Abstract
Alkaptonuria (AKU) is an autosomal recessive disorder caused by mutations in homogentisate-1,2-dioxygenase (HGD) gene leading to the deficiency of HGD enzyme activity. The DevelopAKUre project is underway to test nitisinone as a specific treatment to counteract this derangement of the phenylalanine-tyrosine catabolic pathway. We analysed DNA of 40 AKU patients enrolled for SONIA1, the first study in DevelopAKUre, and of 59 other AKU patients sent to our laboratory for molecular diagnostics. We identified 12 novel DNA variants: one was identified in patients from Brazil (c.557T>A), Slovakia (c.500C>T) and France (c.440T>C), three in patients from India (c.469+6T>C, c.650–85A>G, c.158G>A), and six in patients from Italy (c.742A>G, c.614G>A, c.1057A>C, c.752G>A, c.119A>C, c.926G>T). Thus, the total number of potential AKU-causing variants found in 380 patients reported in the HGD mutation database is now 129. Using mCSM and DUET, computational approaches based on the protein 3D structure, the novel missense variants are predicted to affect the activity of the enzyme by three mechanisms: decrease of stability of individual protomers, disruption of protomer-protomer interactions or modification of residues in the region of the active site. We also present an overview of AKU in Italy, where so far about 60 AKU cases are known and DNA analysis has been reported for 34 of them. In this rather small group, 26 different HGD variants affecting function were described, indicating rather high heterogeneity. Twelve of these variants seem to be specific for Italy.
Similar content being viewed by others
Introduction
Alkaptonuria (AKU) [OMIM 203500] is caused by deficiency of homogentisate-1,2-dioxygenase (HGD, EC 1.13.11.5).1 Darkening of the urine upon standing is usually the first sign of AKU, characteristic of homogentisic acid excretion. Homogentisic acid accumulated in the body forms a melanin-like polymer that is deposited in the connective tissues, causing a pathologic pigmentation known as ochronosis. In their late 20s, patients start to suffer severe pain due to degenerative ochronotic arthropathy of intervertebral discs and joints, especially in shoulders, hips and knees.
Since November 2012, the DevelopAKUre project has been underway and is focused on clinical testing of nitisinone as a possible treatment for AKU. Its first part, the SONIA1 study, showed that this drug decreases urine homogentisic acid in a dose-dependent manner.2
The enzymatic defect in AKU is caused by recessive mutations within the HGD gene (HGNC:4892), a single-copy gene that spans 54 363 bp of genomic sequence (3q13.33) and is split into 14 exons and codes for the HGD protomer composed of 445 amino acids.3, 4 The active form of the HGD protein is organised as a hexamer comprising two disc-like trimers.5 An intricate network of non-covalent interactions is required to maintain the spatial structure of the protomer, of the trimer and finally of the hexamer, which can be easily disrupted by variants leading to effects on enzyme function.
AKU has a very low prevalence (1:1 000 000–250 000) in most ethnic groups, but it presents a remarkable allelic heterogeneity—149 different HGD variants have been identified, out of which 116 were reported as mutations and 33 as polymorphisms. All variants are summarised in the HGD mutation database (http://hgddatabase.cvtisr.sk/).6, 7, 8 Some of the reported HGD variants are spread throughout the world, such as c.175delA (p.(Ser59Alafs*52)), one of the first identified AKU mutations, c.899T>G (p.(Val300Gly)).7 On the other hand, there are variants rather specific for some countries or regions; for example, c.342+1G>A (ivs5+1G>A) for Slovakia and the Czech Republic,9 c.87+1G>A (ivs2+1G>A) for the gypsy community Narikuravar in India10 or c.360T>G (p.(Cys120Trp)) for the Dominican Republic.11
So far, about 950 AKU patients have been reported in 61 countries worldwide (AKU Society, www.akusociety.org). The highest number of AKU patients, 208, was reported in Slovakia, including 110 children.12 This country and the Dominican Republic exhibit prevalence of AKU of up to 1:19 000.13, 14 Recently, a high number of AKU cases were also found in Jordan15 and India,10 indicating that the overall prevalence of this disease in some countries might be underestimated.
