Similarly, yellow fluorescence was detected only when C-terminal half of yellow fluorescence protein (YFP) fused with 2b is co-expressed with N-terminal half of YFP fused with either JAZ1 or JAZ10 in the bimolecular fluorescence complementation (BiFC) assay (Figure 2E and Supplementary information, Figure S4). Taken together, our findings demonstrate that 2b interacts with Arabidopsis JAZ proteins in vitro and in vivo.
As an overlapping gene thought to have emerged by a mechanism known as evolutionary overprinting, 2b is the most variable among the five viral genes encoded by different CMV strains41. Nevertheless, we found that 2b encoded by Q-CMV, a subgroup II strain of CMV, also interacted with multiple Arabidopsis JAZ proteins (Figure 3A and Supplementary information, Figure S5).
Interaction between 2b and host JAZ proteins is a conserved feature of CMV infection. (A) Q2b interacts with multiple Arabidopsis JAZ proteins. Precipitation of MBP-JAZs and MBP with equal amount of immobilized GST-Q2b (2b encoded by Q-CMV) was detected by immunoblots using anti-MBP antibody. Input abundances of MBP-JAZs were detected by Coomassie blue staining (Input). (B-E) 2b interacts with JAZ proteins of Nicotiana tabacum. Interactions of 2b proteins encoded by SD-CMV (2b) and Q-CMV (Q2b) with NtJAZ1 (B, D) and NtJAZ2 (C, E) were detected by LCI assays.
Tobacco plant (Nicotiana tabacum), another natural host of CMV, encodes a large JAZ family of proteins, of which NtJAZ1 and NtJAZ2 show the highest similarity with Arabidopsis t. JAZ1 (AtJAZ1) and AtJAZ3, respectively (Supplementary information, Figure S6). In the LCI assay, both NtJAZ1 and NtJAZ2 interacted with the 2b protein from CMV strains of both subgroup I and II (Figure 3B-3E). These results indicate that the interaction between the viral 2b protein and host JAZ proteins is a conserved feature of CMV infection.
2b prevents JA-induced recruitment of JAZ1 by COI1
Previous studies showed that recruitment of JAZs by COI1 depends on the interaction of COI1 with the C-terminal Jas motif of JAZ proteins24,25,42. Consistently, recruitment of JAZ1 by COI1 was abolished when the Jas motif was deleted or disrupted by point mutations (R205A and R206A) (Figure 4A). Furthermore, both the deletion and the point mutations in the Jas motif markedly reduced the interaction between 2b and JAZ1 (Figure 4B) as it reduced the interaction between COI1 and JAZ1 interaction (Figure 4A).
2b prevents JA-induced recruitment of JAZ1 by COI1. (A, B) The Jas motif is essential for the interaction of JAZ1 with COI1 and 2b. Precipitation of flag-COI1 (equal input) with the immobilized MBP-JAZ1, MBP-JAZ1ΔJas and MBP-JAZ1A205A206, in the presence of 0.1 μM coronatine, was detected by immunoblots using anti-flag antibody (A). Precipitation of MBP-JAZ1, MBP-JAZ1ΔJas, MBP-JAZ1A205A206 and MBP with equal amount of immobilized GST-2b was detected by immunoblots using anti-MBP antibody (B). Input abundances of MBP and MBP-fused JAZ1 derivatives were detected by Coomassie blue staining (Input). 2b attenuates COI1-JAZ interaction (C). Precipitation of MBP-JAZ1 with the immobilized flag-COI1 in the presence of 0.1 μM coronatine and the indicated amount (in μg) of 2b-His (or buffer and GST as controls) was detected by immunoblots using anti-MBP antibody. Equal amounts of flag-COI1 and MBP-JAZ1 were used for each reaction in this experiment. Abundances of flag-COI1 on the affinity resins after washes were detected with Memstain.
Coronatine is a JA mimic known to efficiently activate the association of COI1 with JAZ122. We found that 2b was unable to pull down COI1 through JAZ1 in the presence of coronatine (Supplementary information, Figure S7), suggesting that the interaction between JAZ1 and 2b is incompatible with JAZ1-COI1 interaction. Furthermore, competitive binding assay with COI1 as prey showed that the abundance of JAZ1, which was pulled down by COI1, decreased when 2b (Figure 4C) was increased. Taken together, these findings suggest that 2b competes with COI1 in binding to the same domain of JAZ1, thereby preventing JA-induced recruitment of JAZ1 by COI1.
2b prevents JA-induced degradation of JAZ1 thereby suppressing JA signaling
JA-induced degradation of JAZ proteins depends on their interaction with the F-box protein COI124,25. We then determined if the observed inhibition of the interaction between COI1 and JAZ1 by 2b attenuates the COI1-dependent degradation of JAZ proteins. In our in vitro assay conditions, recombinant MBP-JAZ1 protein was rapidly degraded after the addition of crude protein extracts from wild-type Arabidopsis, but neither the extract from coi1-1 mutant plants nor the extract from wild-type plant treated with MG132 was effective (Figure 5A and 5B; Supplementary information, Figure S8A and S8B), consistent with COI1-dependent degradation of JAZ1 by the 26S proteasome pathway as shown previously24,25.
