Abstract
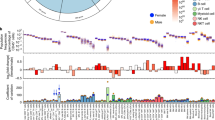
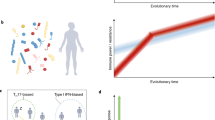
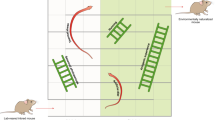
The Collaborative Cross (CC) is an emerging panel of recombinant inbred (RI) mouse strains. Each strain is genetically distinct but all descended from the same eight inbred founders. In 66 strains from incipient lines of the CC (pre-CC), as well as the 8 CC founders and some of their F1 offspring, we examined subsets of lymphocytes and antigen-presenting cells. We found significant variation among the founders, with even greater diversity in the pre-CC. Genome-wide association using inferred haplotypes detected highly significant loci controlling B-to-T cell ratio, CD8 T-cell numbers, CD11c and CD23 expression. Comparison of overall strain effects in the CC founders with strain effects at QTL in the pre-CC revealed sharp contrasts in the genetic architecture of two traits with significant loci: variation in CD23 can be explained largely by additive genetics at one locus, whereas variation in B-to-T ratio has a more complex etiology. For CD23, we found a strong QTL whose confidence interval contained the CD23 structural gene Fcer2a. Our data on the pre-CC demonstrate the utility of the CC for studying immunophenotypes and the value of integrating founder, CC and F1 data. The extreme immunophenotypes observed could have pleiotropic effects in other CC experiments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Conley ME . Genetics of primary immunodeficiency diseases. Rev Immunogenet 2000; 2: 231–242.
Ober C, Yao TC . The genetics of asthma and allergic disease: a 21st century perspective. Immunol Rev 2011; 242: 10–30.
Todd JA . Etiology of type 1 diabetes. Immunity 2010; 32: 457–467.
Flint J, Eskin E . Genome-wide association studies in mice. Nat Rev Genet 2012; 13: 807–817.
Collaborative Cross Consortium. The genome architecture of the Collaborative Cross mouse genetic reference population. Genetics 2012; 190: 389–401.
Churchill GA, Airey DC, Allayee H, Angel JM, Attie AD, Beatty J et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat Genet 2004; 36: 1133–1137.
Threadgill DW, Hunter KW, Williams RW . Genetic dissection of complex and quantitative traits: from fantasy to reality via a community effort. Mamm Genome 2002; 13: 175–178.
Broman KW . The genomes of recombinant inbred lines. Genetics 2005; 169: 1133–1146.
Roberts A, Pardo-Manuel de Villena F, Wang W, McMillan L, Threadgill DW . The polymorphism architecture of mouse genetic resources elucidated using genome-wide resequencing data: implications for QTL discovery and systems genetics. Mamm Genome 2007; 18: 473–481.
Chesler EJ, Miller DR, Branstetter LR, Galloway LD, Jackson BL, Philip VM et al. The Collaborative Cross at Oak Ridge National Laboratory: developing a powerful resource for systems genetics. Mamm Genome 2008; 19: 382–389.
Iraqi FA, Churchill G, Mott R . The Collaborative Cross, developing a resource for mammalian systems genetics: a status report of the Wellcome Trust cohort. Mamm Genome 2008; 19: 379–381.
Morahan G, Balmer L, Monley D . Establishment of ‘The Gene Mine’: a resource for rapid identification of complex trait genes. Mamm Genome 2008; 19: 390–393.
Aylor DL, Valdar W, Foulds-Mathes W, Buus RJ, Verdugo RA, Baric RS et al. Genetic analysis of complex traits in the emerging Collaborative Cross. Genome Res 2011; 21: 1213–1222.
Welsh CE, Miller DR, Manly KF, Wang J, McMillan L, Morahan G et al. Status and access to the Collaborative Cross population. Mamm Genome 2012; 23: 706–712.
Kelada SN, Aylor DL, Peck BC, Ryan JF, Tavarez U, Buus RJ et al. Genetic analysis of hematological parameters in incipient lines of the collaborative cross. G3 (Bethesda) 2012; 2: 157–165.
Ferris MT, Aylor DL, Bottomly D, Whitmore AC, Aicher LD, Bell TA et al. Modeling host genetic regulation of influenza pathogenesis in the collaborative cross. PLoS Pathog 2013; 9: e1003196.
Ford JW, Sturgill JL, Conrad DH . 129/SvJ mice have mutated CD23 and hyper IgE. Cell Immunol 2009; 254: 124–134.
Becker-Herman S, Meyer-Bahlburg A, Schwartz MA, Jackson SW, Hudkins KL, Liu C et al. WASp-deficient B cells play a critical, cell-intrinsic role in triggering autoimmunity. J Exp Med 2011; 208: 2033–2042.
Bouma G, Burns SO, Thrasher AJ . Wiskott-Aldrich Syndrome: immunodeficiency resulting from defective cell migration and impaired immunostimulatory activation. Immunobiology 2009; 214: 778–790.
Kliche S, Worbs T, Wang X, Degen J, Patzak I, Meineke B et al. CCR7-mediated LFA-1 functions in T cells are regulated by 2 independent ADAP/SKAP55 modules. Blood 2012; 119: 777–785.
Lenarcic AB, Svenson KL, Churchill GA, Valdar W . A general Bayesian approach to analyzing diallel crosses of inbred strains. Genetics 2012; 190: 413–435.
Lewis G, Rapsomaniki E, Bouriez T, Crockford T, Ferry H, Rigby R et al. Hyper IgE in New Zealand black mice due to a dominant-negative CD23 mutation. Immunogenetics 2004; 56: 564–571.