In this report, we present 12 novel variants identified in thr AKU gene, and as most of them were found in the patients from Italy, we summarize data on AKU in this country.
Materials and methods
Patients
We analysed 99 patients from 20 countries for HGD variants. Forty were enrolled in the SONIA1 clinical study, whereas the remaining 59 were sent to our laboratory for routine DNA diagnostics. Diagnoses of AKU were established based on documented elevated homogentisic acid in urine and/or the bluish-black pigmentation in connective tissue (ochronosis). All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2000. Informed consent was obtained from all patients included in the study.
Variant identification
All 14 HGD exons in each patient were screened for variants by DNA sequencing as previously described.8 Variants were reported according to the Human Genome Variation Society (HGVS) nomenclature additions16 and their description is based on coding DNA Reference Sequence NM_000187.3 (genomic reference sequence NG_011957.1). Exons are numbered like in the study by Granadino et al.4 Variants and other patient data were deposited in the HGD gene mutation database (http://hgddatabase.cvtisr.sk/),8 using a specific family/allele code. Variant nomenclature was verified using MUTALYSER Name Checker (https://www.mutalyzer.nl/). In order to make it easier to recognize variants, in the tables, we also indicate their brief names used in AKU scientific community.
Variant verification
For novel missense variants, the conservation of the affected amino acid position between Homo sapiens and Mus musculus, Rattus norvegicus, Danio rerio, Drosophila melanogaster, Arabidopsis thaliana and Aspergillus nidulans was checked using ClustalW2. Where available, carrier status was tested in parents and other relatives, to confirm segregation of the variant with the disease. PolyPhen-2 (Polymorphism Phenotyping v2)17 and SNAP (Screening for non-acceptable polymorphisms)18 were used to predict the possible effect of amino acid substitutions on the structure and function of the human HGD protein (NP_000178.2). These tools were selected as they use information on 3D structure of the protein and may be more reliable for HGD variants, as was shown before.19
We also used mCSM20 and DUET21 in order to predict the effects of variants on a structural basis. These latter approaches are novel machine-learning algorithms that use the three dimensional structure in order to predict quantitatively the effects of point variants on protein stability and protein-protein and protein-nucleic acid affinities. Two crystal structures of human HGD were used in this analysis (PDB code 1EY2 and 1EYB5). The effect of the variants was assessed in the context of the molecular interactions of the wild-type residue, and mCSM and DUET were used to predict the effects of the variants on protomer and hexamer thermal stability, and mCSM-PPI20 to predict the effects of the variants on the affinity of the protomers to interact with each other. To examine Lys353, a loop missing in the HGD crystal structures (residues 348–355) was modelled and minimised using ModLoop22 and Schrodinger.23
The effect of the splicing variants was predicted using Splice Site Prediction by Neural Network (SSPNN) (http://www.fruitfly.org/seq_tools/splice.html) and Human Splicing Finder (HSF).24
Results
Novel HGD gene variants
Genomic sequencing in 40 SONIA1 AKU patients identified 20 different variants, and variants on both alleles were identified in all of them (Supplementary table 1). Two HGD variants were novel (Table 1): c.158G>A (p.(Arg53Gln)) present in the homozygous state in one patient of Indian origin; and c.500C>T (p.(Tyr167Ile)) identified in one copy in a Slovak AKU patient.
In the remaining 59 AKU cases, 32 different HGD variants affecting protein function were found, including 10 novel (Table 1,Supplementary table 1).
In all 99 patients, all exons were sequenced in order to exclude the presence of other potentially pathogenic variant.