2b inhibits JA-induced degradation of JAZ1. (A-D) 2b suppresses JAZ1 degradation in the in vitro assays. Purified MBP-JAZ1 was incubated with total crude extract from WT plants and coi1-1 plants (A), total WT crude extract plus MG132 or ethanol (Mock) (B), total crude extract from WT plants infected by CMV Shandong strain (CMV) or the Mock control (C), total WT crude extract plus purified GST-2b or GST control (D), total crude extract from WT plants and myc-tagged 2b transgenic plants (myc-2b) (E). MBP-JAZ1 was detected by immunoblots using anti-MBP antibody after the indicated incubation time. Large subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase (LSU) detected with Memstain was used as a control for equal loading. (F-H) 2b suppresses JA-induced JAZ1 degradation in vivo. Endogenous flag-JAZ1, before (F) or after MeJA treatment (G, H), was detected by immunoblot using anti-flag antibody in paired plants, including CMV-infected (CMV) and mock-inoculated (Mock) plants transgenic for flag-JAZ1 (flag-JAZ1) (F, G), or flag-JAZ1 plants and plants transgenic for both flag-JAZ1 and myc-2b (flag-JAZ1/myc-2b) (F, H).
Furthermore, we found that JAZ1 degradation in the crude extract from wild-type plants infected with CMV was markedly slower compared with the extract from mock-inoculated plants (Figure 5C and Supplementary information, Figure S8C). The degradation of JAZ1 was also inhibited either by adding recombinant 2b protein (GST-2b) to the crude extract from healthy wild-type plants (Figure 5D and Supplementary information, Figure S8D), or by the crude extract from transgenic plants expressing 2b (myc-2b) (Figure 5E and Supplementary information, Figures S8E, S8F and S9).
Furthermore, our in vivo degradation assays showed that endogenous JAZ1, which is stably expressed in plants carrying flag-JAZ1 transgene, accumulated to a higher level in plants infected by CMV or with transgenic expression of 2b, compared with the control plants (Figure 5F). Furthermore, rapid degradation of endogenous flag-JAZ1 protein, which was induced by exogenous JA treatment, was also suppressed by either CMV infection (Figure 5G) or transgenic expression of 2b (Figure 5H). Together, these findings demonstrate that both CMV infection and transgenic expression of 2b inhibit JA-induced, COI1-dependent degradation of JAZ proteins.
JA-induced degradation of JAZ proteins results in the activation of the JA pathway18. Consistent with the previous reports40,43, our quantitative RT-PCR analysis showed that either CMV infection or the transgenic expression of 2b in Arabidopsis plants significantly inhibited JA-dependent induction of many genes (Supplementary information, Figure S10A and S10B), including LOX2, TAT1 and VSP1, as well as CYP81D11, MYC2 and TPS10, the latter three genes encoding regulators of plant volatile biosynthesis. Furthermore, we generated Arabidopsis plants transgenic for β-glucuronidase reporter gene under the control of promoter of the JA-inducible VSP1 gene (PVSP1-GUS) either alone or together with a 2b transgene. Histochemical staining revealed that transgenic expression of 2b markedly inhibited JA-induced GUS activity in Arabidopsis plants (Supplementary information, Figure S10C), providing an independent verification of the inhibition of JA signaling by 2b.
2b targets the JA pathway for viral induction of host attractiveness to aphid vectors
Having shown that 2b interacts with and represses JA-induced degradation of host JAZ proteins thus suppressing JA signaling (Figures 2, 3, 4, 5), we next examined the biological effect of 2b repression on JAZ degradation by comparing feeding preferences of aphids between wild-type Arabidopsis plants and coi1-1 mutant, which is defective in JA perception and accumulates higher amounts of JAZ proteins18. As shown in Figure 6A, leaves from coi1-1 mutant plants attracted significantly more aphids than those from wild-type plants in the circular dish bioassay (Figure 6A). Consistently, Y-tube olfactometer bioassay showed that less aphids approached wild-type plants than the coi1-1 mutant plants (Figure 6B). Furthermore, leaves of JA-treated wild-type plants attracted fewer aphids than leaves from the untreated wild-type plants in the circular dish bioassay (Figure 6C). On the contrary, JA treatment had no obvious effect on the aphid attraction of coi1-1 (Figure 6D). These results demonstrate that the genetic disruption of JA signaling induces the plant attractiveness to aphid vectors and that JA signaling represses the odor-dependent plant attractiveness to aphid vectors in a COI1-dependent manner.
JA suppresses aphid attraction. (A, B) coi1-1 mutant showed higher attractiveness to aphids than the WT in circular-dish bioassay (A) and Y-tube olfactometer bioassay (B). For the circular-dish bioassay, data are mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. For the Y-tube olfactometer bioassay, number of aphids that made a choice in this comparison was 56, data are shown as percentages. **P< 0.01, χ2-test. (C, D) JA treatment reduces attractiveness of WT leaves but not coi1-1 leaves to aphids in circular-dish bioassay. Three-week-old WT or coi1-1 plants grown on MS medium were pretreated with 100 μM MeJA (JA) or equal concentration of ethanol (Mock) 8 h before the aphid bioassays. Data are mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. (E, F) myc2, myc3 and myc4 (myc2/3/4) mutant plants showed higher attractiveness to aphids than WT plants in circular-dish bioassay (E) and Y-tube olfactometer bioassay (F). For the circular-dish bioassay, data are mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. For the Y-tube olfactometer bioassay, number of aphids that made a choice in this comparison was 66. Data are shown as percentages. **P< 0.01, χ2-test.