Keane TM, Goodstadt L, Danecek P, White MA, Wong K, Yalcin B et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 2011; 477: 289–294.
Yalcin B, Flint J, Mott R . Using progenitor strain information to identify quantitative trait nucleotides in outbred mice. Genetics 2005; 171: 673–681.
Valdar WSJ . Scoring residue conservation. Proteins 2002; 48: 227–241.
Binkley J, Karra K, Kirby A, Hosobuchi M, Stone EA, Sidow A . ProPhylER: a curated online resource for protein function and structure based on evolutionary constraint analyses. Genome Res 2010; 20: 142–154.
Svenson KL, Gatti DM, Valdar W, Welsh CE, Cheng R, Chesler EJ et al. High-resolution genetic mapping using the Mouse Diversity outbred population. Genetics 2012; 190: 437–447.
Valdar W, Solberg LC, Gauguier D, Burnett S, Klenerman P, Cookson WO et al. Genome-wide genetic association of complex traits in heterogeneous stock mice. Nat Genet 2006; 38: 879–887.
Belknap JK . Effect of within-strain sample size on QTL detection and mapping using recombinant inbred mouse strains. Behav Genet 1998; 28: 29–38.
Durrant C, Tayem H, Yalcin B, Cleak J, Goodstadt L, de Villena FP et al. Collaborative Cross mice and their power to map host susceptibility to Aspergillus fumigatus infection. Genome Res 2011; 21: 1239–1248.
Yalcin B, Nicod J, Bhomra A, Davidson S, Cleak J, Farinelli L et al. Commercially available outbred mice for genome-wide association studies. PLoS Genet 2010; 6: 9.
Koarada S, Wu Y, Yim YS, Wakeland EW, Ridgway WM . Nonobese diabetic CD4 lymphocytosis maps outside the MHC locus on chromosome 17. Immunogenetics 2004; 56: 333–337.
Hsu J . Multiple Comparisons: Theory and Methods. Chapman and Hall/CRC, 1996.
Team RDA . Language and Environment for Statistical Computing In R Foundation of Statistical Computing: Vienna, Austria, 2012.
Pinheiro J, Bates D . Mixed-Effects Models in S and S-PLUS. Springer, 2009.
Hothorn T, Bretz F, Westfall P . Simultaneous inference in general parametric models. Biom J 2008; 50: 346–363.
Lynch M, Walsh B . Genetics and Analysis of Quantitative Traits. Sinauer: Sunderland, Mass., 1998.
Yang H, Ding Y, Hutchins LN, Szatkiewicz J, Bell TA, Paigen BJ et al. A customized and versatile high-density genotyping array for the mouse. Nat Methods 2009; 6: 663–666.
Mott R, Talbot CJ, Turri MG, Collins AC, Flint J . A method for fine mapping quantitative trait loci in outbred animal stocks. Proc Natl Acad Sci USA 2000; 97: 12649–12654.
Valdar W, Holmes CC, Mott R, Flint J . Mapping in structured populations by resample model averaging. Genetics 2009; 182: 1263–1277.
Johnsen AK, Valdar W, Golden L, Ortiz-Lopez A, Hitzemann R, Flint J et al. Genome-wide and species-wide dissection of the genetics of arthritis severity in heterogeneous stock mice. Arthritis Rheum 2011; 63: 2630–2640.
Robinson G . That BLUP is a good thing: the estimation of random effects. Statistical Science 1991; 6: 15–51.
Solberg Woods LC, Holl KL, Oreper D, Xie Y, Tsaih SW, Valdar W . Fine-mapping diabetes-related traits, including insulin resistance, in heterogeneous stock rats. Physiol Genomics 2012; 44: 1013–1026.
Dupuis J, Siegmund D . Statistical methods for mapping quantitative trait loci from a dense set of markers. Genetics 1999; 151: 373–386.
Solberg Woods LC, Holl K, Tschannen M, Valdar W . Fine-mapping a locus for glucose tolerance using heterogeneous stock rats. Physiol Genomics 2010; 41: 102–108.
Visscher PM, Thompson R, Haley CS . Confidence intervals in QTL mapping by bootstrapping. Genetics 1996; 143: 1013–1020.
Schwarz G . Estimating Dimension of a Model. Annals of Statistics 1978; 6: 461–464.
Acknowledgements
The authors thank Dr Ellen Young, Cindy Hensley and Shaun Steele for their substantial assistance in data collection. The authors acknowledge NIH grants GM070683 (genotyping), P50 MH0903380/HG006582 (partial support for AB, FPMV, WV), GM104125 (partial support for AB, WV), F32 GM090667 (DLA), CA105417 (DWT, FVPM), and the Lineberger Comprehensive Cancer Center (JAF).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Rights and permissions
About this article
Cite this article
Phillippi, J., Xie, Y., Miller, D. et al. Using the emerging Collaborative Cross to probe the immune system. Genes Immun 15, 38–46 (2014). https://doi.org/10.1038/gene.2013.59
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2013.59
Keywords
This article is cited by
-
Onchocerca volvulus bivalent subunit vaccine induces protective immunity in genetically diverse collaborative cross recombinant inbred intercross mice
npj Vaccines (2021)
-
Genomic, microbial and environmental standardization in animal experimentation limiting immunological discovery
BMC Immunology (2020)
-
A comprehensive and comparative phenotypic analysis of the collaborative founder strains identifies new and known phenotypes
Mammalian Genome (2020)
-
Variable outcomes of human heart attack recapitulated in genetically diverse mice
npj Regenerative Medicine (2019)
-
The Collaborative Cross mouse model for dissecting genetic susceptibility to infectious diseases
Mammalian Genome (2018)