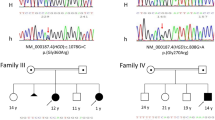
Nine of the tested patients come from Jordan, three of whom were reported before (patients 2,5,8; Supplementary table 1).25 Seven of these cases carried the homozygous variant c.365C>T (p.(Ala122Val)) in exon 6, indicating a founder effect (Figure 1, Supplementary table 1). Two siblings carried also c.16–1G>A (ivs1-1G>A) in the heterozygous state (Figure 1). We did not confirm the presence of variants in exons 11, 13 and 10 published in patient 2, 5, and 8 before.25
c.365C>T (p.(Ala122Val)) founder variant identified in Jordanian families. (a) Chromatogram showing exon 6 of HGD gene sequence. Nucleotide c.365 is indicated by red arrow. Wild-type sequence is in the upper panel, heterozygous carrier in the middle and the patient homozygous for c.365C>T variant in the lower one. (b) Pedigrees of consanguineous Jordanian families AKU_DB_139, AKU_DB_140 and AKU_DB_144 tested for HGD variants. All families are included in the HGD mutation database, under the code indicated. DNA was analysed in individuals where mutation status is indicated. In the family AKU_DB_140, a splicing variant c.16-1G>A in intron 1 was found in heterozygous state. wt indicates a wild-type allele in carriers. The numbers under analysed patients correspond to the patient’s numbers in Supplementary table 1. The full colour version of this figure is available at European Journal of Human Genetics online.
In one patient from Israel, a recently reported genomic deletion of exon 2 (c.16-272_c.87+305del) including flanking intronic sequences was identified (Supplementary table 1).26
In two patients, we were not able to identify any HGD gene variant affecting function, and in two other cases, only one variant was found. All of these cases were of Italian origin (Supplementary table 1).
In summary, including the novel variants, the total number of different HGD function affecting variants reported in the HGD mutation database is now 129 across 380 patients.
Missense variant verification
ClustalW2 showed that all novel missense variants affected conserved amino acid position (data not shown). Moreover, at least one of the prediction programs PolyPhen-2 or SNAP indicated a pathogenic effect of all but one novel amino acid substitutions on the structure and function of the human HGD protein (Table 1). In addition, mCSM and DUET provided a structural understanding behind the inactivation of HGD activity. The variants are spread throughout the structure: three are located along protomer interaction interfaces (Arg53, Tyr40, Gly309), several are buried in the core of the protomers (Phe147, Thr167, Met186, Gly205, Lys248, Gly251) and one lies on a solvent-exposed loop (Lys353). The structural analysis of the identified variants allowed their classification based on the predicted effects into three classes: (i) those that alter the active site, reducing activity; (ii) those that destabilize the protein, reducing activity; and (iii) those that prevent formation of the homohexamer, disrupting activity.
The first class of variants are predicted to affect HGD activity through direct alteration of the active site. The novel variant c.1056A>C (p.(Lys353Gln)) is located on a loop at the mouth of the active site (Supplementary Figure 1). This loop is disordered in the human crystal structure, reflecting its flexibility and possible role in regulating access to the active site. Consistent with its solvent-exposed, flexible nature, the c.1056A>C variant is predicted by mCSM and DUET to have minimal effect on either stability or hexamer formation. However, the loss of the positive charge will alter potential interactions, and it is likely that this variant may alter the flexibility of the loop and consequently catalytic activity. Molecular dynamics simulations using Desmond suggests that the mutant loop indeed has greater flexibility, showing an increase in average RMSDs for the backbone of approximately 20% (data not shown).
The second group of variants affect the production of active HGD enzyme through destabilization of the protomer structure. These variants typically perturb the local secondary structure by the introduction of energetically unfavourable changes, disrupting the interactions made by the wild-type residues (Supplementary Figure 2). Examples of the disruptive nature of these variants include: change of Phe147 to serine would likely disrupt a series of strong intramolecular hydrophobic and pi-pi interactions, strongly destabilising the protomer; the variant c.742A>G (p.(Lys248Glu)) would add an intramolecular ionic repulsive force; mutation of Thr168 to isoleucine would expose a destabilising free main chain NH and would sterically hinder formation of the protomer; and Gly205 and Gly251 are both buried glycines that adopt a positive phi angle in the crystal structures that would usually be energetically unfavourable for most non-glycine amino acids.