Among JAZ-repressed downstream components, MYC2, MYC3 and MYC4 were previously found to have important roles in JA-regulated plant defense including plant volatile biosynthesis18,30,33. Thus, we next examined whether these MYCs were involved in JA-regulated odor-dependent plant attractiveness to aphids. As shown in Figure 6E and 6F, the myc2/3/4 triple mutant plants exhibited higher aphid attraction than wild-type plants in both the circular dish bioassay and Y-tube olfactometer bioassay (Figure 6E and 6F). These results suggest that JA signaling represses the odor-dependent plant attractiveness to aphid vectors through MYC2, MYC3 and MYC4.
VIA function of 2b is independent of its VSR activity
Having uncovered a novel VIA function for 2b, we next investigated whether the VSR activity of 2b is required for the VIA function. Previous study showed that Fny2blm, a double mutant 2b (L15A and M18A) derived from Fny-CMV, is drastically compromised in VSR activity44. Fny2blm nevertheless was able to efficiently interact with JAZ1 (Figure 7A) and inhibit the JA-induced JAZ1 degradation (Figure 7B; Supplementary information, Figure S8G and S8H). Moreover, transgenic expression of Fny2blm effectively enhanced plant attractiveness to aphid vectors in both the circular dish and Y-tube olfactometer bioassays (Figure 7C-7E). These results demonstrate that Fny2blm, which lacks the VSR activity44, still retains the VIA activity.
VIA function of 2b is independent of its VSR activity. (A) Both Fny2b and Fny2blm interact with JAZ1 in pull-down assay. Precipitation of MBP-Fny2b, MBP-Fny2blm and MBP with equal amount of immobilized GST-JAZ1 was detected by immunoblots using anti-MBP antibody. Input abundances of MBP-Fny2b, MBP-Fny2blm and MBP were detected by Coomassie blue staining (Input). (B) Both Fny2b and Fny2blm inhibit JAZ1 degradation in the in vitro degradation assay. Amount of purified MBP-JAZ1 in total crude protein extract from WT and Fny2b transgenic plants, or from WT and Fny2blm transgenic plants was detected by immunoblots using anti-MBP antibody after the indicated incubation time. LSU detected with Memstain was used as a control for equal loading. (C-E) Both Arabidopsis plants transgenic for Fny2b and Fny2blm showed higher attractiveness to aphids than WT in circular-dish bioassay (C, D) and Y-tube bioassay (E). For the circular-dish bioassays, data are shown as mean percentages ± SEM, n = 5. **P< 0.01, paired samples Student's t-test. For the Y-tube bioassays, number of aphids that made a choice in the comparison of WT vs Fny2b and WT vs Fny2blmwas 60 and 64, respectively. Data are shown as percentages. **P< 0.01, χ2-test. (F) Fny2blm suppresses JA-inducible gene expression. The relative mRNA abundance of MYC2, CYP81D11, TPS10, LOX2, TAT1 and VSP1 was detected by quantitative RT-PCR in 14-day-old MS medium-cultivated WT and Arabidopsis plants transgenic for Fny2b or Fny2blm, treated with MeJA (100 μM) for the indicated time periods. Data are means ± SEM, n = 3. *P< 0.05, **P< 0.01, Student's t-test significance compared with WT.
Further quantitative RT-PCR analysis of Arabidopsis plants transgenic for Fny2b or Fny2blm showed that JA-dependent induction of many genes was severely repressed in both Fny2b- and Fny2blm-transgenic plants compared with wild-type plants (Figure 7F). These data demonstrate that Fny2blm still maintains the ability to inhibit the expression of JA response genes, indicating that the 2b inhibition of JA response genes shown in this work (Figure 7F; Supplementary information, Figure S10) and in the previous studies40,43 may contribute largely to its VIA action (induction of host attractiveness to insect vectors) but not to the VSR activity (suppression of host defense against virus infection). In summary, our results demonstrate that VIA function of 2b, which inhibits JA signaling by stabilizing host JAZ proteins and suppressing MYC factors, is independent of its VSR activity.
Discussion
Recent studies have uncovered novel strategies arthropod-borne pathogens adopt to manipulate their hosts45,46. In principle, these novel functions represent a core feature of arthropod-borne pathogens and are vital to their adaptive success during the co-evolution with their host and vector organisms. However, the molecular mechanisms involved in host manipulation by arthropod-borne pathogens are currently largely unclear. For example, it is unknown whether pathogen-induced host odor-dependent attractiveness to vectors is elicited by specific pathogen effectors and how these pathogen effectors target host pathways to effect host manipulation. In this study, we used an Arabidopsis model to uncover the mechanisms underlying odor-dependent vector attraction induced by CMV, an economically important aphid-borne plant pathogen11. We show that the CMV-induced vector attraction is under the control of the signaling pathway of the host hormone JA, which represses odor-dependent host attractiveness to aphids through MYC2 and its homologs MYC3 and MYC4. We define viral 2b protein, the previously known VSR of CMV, as a VIA. 2b physically interacts with host JAZ proteins and prevents JA-induced JAZ degradation, thus attenuating JA signaling to enhance host attractiveness to aphid vectors. 2b acts differently from the bacterial effectors HopZ1a and HopX1, which promote JAZ degradation and activate JA signaling47,48.
