Abstract
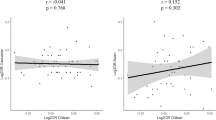
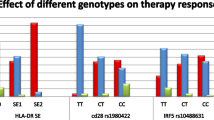
Notwithstanding the well-established association of HLA–DRB1 shared epitope alleles, interest remains in identifying additional major histocompatibility complex (MHC) region variants associated with rheumatoid arthritis (RA). We used a panel of 1201 haplotype-tagging single nucleotide polymorphisms (SNPs) designed for African Americans to find genetic variants associated with RA in a 3.8-Mb region encompassing the MHC. Conditioning on seven covariates, including HLA–DRB1 risk alleles and population structure, we identified an SNP in HLA–DOA (rs9276977) significantly associated with RA; minor allele frequency (MAF) 0.27 in cases versus 0.21 in controls, odds ratio (±95% confidence interval)=2.86 (1.61, 5.31). Genotyping of rs9276977 in an independent sample of African-American RA patients and controls did not replicate the association (MAF 0.28 in cases versus 0.27 in controls). This study points to the potential association of a SNP in the HLA–DOA gene with RA in African Americans, but also underscores the importance of replication of findings in larger patient cohorts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Gregersen PK, Silver J, Winchester RJ . The shared epitope hypothesis: an approach to understanding the molecular genetics of susceptibility to rheumatoid arthritis. Arthritis Rheum 1987; 30: 1205–1213.
Ding B, Padyukov L, Lundstrom E, Seielstad M, Plenge RM, Oksenberg JR et al. Different patterns of associations with anti-citrullinated protein antibody-positive and anti-citrullinated protein antibody-negative rheumatoid arthritis in the extended major histocompatibility complex region. Arthritis Rheum 2009; 60: 30–38.
Vignal C, Bansal AT, Balding DJ, Binks MH, Dickson MC, Montgomery DS et al. Genetic association of the major histocompatibility complex with rheumatoid arthritis implicates two non-DRB1 loci. Arthritis Rheum 2009; 60: 53–62.
Hughes L, Morrison D, Kelley JM, Padilla MA, Vaughn LK, Westfall AO et al. The HLA-DRB1 shared epitope is associated with susceptibility to rheumatoid arthritis in African Americans through European genetic admixture. Arthritis Rheum 2008; 58: 349–358.
Kelley JM, Hughes LB, Feng R, Liu N, Padilla MA, Vaughan LK et al. Evaluating linkage disequilibrium and recombination provides a haplotype-tagging SNP panel of the major histocompatibility complex in African Americans. Genes Immun 2008; 9: 271–273.
R Development Core Team. R: a Language and Environment for Statistical Computing 2008. http://www.R-project.org.
O'Gorman TW . The performance of randomization tests that use permutations of independent variables. Commun Stat Simulat 2005; 34: 895–908.
van Lith M, van Ham M, Neefjes J . Novel polymorphisms in HLA-DOA and HLA-DOB in B-cell malignancies. Immunogenetics 2008; 54: 591–595.
Denzin LK, Fallas JL, Prendes M, Yi W . Right place, right time, right peptide: DO keeps DM focused. Immunol Rev 2005; 207: 279–292.
Mikuls TR, Holers VM, Parrish L, Kuhn KA, Conn DL, Gilkeson G et al. Anti-cyclic citrullinated peptide antibody and rheumatoid factor isotypes in African Americans with early rheumatoid arthritis. Arthritis Rheum 2006; 54: 3057–3059.
Balding DJ . A tutorial on statistical methods for population association studies. Nat Rev Gen 2006; 7: 781–791.
Gao X, Starmer J, Martin ER . A multiple testing correction method for genetic association studies using correlated single nucleotide polymorphisms. Genet Epidemiol 2008; 32: 361–369.
Acknowledgements
The CLEAR Investigators are as follows: Graciela S Alarcón, MD, MPH and George Howard, DrPH from the University of Alabama at Birmingham, Birmingham, AL; Doyt L Conn, MD from the Emory University, Atlanta, GA; Beth L Jonas, MD and Leigh F Callahan, PhD from the University of North Carolina, Chapel Hill, NC; Edwin A Smith, MD from the Medical University of South Carolina, Charleston, SC; Richard D Brasington Jr, MD from Washington University, St Louis, MO; and Larry W Moreland, MD, University of Pittsburgh. We gratefully acknowledge the following physicians who enrolled patients: Jacob Aelion, MD, Jackson, TN; Charles Bell, Birmingham, AL; Sohrab Fallahi, MD, Montgomery, AL; Richard Jones, PhD, MD, Tuscaloosa, AL; Maura Kennedy, MD, Birmingham, AL; Adahli Estrada Massey, MD, Auburn, AL; John Morgan, MD, Birmingham, AL; Donna Paul, MD, Montgomery, AL; Runas Powers, MD, Alexander City, AL; William Shergy, MD, Huntsville, AL; Cornelius Thomas, MD, Birmingham, AL; Ben Wang, MD, Memphis, TN. We gratefully acknowledge the staff and coordinators at the following sites: University of Alabama at Birmingham: Stephanie Ledbetter, MS; Zenoria Causey, MS; Selena Luckett, RN, CRNC; Laticia Woodruff, RN, MSN; Candice Miller; Emory University: Joyce Carlone, RN, FNP-BC; Karla Caylor, BSN, RN; Sharon Henderson, RN; University of North Carolina: Diane Bresch, BSN; Medical University of South Carolina: Trisha Sturgill. We gratefully acknowledge Nicholas Pajewski, Kelly Vaughan and Miguel Padilla for discussions of the data analysis. The CLEAR Registry is a national resource, with clinical data, DNA and other biological samples available. For details, see the following website: http://www.dom.uab.edu/rheum/CLEAR%20home.htm. This work was supported by NIH N01 AI40068, N01 AR02247, P01 AR40894, R01 AR51394, and NIH GCRC/CTSA Grants M01 RR00032, U54 RR025777, R01 GM069430, T32 HL072757 (University of Alabama at Birmingham) and M01 RR 000046 (University of North Carolina).
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
About this article
Cite this article
Reynolds, R., Kelley, J., Hughes, L. et al. Genetic association of htSNPs across the major histocompatibility complex with rheumatoid arthritis in an African-American population. Genes Immun 11, 94–97 (2010). https://doi.org/10.1038/gene.2009.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2009.69
Keywords
This article is cited by
-
What to do with HLA-DO/H-2O two decades later?
Immunogenetics (2019)