The mutations in the third group are at interfaces between protomers and are likely to affect the enzyme activity by lowering stability of the symmetrical homohexameric structure. Both Arg53 and Tyr40 make strong intermolecular interactions between protomers, with Arg53 forming an ionic/hydrogen bond network and Tyr40 making a series of strong hydrophobic and pi-pi interactions (Supplementary Figure 3). Mutation of Arg53 to glutamine, p.(Arg53Gln), was not predicted by mCSM or DUET to affect protomer stability, however, mCSM-PPI predicted the mutation to highly destabilise the formation of the hexamer, because of the loss of the interactions made by the arginine. Mutation of Tyr40 to serine, introducing a polar amino acid residue in the middle of a series of hydrophobic interactions is predicted to be highly destabilising of both protomer and hexamer. Gly309 is sitting along one of the protomer-protomer interfaces with a positive phi angle and mutation to a valine would disrupt the local secondary structure and this interaction.
Splicing variants
We also identified two novel mutations potentially affecting mRNA splicing (Table 1). One is a variant c.469+6T>C in intron 7, identified in the homozygous state in a patient from India. This change is predicted to weaken acceptor splice site by both SSPNN and HSF (data not shown).
The second variant, located 85 nucleotides upstream from the intron 9 acceptor splice site (c.650–85A>G), was found in the homozygous state in a patient from India (Supplementary table 1). No other change was found in this individual by sequencing or by the genomic PCR designed to uncover exon 2 genomic deletion. This substitution was predicted by HSF to potentially activate a cryptic splice site at this location, indicated by improved splice site score (+58.01%, data not shown). This site might compete with the natural intron 9-acceptor splice site. However, no cDNA of the patient was available for analysis in order to prove the splicing effect of this change.
AKU in Italy
So far, about 60 AKU cases are known (aimAKU, www.aimaku.it) and the results of mutation analysis are now reported for 34 families,27, 28 HGD mutation database, present work).
It is remarkable that in 68 AKU chromosomes identified in Italy and published in HGD mutation database, 26 different HGD variants were found, indicating extremely high allelic heterogeneity (Table 2). Twelve of these variants seem to be specific for Italy (Table 2). However, for four Italian patients, only one variant has been identified so far (two in our study: P56 (c.4697+2T>C/?), P46 (c.752G>A/?) (Supplementary table 1) and two reported before (c.650–56G>A/?, c.31_32delGGinsATT/?).29 In other three Italian probands no variant was identified at all (one in our study P22, and two previously reported, including P21 also analysed by us (Supplementary table 1)27).
Discussion
We describe 12 novel variants identified in 99 AKU patients from 20 countries, increasing to 129 the total number of HGD function-affecting variants described so far in 380 patients with this rare disease, as reported in the HGD mutation database (http://hgddatabase.cvtisr.sk/).
The local effect of the novel missense variants was assessed in the context of the molecular interactions of the wild-type residue using mCSM20 and DUET.21 For all variants, disruptive effects were predicted, with change resulting in destabilisation of the protomer, reduced protein-protein affinity or altered activity. With the exception of c.614G>A (p.(Gly205Asp)), a pathogenic effect of amino acid substitutions on the structure and function of the human HGD protein was also predicted by PolyPhen-2 or SNAP.
The effect of potential splicing-affecting variant c.650–85A>G was tested by SSPNN and HSF, and a potential activation of cryptic splice site in intron 9 was predicted. If this novel site is used, 84 bases can be included into HGD transcript between exons 9 and 10, causing insertion of 28 amino acids into HGD protein sequence (c.649_650insGAAAGCCTTTCTCTTCATGCAACCATGGGCATCTTT CCTATGTTTTGGAAGTTTCTAAAAGACTTTTGGGTTACTGTTTTCTAG, p.(Gly217_Ala218ins LysProPheSerSerCysAsnHisGlyHisLeuSerTyrValLeuGluValSerLysArgLeuLeuGlyTyrCysPheLeuGly). The insertion is located on a loop that sits at the interface between two different protomers. There is no room in this region to accommodate a significant change, and so, this mutation would be very disruptive to hexamer formation, as it would interfere and block two separate protomer-protomer interactions. Interestingly, polymorphism c.650–86A>G was described by Vilboux et al.19 Unfortunately, the cDNA of the patient is not available for analysis in order to confirm the splicing effect. We plan to perform MLPA analysis in this patient, in order to test for a presence of possible larger deletions.