2b protein from either of the two CMV subgroup strains interacts with JAZ proteins from both Arabidopsis and tobacco plants, indicating that targeting the repressor of JA signaling to inhibit JA-induced responses is an evolutionarily conserved function of the aphid-borne virus. Notably, odor-dependent vector attraction is suppressed by the induction of JA signaling and is enhanced in the Arabidopsis coi1-1 mutant defective in JA signaling. Further studies are expected to identify the exact volatile(s) that mediates the JA-regulated and CMV-induced aphid attraction.
Interestingly, our study shows that 2b interacts with JAZ proteins known to bind both COI1 and MYCs (Figures 2, 3, 4)18. It has been reported that, in the resting stage, JAZs exist in complex with MYCs (JAZ-MYC complex)27. After perceiving JA, COI1 interacts with JAZ proteins and recruits them for degradation, thereby releasing MYCs to activate JA signaling18,27. Upon CMV infection, 2b binds JAZs and inhibits COI1-mediated JAZ degradation, resulting in an elevated level of JAZs. As 2b-JAZ interaction is dynamic and reversible, an increase in JAZ proteins by 2b could conceivably cause an increase in the amount of JAZ-MYC complex in the cells by attenuating the dissociation of pre-existing JAZ-MYC complex and promoting the formation of new JAZ-MYC complex. This model is consistent with both the suppression of JA signaling by 2b and the observation that genetic suppression of MYC function in myc2/3/4 triple mutant plants also induces aphid attraction.
Suppression of antiviral RNAi defense of the host is a widespread strategy in plant and invertebrate viruses12. 2b of CMV is a well-characterized VSR that effectively suppresses host RNAi-directed antiviral machinery12. In this study, we show that the VSR 2b also serves as a VIA to control viral induction of host attractiveness to insect vectors, and its VIA activity is independent of its VSR activity. Moreover, our study demonstrates how 2b interferes with the JA pathway and inhibits expression of many JA-regulated Arabidopsis genes that are essential for generation of both insect-repellent volatiles and feeding-deterrent/toxic products, thereby providing mechanistic understanding for our observation that CMV 2b induces odor-dependent host attractiveness to aphids, and also for the previously reported phenomenon that CMV 2b suppresses host resistance against insect feeding49.
Our study shows that JA represses plant attractiveness to insect vectors (Figure 6), and the previously defined VSR 2b inhibits JA signaling to serve as a VIA, independent of its VSR activity, for inducing odor-dependent vector attraction (Figures 1, 2, 3, 4, 5 and 7). Interestingly, Carr and colleagues reported recently that four unrelated VSRs encoded by both insect-transmissible and non-insect-transmissible plant viruses have a shared activity to suppress JA-induced host gene expression43. Future studies are required to experimentally verify whether all these VSRs, which repress JA signaling43, induce odor-dependent host attractiveness to insects. Although some viruses may never be insect-transmissible due to their physiochemical properties including instability of virions inside the body of insects or inability to be acquired due to the lack of interaction with insect stylets50,51, we propose that such a virus-induced odor-dependent insect attraction has a role in the emergence and evolution of arthropod-borne pathogens. Pathogen-induced odor-dependent insect attraction will allow more insects to move toward the infected plants, thus providing more opportunity for viruses to interact with insects, which might be essential for some viruses to evolve and adapt to a vector-borne lifestyle.
Materials and Methods
CMV inoculation
For CMV infection, 4-week-old Arabidopsis plants that grown under a 10-h (23 °C)/14-h (19 °C) light/dark photoperiod were lightly dusted with silicon carbide and then rub-inoculated with purified virions at the concentration of 20 μg/ml. Mock-treated plants were inoculated with buffer (5 mM sodium borate, 0.5 mM disodium EDTA, pH = 9.0) only. The CMV-infected and mock-inoculated plants were used for aphid attraction bioassays, gene expression assays and protein degradation assays 2 weeks post inoculation.
Generation of transgenic plants
The full-length coding sequence of 2b was amplified from the cDNA of SD-CMV and inserted into modified pROK2 binary vector to generate the pMyc-2b construct. The full-length coding sequence of JAZ1 was amplified from Arabidopsis cDNA and constructed into modified pCAMBIA1300 binary vector (driven by 35S promoter and fused with 3× flag tag) to generate the pCAMBIA1300-3flag-JAZ1 construct. The pMyc-2b and pCAMBIA1300-3flag-JAZ1 constructs were then transformed into Arabidopsis Col-0 wild-type plants using the Agrobacterium-mediated floral dip method to get the myc-2b and flag-JAZ1 transgenic plants, respectively. Two individual transgenic lines of myc-2b, including myc-2b-2 (myc-2b) and myc-2b-4 were used in this study. Arabidopsis plants transgenic for both myc-2b and flag-JAZ1 (flag-JAZ1/myc-2b) were obtained by crossing myc-2b plants with flag-JAZ1 plants.
The Fny2b or Fny2blm constructs, the Fny-CMV or Fny-CMVΔ2b viruses, and the Arabidopsis plants transgenic for Fny2b or Fny2blm were described previously35,37,44.
Aphid attraction bioassays
Wingless M. persicae aphids maintained on Raphanus sativus were used for aphid bioassays (as shown in Supplementary information, Figure S1A). Aphids were isolated from R. sativus and starved for 6 h before testing.