AKU generally has low prevalence worldwide, with the notable exceptions of Slovakia30, 31 and the Dominican Republic.13 Although a clear founder effect was observed in the Dominican republic,11 in Slovakia, already 13 different HGD variants affecting function are reported in 121 AKU chromosomes, though most of the patients in this country come from a previously genetically isolated region.7, 8, 9, 32, 33, 34, 35 The most frequent variant found in Slovakia is p.Gly161Arg, present in 45% of all AKU alleles (HGD mutation database).
Recently, several new AKU cases were also reported in southern Jordan,15 in villages characterised by a high level of consanguineous marriages, thus a founder effect is expected. Surprisingly, five different variants were reported in them.15, 25 We re-analysed three patients from this study, however, we were not able to confirm results reported before. None of the variants reported by Al-Sbou25 was found in the DNA samples from the patients. Instead, we show a clear founder effect in Jordan, where most of the patients carried the c.365C>T (p.(Ala122Val)) missense variant in exon 6.
As the majority of novel mutations originate from Italy, we focus on a brief overview on AKU in this country, where about 60 patients have been identified so far. This was possible through early call-for-patients at relevant National Congresses28 and, more recently, with the important contribution of the Italian AKU patient Association (aimAKU). Even though epidemiological estimates specific to Italy are not available, on the basis of the commonly held figures of disease incidence worldwide (1:250 000–1:1 000 000 live births) in a population of about 60 million inhabitants with a birth rate of 9/1000, 0.5–2 new cases per year are expected. As AKU does not generally reduce the lifespan of affected individuals, 40–160 cases are predicted. Therefore, it would seem that the number of ascertained patients fall well within the expected range of AKU prevalence, which means that a good sensitisation work has been made in this country. Including present data, there are 34 Italian families reported in the HGD mutation database and previously published.7, 27, 28, 29, 32, 33 In this rather small group, 26 different AKU-causing mutations were described, indicating high heterogeneity. Moreover, 12 of these mutations seem to be specific for Italy.
In four Italian patients, so far, only one mutation has been identified, and in three patients, mutations have not been discovered at all. Altogether, for 8 AKU patients reported in the database, no mutations were identified, and 22 with only one mutation were found. Whether deep intronic mutations or large deletions encompassing one or more exons are occurring in these cases remains to be investigated. Indeed, recently, the first large deletion of exon 2 including intronic sequences was reported in one case from Lebanon,26 that was also identified in our study in two siblings from Israel. This might indicate that this type of mutations needs to be also considered in other cases where genomic sequencing does not lead to mutation identification.
For a patient to display AKU symptoms, a loss of more than 99% of the enzymatic activity is required.36 So far, observed absence of a clear correlation between genotype and phenotype may be explained by the variability in residual HGD enzymatic activities caused by different mutations.19 Owing to the complex hexameric structure of the HGD enzyme, genotype-phenotype correlation studies are even more complicated when in the patients two different mutations are to be considered. In addition, an effective AKU severity scoring system, AKUSSI, has been developed only recently.37 One of the planned outcomes of the DevelopAKUre project will be detailed clinical characterisations of AKU patients, including mutations, various biochemical parameters as well as AKUSSI. We believe that the tools utilised here for studying the effects of HGD mutations identified in AKU patients provide an important basis for understanding and predicting genotype-phenotype correlations in AKU, the first described inborn error of metabolism.38
References
La Du BN, Zannoni VG, Laster L, Seegmiller JE : The nature of the defect in tyrosine metabolism in alcaptonuria. J Biol Chem 1958; 230: 251–260.