For the circular dish bioassays, plants were cultivated on MS medium under a 16-h (23 °C)/8-h (19 °C) light/dark photoperiod for 3 weeks. As shown in the schematic diagram (Supplementary information, Figure S1B), in each pair-wised comparison, 12-15 leaves from each genotype were placed alternately in a circular dish, which was 15 cm in diameter and padded with three layers of water wet filter papers to prevent water loss of the leaves. In each time of the experiment, 60-100 aphids were released from the center of the dish for free walk, and their trace were recorded by a digital camera. An aphid was considered to have made its choice once it reached the leaf of either genotype and didn't walk directly away without a stop. The choices of aphids were counted according to the video records and the ratio between the numbers of aphids choosing each of the paired plants was calculated.
For the Y-tube olfactometer bioassays, plants grown in soil under a 10-h (23 °C)/14-h (19 °C) light/dark photoperiod for 5 weeks were used. As CMV infection or 2b transgenic expression causes morphological changes in Arabidopsis, such as dwarf and curled leaves (data not shown)14,37,44, to ensure that equal weight of each kind of plants were used for the pair-wise comparisons in Y-tube olfactometer bioassays, we firstly weighed plants with CMV infection or 2b transgenic expression and the corresponding control plants, and then used the soil-grown plants (which grew at the same batch and had similar size with the above-weighed plants) for the pair-wise comparisons. As shown in the schematic diagram (Supplementary information, Figure S1C), plants that used for pair-wise comparison were placed into two odor source bottles, which connected to the two arms of the Y-tube. Under the action of the aspirator, air that was cleaned by activated charcoal and humidified by water flowed through the odor source bottles and flow meters into the arms of the Y-tube, which was placed in an enclosed incubator with diffused light. The rate of airflow in each arm was maintained at 200 ml/min. The inner diameter of the Y-tube was 1.0 cm. The stem and the arms of the Y-tube were 6 and 10 cm in length, respectively. In each test, an individual aphid was introduced at the end of the Y-tube stem and walked upwind toward the arms. The aphid was considered to have made its choice once it entered one arm and walked across the decision line that marked 3 cm away from the Y-junction. A “no choice” outcome was recorded and the aphid was replaced if it failed to make decision during the 5 min testing period. To compensate any unknown asymmetry in the setup, the odor sources were interchanged after every 6 tests, and the Y-tube and plants were replaced after every 12 tests. The used Y-tubes were cleaned and dried with 180 °C oven before reuse.
Preparation of recombinant proteins
To obtain recombinant GST-tagged 2b and JAZ1 proteins, the coding sequences of 2b from SD-CMV or Q-CMV and Arabidopsis JAZ1 were cloned into modified pGEX-4T-2 vector. To obtain recombinant MBP-JAZ proteins, the coding sequences of various JAZ genes were cloned into pMAL-C2X vector. These constructs were then introduced into Escherichia coli BL21 strain, and the recombinant proteins were purified using corresponding affinity resin followed by size exclusion chromatography.
Quantitative RT-PCR
Total RNAs were extracted from indicated plant materials using Transzol reagent (Transgen) and reverse transcribed using TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix (Transgen).Quantitative RT-PCR was performed with ABi7500 real-time PCR system using SYBR Select Master Mix (Life Technologies). ACTIN8 was used as the internal control.
GUS staining
For the GUS staining assay, homozygous PVSP1-GUS (GUS reporter gene dirven by VSP1 promoter) transgenic plants were crossed with heterozygous myc-2b transgenic plant, and the PVSP1-GUS plants in background of WT (PVSP1-GUS/WT) and myc-2b (PVSP1-GUS/myc-2b) were identified in the F1 population. Two-week-old PVSP1-GUS/WT and PVSP1-GUS/myc-2b seedlings were treated with 100 μM MeJA or equal amount of ethanol (Mock) for 8 h and then incubated in GUS staining buffer (1 mM X-glucuronide, 0.5 mM potassium ferricyanide, 0.5 mM potassium ferrocyanide, 10 mM pH 8.0 EDTA, 1% Triton X-100) at 37 °C overnight. The stained seedlings were cleared by washing with 70% ethanol.
Pull-down assay
For pull-down assays, about 50 μg of bait proteins were used to bind with corresponding resin (amylose resin for MBP-fused proteins, glutathione resin for GST-fused proteins and anti-flag resin for flag-fused proteins), and then were further incubated with about 10 μg prey proteins at 4 °C for 1 h in RB buffer (50 mM Tris-HCl, pH 7.8, 100 mM NaCl, 0.1% (v/v) Tween 20, 10% (v/v) glycerol and 20 mM β-mercaptoethanol). Precipitated prey proteins on the resin were released by boiling at 100 °C for 5 min and detected by immunoblots using corresponding antibodies. The relative input abundances of the bait or prey proteins were detected by Coomassie blue staining.
For pull-down assay in Supplementary information, Figure S3, 50 μg of MBP and MBP-JAZ1 proteins were used to bind with amylose resin, and then further incubated with 1 ml concentrated total proteins extracted from 2 g myc-2b transgenic plants. Precipitated myc-2b on the beads was detected by immunoblots using anti-myc antibody.
For competitive pull-down assay in Figure 4C, 50 μg of flag-COI1 proteins were used to bind with anti-flag resin, and were further incubated with 10 μg MBP-JAZ1 plus increasing amount of 2b-His (10, 20 and 50 μg) or buffer and GST(50 μg) as controls, in the presence of 0.1 μM coronatine. Precipitated MBP-JAZ1 on the beads were detected by immunoblots using anti-MBP antibody.
Co-IP assay
For Co-IP assay, Arabidopsis plants transgenic for both myc-2b and flag-JAZ1 (flag-JAZ1/myc-2b) or for myc-2b, flag-JAZ1 alone, were used in the Co-IP assay. Total proteins were extracted from 2 g seedlings using 15 ml RB-plus buffer (RB buffer plus 1× EDTA-free complete mini protease inhibitor cocktail and 20 μM MG132) and concentrated to 1 ml using 10 kD ultrafiltration tubes (Millipore). About 50 μl anti-flag resin beads were added into 800 μl concentrated total proteins and incubated at 4 °C for 1 h. The beads were then washed with RB plus buffer for five times and resuspended with SDS loading buffer. The precipitated proteins on the beads were released by boiling at 100 °C for 5 min and detected by immunoblots using anti-myc and anti-flag antibody.
BiFC and LCI assays
For BiFC assays, the full-length coding sequence of 2b from SD-CMV was cloned into cYFP vector to get the cYFP-2b construct, and the full-length coding sequences of Arabidopsis JAZ1 and JAZ10 were cloned into the nYFP vector to get the JAZ1-nYFP and JAZ10-nYFP constructs. These constructs were then transformed into Agrobacterium tumefaciens strain GV3101 and transiently co-expressed with indicated combinations in the N. benthamiana leaves using Agrobacterium infiltration method. The YFP fluorescence signal for each combination was detected using confocal microscope (Zeiss LSM710) 50 h after infiltration.
For LCI assays, the full-length coding sequences of 2b from SD-CMV and Q-CMV were cloned into cLUC vector, and the full-length coding sequences of JAZ1 and JAZ2 from Arabidopsis and N. tabacum were cloned into nLUC vector. These constructs were transformed into A. tumefaciens strain GV3101 and co-expressed with indicated combinations in N. benthamiana leaves for 50 h. The LUC substrate (luciferin) was then sprayed onto the surface of the leaves and images were taken using a low-light cooled imaging apparatus (Andor iXon).
In vitro and in vivo protein degradation assay
For the in vitro degradation assays of JAZ1 in various conditions, 5 μg of purified MBP-JAZ1 protein was added to 600 μl crude total proteins (1 μg/μl) extracted from indicated plants using RB buffer. The mixture was incubated at 22 °C and sampled at indicated time points. The remained abundance of MBP-JAZ1 in these samples was detected by immunoblots using anti-MBP antibody. For the degradation assay in Figure 5B, 10 μM of MG132 or equal amount of ethanol (Mock) was added into the crude extracts from wild-type plants before adding MBP-JAZ1. For the degradation assay in Figure 5D, MBP-JAZ1 was incubated with 20 μg GST-2b or GST at 4 °C for 1 h before added to the crude extracts from wild-type plants. For quantitative analysis, the band densities of MBP-JAZ1 were quantified using ImageJ2x software.
For the in vivo degradation assay in Figure 5G, 4-week-old flag-JAZ1 transgenic plants that are grown under a 10-h (23 °C)/14-h (19 °C) light/dark photoperiod were inoculated with SD-CMV (CMV) or buffer (Mock), and then treated with MeJA (30 μM) for indicated time periods 2 weeks after inoculation. The remained abundance of flag-JAZ1 in these samples was detected by immunoblots using anti-flag antibody. For the in vivo degradation assay in Figure 5H, 3-week-old flag-JAZ1 or flag-JAZ1/myc-2b plants that are grown on MS were treated with 10 μM MeJA for indicated time periods before immunoblot detection.
Accession numbers
Sequence data described in this article can be found in GenBank/EMBL or TAIR (www.Arabidopsis.org) under the following accession numbers: COI1 (AT2G39940), LOX2 (AT3G45140), MYC2 (AT1G32640), CYP81D11 (AT3G28740), TPS10 (AT2G24210), VSP1 (AT5G24780), TAT1 (AT4G23600), JAZ1 (AT1G19180), JAZ3 (AT3G17860), JAZ6 (AT1G72450), JAZ10 (AT5G13220), ACTIN8 (AT1G49240), SD2b (D86330), Q2b (Z21863), Fny2b (NC_002035), NtJAZ1 (AB433896) and NtJAZ2 (AB433897).
Author Contributions
DX, SWD and LK designed the research; DW and TQ performed the experiments with the help of WL, HT, HG, JW, JG and RY; DX, SWD, LK, DW, TQ, CR, XW and YL analyzed the data; DX, DW, TQ and SWD wrote the manuscript with contributions of all the authors.
Competing Financial Interests
The authors declare no competing financial interests.
References
Liang G, Gao X, Gould EA . Factors responsible for the emergence of arboviruses; strategies, challenges and limitations for their control. Emerg Microbes Infect 2015; 4:e18.
Caplan J, Padmanabhan M, Dinesh-Kumar SP . Plant NB-LRR immune receptors: from recognition to transcriptional reprogramming. Cell Host Microbe 2008; 3:126–135.
Ng JC, Falk BW . Virus-vector interactions mediating nonpersistent and semipersistent transmission of plant viruses. Annu Rev Phytopathol 2006; 44:183–212.
McLeod G, Gries R, von Reuss SH, et al. The pathogen causing Dutch elm disease makes host trees attract insect vectors. Proc Biol Sci 2005; 272:2499–2503.
Lacroix R, Mukabana WR, Gouagna LC, Koella JC . Malaria infection increases attractiveness of humans to mosquitoes. PLoS Biol 2005; 3:e298.