Ranganath LR, Milan AM, Hughes AT et al: Suitability Of Nitisinone In Alkaptonuria 1 (SONIA 1): An international, multicenter, randomized, open-label, no-treatment controlled, parallel-group, dose-response study to investigate the effect of once daily nitisinone on 24-hour urinary homogentisic acid excretion in patients with alkaptonuria after 4 weeks of treatment. Ann Rheum Dis 2014, in press.
Fernández-Cañón JM, Granadino B, Beltrán-Valero de Bernabé D et al: The molecular basis of alkaptonuria. Nat Genet 1996; 14: 19–24.
Granadino B, Beltrán-Valero de Bernabé D, Fernández-Cañón JM, Peñalva MA, Rodríguez de Córdoba S : The human homogentisate 1,2-dioxygenase (HGO) gene. Genomics 1997; 43: 115–122.
Titus GP, Mueller HA, Burgner J, Rodríguez de CS, Peñalva MA, Timm DE : Crystal structure of human homogentisate dioxygenase. Nat Struct Biol 2000; 7: 542–546.
Yang YJ, Guo JH, Chen WJ et al: First report of HGD mutations in a Chinese with alkaptonuria. Gene 2013; 518: 467–469.
Zatkova A : An update on molecular genetics of Alkaptonuria (AKU). J Inherit Metab Dis 2011; 34: 1127–1136.
Zatkova A, Sedlackova T, Radvansky J et al: Identification of 11 novel homogentisate 1,2 dioxygenase variants in alkaptonuria patients and establishment of a novel LOVD-based HGD mutation database. JIMD Rep 2012; 4: 55–65.
Zatkova A, Beltrán-Valero de Bernabé D, Polakova H et al: High frequency of alkaptonuria in Slovakia: evidence for the appearance of multiple mutations in HGO involving different mutational hot spots. Am J Hum Genet 2000; 67: 1333–1339.
Sakthivel S, Zatkova A, Nemethova M, Surovy M, Kadasi L, Saravanan MP : Mutation screening of the HGD gene identifies a novel alkaptonuria mutation with significant founder effect and high prevalence. Ann Hum Genet 2014; 78: 155–164.
Goicoechea De Jorge E, Lorda I, Gallardo ME et al: Alkaptonuria in the Dominican Republic: identification of the founder AKU mutation and further evidence of mutation hot spots in the HGO gene. J Med Genet 2002; 39: E40.
Srsen S, Muller CR, Fregin A, Srsnova K : Alkaptonuria in Slovakia: thirty-two years of research on phenotype and genotype. Mol Genet Metab 2002; 75: 353–359.
Milch RA : Studies of Alcaptonuria: Inheritance of 47 Cases in Eight Highly Inter-related Dominican Kindreds. Am J Hum Genet 1960; 12: 76–85.
Srsen S, Varga F : Screening for alkaptonuria in the newborn in Slovakia. Lancet 1978; 2: 576.
Al-Sbou M, Mwafi N : Nine cases of Alkaptonuria in one family in southern Jordan. Rheumatol Int 2012; 32: 621–625.
den Dunnen JT, Antonarakis SE : Mutation nomenclature extensions and suggestions to describe complex mutations: a discussion. Hum Mutat 2000; 15: 7–12.
Adzhubei IA, Schmidt S, Peshkin L et al: A method and server for predicting damaging missense mutations. Nat Methods 2010; 7: 248–249.
Bromberg Y, Rost B : SNAP: predict effect of non-synonymous polymorphisms on function. Nucleic Acids Res 2007; 35: 3823–3835.
Vilboux T, Kayser M, Introne W et al: Mutation spectrum of homogentisic acid oxidase (HGD) in alkaptonuria. Hum Mutat 2009; 30: 1611–1619.
Pires DE, Ascher DB, Blundell TL : mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 2014; 30: 335–342.
Pires DE, Ascher DB, Blundell TL : DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res 2014; W314–W319.