O'Shea B, Rebollar-Tellez E, Ward RD, et al. Enhanced sandfly attraction to Leishmania-infected hosts. Trans R Soc Trop Med Hyg 2002; 96:117–118.
De Moraes CM, Stanczyk NM, Betz HS, et al. Malaria-induced changes in host odors enhance mosquito attraction. Proc Natl Acad Sci USA 2014; 111:11079–11084.
Mann RS, Ali JG, Hermann SL, et al. Induced release of a plant-defense volatile 'deceptively' attracts insect vectors to plants infected with a bacterial pathogen. PLoS Pathog 2012; 8:e1002610.
Mauck KE, De Moraes CM, Mescher MC . Deceptive chemical signals induced by a plant virus attract insect vectors to inferior hosts. Proc Natl Acad Sci USA 2010; 107:3600–3605.
Eigenbrode SD, Ding HJ, Shiel P, Berger PH . Volatiles from potato plants infected with potato leafroll virus attract and arrest the virus vector, Myzus persicae (Homoptera : Aphididae). Proc Biol Sci 2002; 269:455–460.
Palukaitis P, Garcia-Arenal F . Cucumoviruses. Adv Virus Res 2003; 62:241–323.
Ding SW . RNA-based antiviral immunity. Nat Rev Immunol 2010; 10:632–644.
Gonzalez I, Rakitina D, Semashko M, et al. RNA binding is more critical to the suppression of silencing function of cucumber mosaic virus 2b protein than nuclear localization. RNA 2012; 18:771–782.
Duan CG, Fang YY, Zhou BJ, et al. Suppression of Arabidopsis ARGONAUTE1-mediated slicing, transgene-induced RNA silencing, and DNA methylation by distinct domains of the cucumber mosaic virus 2b protein. Plant Cell 2012; 24:259–274.
Diaz-Pendon JA, Li F, Li WX, Ding SW . Suppression of antiviral silencing by cucumber mosaic virus 2b protein in Arabidopsis is associated with drastically reduced accumulation of three classes of viral small interfering RNAs. Plant Cell 2007; 19:2053–2063.
Zhang XR, Yuan YR, Pei Y, et al. Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes Dev 2006; 20:3255–3268.
Yan C, Xie D . Jasmonate in plant defence: sentinel or double agent? Plant Biotechnol J 2015; 13:1233–1240.
Wasternack C, Hause B . Jasmonates: biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann Bot 2013; 111:1021–1058.
Browse J . Jasmonate passes muster: a receptor and targets for the defense hormone. Annu Rev Plant Biol 2009; 60:183–205.
Sheard LB, Tan X, Mao H, et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 2010; 468:400–407.
Yan JB, Zhang C, Gu M, et al. The Arabidopsis CORONATINE INSENSITIVE1 protein is a jasmonate receptor. Plant Cell 2009; 21:2220–2236.
Katsir L, Schilmiller AL, Staswick PE, He SY, Howe GA . COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc Natl Acad Sci USA 2008; 105:7100–7105.
Yan Y, Stolz S, Chételat A, et al. A downstream mediator in the growth repression limb of the jasmonate pathway. Plant Cell 2007; 19:2470–2483.
Thines B, Katsir L, Melotto M, et al. JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature 2007; 448:661–665.
Chini A, Fonseca S, Fernández G, et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 2007; 448:666–671.
Xie DX, Feys BF, James S, Nieto-Rostro M, Turner JG . COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science 1998; 280:1091–1094.
Zhang F, Yao J, Ke J, et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 2015; 525:269–273.
Song S, Qi T, Wasternack C, Xie D . Jasmonate signaling and crosstalk with gibberellin and ethylene. Curr Opin Plant Biol 2014; 21:112–119.
Pauwels L, Barbero GF, Geerinck J, et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 2010; 464:788–791.
Hong GJ, Xue XY, Mao YB, Wang LJ, Chen XY . Arabidopsis MYC2 interacts with DELLA proteins in regulating sesquiterpene synthase gene expression. Plant Cell 2012; 24:2635–2648.
Howe GA, Jander G . Plant immunity to insect herbivores. Annu Rev Plant Biol 2008; 59:41–66.
Liechti R, Farmer EE . The jasmonate pathway. Science 2002; 296:1649–1650.
Li R, Weldegergis BT, Li J, et al. Virulence factors of geminivirus interact with MYC2 to subvert plant resistance and promote vector performance. Plant Cell 2014; 26:4991–5008.
Bruce TJA, Matthes MC, Chamberlain K, et al. cis-Jasmone induces Arabidopsis genes that affect the chemical ecology of multitrophic interactions with aphids and their parasitoids. Proc Natl Acad Sci USA 2008; 105:4553–4558.
Ryabov EV, Fraser G, Mayo MA, Barker H, Taliansky M . Umbravirus gene expression helps potato leafroll virus to invade mesophyll tissues and to be transmitted mechanically between plants. Virology 2001; 286:363–372.
Wang XB, Jovel J, Udomporn P, et al. The 21-nucleotide, but not 22-nucleotide, viral secondary small interfering RNAs direct potent antiviral defense by two cooperative argonautes in Arabidopsis thaliana. Plant Cell 2011; 23:1625–1638.
Wang XB, Wu Q, Ito T, et al. RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc Natl Acad Sci USA 2010; 107:484–489.
Westwood JH, McCann L, Naish M, et al. A viral RNA silencing suppressor interferes with abscisic acid-mediated signalling and induces drought tolerance in Arabidopsis thaliana. Mol Plant Pathol 2013; 14:158–170.
Ji LH, Ding SW . The suppressor of transgene RNA silencing encoded by cucumber mosaic virus interferes with salicylic acid-mediated virus resistance. Mol Plant Microbe Interact 2001; 14:715–724.
Lewsey MG, Murphy AM, MacLean D, et al. Disruption of two defensive signaling pathways by a viral RNA silencing suppressor. Mol Plant Microbe Interact 2010; 23:835–845.
Ding SW, Li WX, Symons RH . A novel naturally occurring hybrid gene encoded by a plant RNA virus facilitates long distance virus movement. EMBO J 1995; 14:5762–5772.
Melotto M, Mecey C, Niu Y, et al. A critical role of two positively charged amino acids in the Jas motif of Arabidopsis JAZ proteins in mediating coronatine- and jasmonoyl isoleucine-dependent interactions with the COI1 F-box protein. Plant J 2008; 55:979–988.
Westwood JH, Lewsey MG, Murphy AM, et al. Interference with jasmonic acid-regulated gene expression is a general property of viral suppressors of RNA silencing but only partly explains virus-induced changes in plant-aphid interactions. J Gen Virol 2014; 95:733–739.
Dong K, Wang Y, Zhang Z, et al. Two amino acids near the N-terminus of cucumber mosaic virus 2b play critical roles in the suppression of RNA silencing and viral infectivity. Mol Plant Pathol 2016; 17:173–183.
Mauck K, Bosque-Perez NA, Eigenbrode SD, De Moraes CM, Mescher MC . Transmission mechanisms shape pathogen effects on host-vector interactions: evidence from plant viruses. Funct Ecol 2012; 26:1162–1175.
Lefevre T, Thomas F . Behind the scene, something else is pulling the strings: emphasizing parasitic manipulation in vector-borne diseases. Infect Genet Evol 2008; 8:504–519.
Gimenez-Ibanez S, Boter M, Fernandez-Barbero G, et al. The bacterial effector HopX1 targets JAZ transcriptional repressors to activate jasmonate signaling and promote infection in Arabidopsis. PLoS Biol 2014; 12:e1001792.
Jiang S, Yao J, Ma KW, et al. Bacterial effector activates jasmonate signaling by directly targeting JAZ transcriptional repressors. PLoS Pathog 2013; 9:e1003715.
Ziebell H, Murphy AM, Groen SC, et al. Cucumber mosaic virus and its 2b RNA silencing suppressor modify plant-aphid interactions in tobacco. Sci Rep 2011; 1:187.
Hogenhout SA, Ammar el D, Whitfield AE, Redinbaugh MG . Insect vector interactions with persistently transmitted viruses. Annu Rev Phytopathol 2008; 46:327–359.
Ng JC, Falk BW . Virus-vector interactions mediating nonpersistent and semipersistent transmission of plant viruses. Annu Rev Phytopathol 2006; 44:183–212.
Acknowledgements
We thank Rongxiang Fang and Huishan Guo for the gifts of SD-CMV and plasmids. This work was supported by the National Natural Science Foundation of China (31421001, 31630085 and 31500228), the Ministry of Science and Technology of China (2016YFA0500501) and an endowment from College of natural and Agricultural Sciences, University of California, Riverside, USA.
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
Experimental setup for circular-dish and Y-tube aphid attraction bioassays. (PDF 129 kb)
Supplementary information, Figure S2
The expression level of the 2b protein in transgenic plants. (PDF 28 kb)
Supplementary information, Figure S3
2b interacts with Arabidopsis JAZ1. (PDF 26 kb)
Supplementary information, Figure S4
Negative controls for BiFC assay. (PDF 105 kb)
Supplementary information, Figure S5
Phylogenetic analysis of various CMV strains. (PDF 209 kb)
Supplementary information, Figure S6
Phylogenetic analysis of Arabidopsis and tobacco JAZ proteins. (PDF 95 kb)
Supplementary information, Figure S7
The JAZ-2b and JAZ-COI1 interactions are incompatible. (PDF 140 kb)
Supplementary information, Figure S8
Quantitative analysis of in vitro degradation of JAZ1. (PDF 351 kb)
Supplementary information, Figure S9
2b inhibits in vitro degradation of JAZ1. (PDF 40 kb)
Supplementary information, Figure S10
2b suppresses JA-inducible gene expression. (PDF 363 kb)
Rights and permissions
About this article
Cite this article
Wu, D., Qi, T., Li, WX. et al. Viral effector protein manipulates host hormone signaling to attract insect vectors. Cell Res 27, 402–415 (2017). https://doi.org/10.1038/cr.2017.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2017.2
Keywords
This article is cited by
-
Leafhopper salivary vitellogenin mediates virus transmission to plant phloem
Nature Communications (2024)
-
Virus-induced changes in host plant phenotype cue behavioral changes in Aphis glycines that enhance acquisition and transmission of soybean mosaic virus
Journal of Pest Science (2024)
-
A plant cytorhabdovirus modulates locomotor activity of insect vectors to enhance virus transmission
Nature Communications (2023)
-
Molecular basis of methyl-salicylate-mediated plant airborne defence
Nature (2023)
-
Plant viruses induce plant volatiles that are detected by aphid parasitoids
Scientific Reports (2023)