Fiser A, Sali A : ModLoop: automated modeling of loops in protein structures. Bioinformatics 2003; 19: 2500–2501.
Jacobson MP, Pincus DL, Rapp CS et al: A hierarchical approach to all-atom protein loop prediction. Proteins 2004; 55: 351–367.
Desmet FO, Hamroun D, Lalande M, Collod-Beroud G, Claustres M, Beroud C : Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res 2009; 37: e67.
Al-sbou M : Novel mutations in the homogentisate 1,2 dioxygenase gene identified in Jordanian patients with alkaptonuria. Rheumatol Int 2012; 32: 1741–1746.
Habbal MZ, Bou-Assi T, Zhu J, Owen R, Chehab FF : First report of a deletion encompassing an entire exon in the homogentisate 1,2-dioxygenase gene causing alkaptonuria. PLoS One 2014; 9: e106948.
Mannoni A, Selvi E, Lorenzini S et al: Alkaptonuria, ochronosis, and ochronotic arthropathy. Semin Arthritis Rheum 2004; 33: 239–248.
Porfirio B, Chiarelli I, Graziano C et al: Alkaptonuria in Italy: polymorphic haplotype background, mutational profile, and description of four novel mutations in the homogentisate 1,2-dioxygenase gene. J Med Genet 2000; 37: 309–312.
Beltrán-Valero de Bernabé D, Granadino B, Chiarelli I et al: Mutation and polymorphism analysis of the human homogentisate 1, 2-dioxygenase gene in alkaptonuria patients. Am J Hum Genet 1998; 62: 776–784.
Srsen S, Koska L, Kapralik I : [Alkaptonuria in the Upper Hron Region in Slovakia (author's transl)]. Cas Lek Cesk 1978; 117: 1517–1522.
Srsen S : Dark pigmentation of ear cerumen in alkaptonuria. Lancet 1978; 2: 577.
Gehrig A, Schmidt SR, Muller CR, Srsen S, Srsnova K, Kress W : Molecular defects in alkaptonuria. Cytogenet Cell Genet 1997; 76: 14–16.
Muller CR, Fregin A, Srsen S et al: Allelic heterogeneity of alkaptonuria in Central Europe. Eur J Hum Genet 1999; 7: 645–651.
Zatkova A, Chmelikova A, Polakova H, Ferakova E, Kadasi L : Rapid detection methods for five HGO gene mutations causing alkaptonuria. Clin Genet 2003; 63: 145–149.
Zatkova A, Polakova H, Micutkova L et al: Novel mutations in the homogentisate-1,2-dioxygenase gene identified in Slovak patients with alkaptonuria. J Med Genet 2000; 37: 539–542.
Rodríguez JM, Timm DE, Titus GP et al: Structural and functional analysis of mutations in alkaptonuria. Hum Mol Genet 2000; 9: 2341–2350.
Cox TF, Ranganath LR : A quantitative assessment of alkaptonuria: testing the reliability of two disease severity scoring systems. J Inherit Metab Dis 2011; 34: 1153–1162.
Garrod AE : A Contribution to the Study of Alkaptonuria. Med Chir Trans 1899; 82: 367–394.
Acknowledgements
DNA analysis of SONIA1 samples was supported by the European Commission Seventh Framework Programme funding granted in 2012 (DevelopAKUre, project number: 304985). Analysis of remaining patients was performed thanks to the Research & Developmental Operational Programme funded by the ERDF (the project ‘Creating a Competitive Centre for research and development in the field of molecular medicine’ ITMS 26240220071), and the Slovak National Agency VEGA (grant No. 2/0027/12). DBA is supported by a NHMRC CJ Martin Fellowship (GNT1072476). DEVP is supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Nemethova, M., Radvanszky, J., Kadasi, L. et al. Twelve novel HGD gene variants identified in 99 alkaptonuria patients: focus on ‘black bone disease’ in Italy. Eur J Hum Genet 24, 66–72 (2016). https://doi.org/10.1038/ejhg.2015.60
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.60