Abstract
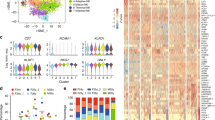
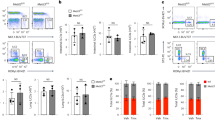
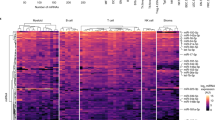
Natural killer (NK) and T lymphocytes share many properties, yet only NK cells respond rapidly to infection and cancer without pre-activation. We found that few microRNAs (miRNAs) differed significantly between human NK and T cells. Among those miRNAs, miR-181a and miR-181b levels rose during NK cell differentiation. Prior studies indicate that miR-181a and miR-181b are critical for human NK cell development and are co-transcribed from genes on chromosome 1 (MIR181A1B1) and on chromosome 9 (MIR181A2B2). We mapped human MIR181A1B1 and MIR181A2B2 transcription start sites to 78.3 kb and 34.0 kb upstream of the mature miRNAs, generating predominantly unspliced transcripts of 80–127 kb and ~60 kb, respectively. Unlike mouse thymocytes, human T cells expressed both MIR181A1B1 and MIR181A2B2. We tested the hypothesis that NK cells differentially transcribe the two genes during development and in response to immune regulatory cytokines. During NK-cell differentiation, MIR181A2B2 expression rose markedly and exceeded that of MIR181A1B1. TGF-β treatment increased NK-cell MIR181A2B2 transcription, whereas IL-2, IL-15 and IL-12/IL-18 treatments upregulated MIR181A1B1. The MIR181A2B2 promoter was strongly transactivated by SMAD3 and SMAD4 transcription factors, suggesting that TGF-β signaling upregulates MIR181A2B2 expression, at least in part, through SMAD-dependent promoter activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Caligiuri MA . Human natural killer cells. Blood 2008; 112: 461–469.
Vivier E, Raulet DH, Moretta A, Caligiuri MA, Zitvogel L, Lanier LL et al. Innate or adaptive immunity? The example of natural killer cells. Science 2011; 331: 44–49.
Fauriat C, Long EO, Ljunggren H-G, Bryceson YT . Regulation of human NK-cell cytokine and chemokine production by target cell recognition. Blood 2010; 115: 2167–2176.
Yu J, Mao HC, Wei M, Hughes T, Zhang J, Park IK et al. CD94 surface density identifies a functional intermediary between the CD56bright and CD56dim human NK-cell subsets. Blood 2010; 115: 274–281.
Xiao C, Rajewsky K . MicroRNA control in the immune system: basic principles. Cell 2009; 136: 26–36.
Kim VN . MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 2005; 6: 376–385.
Wang P, Gu Y, Zhang Q, Han Y, Hou J, Lin L et al. Identification of resting and type I IFN-activated human NK cell miRNomes reveals microRNA-378 and microRNA-30e as negative regulators of NK cell cytotoxicity. J Immunol 2012; 189: 211–221.
Liu X, Wang Y, Sun Q, Yan J, Huang J, Zhu S et al. Identification of microRNA transcriptome involved in human natural killer cell activation. Immunol Lett 2012; 143: 208–217.
Obata-Onai A, Hashimoto S, Onai N, Kurachi M, Nagai S, Shizuno K et al. Comprehensive gene expression analysis of human NK cells and CD8(+) T lymphocytes. Int Immunol 2002; 14: 1085–1098.
Blom B, Spits H . Development of human lymphoid cells. Annu Rev Immunol 2006; 24: 287–320.
Bezman NA, Kim CC, Sun JC, Min-Oo G, Hendricks DW, Kamimura Y et al. Molecular definition of the identity and activation of natural killer cells. Nat Immunol 2012; 13: 1000–1009.
Bezman NA, Cedars E, Steiner DF, Blelloch R, Hesslein DGT, Lanier LL . Distinct requirements of microRNAs in NK cell activation, survival, and function. J Immunol 2010; 185: 3835–3846.
Sullivan RP, Leong JW, Schneider SE, Keppel CR, Germino E, French AR et al. MicroRNA-deficient NK cells exhibit decreased survival but enhanced function. J Immunol 2012; 188: 3019–3030.
Li Q-J, Chau J, Ebert PJR, Sylvester G, Min H, Liu G et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 2007; 129: 147–161.
Cichocki F, Felices M, McCullar V, Presnell SR, Al-Attar A, Lutz CT et al. Cutting Edge: MicroRNA-181 promotes human NK cell development by regulating Notch signaling. J Immunol 2011; 187: 6171–6175.
Okamura K, Phillips MD, Tyler DM, Duan H, Chou YT, Lai EC . The regulatory activity of microRNA* species has substantial influence on microRNA and 3′ UTR evolution. Nat Struct Mol Biol 2008; 15: 354–363.
Chiang HR, Schoenfeld LW, Ruby JG, Auyeung VC, Spies N, Baek D et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev 2010; 24: 992–1009.
Liu G, Min H, Yue S, Chen CZ . Pre-miRNA loop nucleotides control the distinct activities of mir-181a-1 and mir-181c in early T cell development. PLoS ONE 2008; 3: e3592.
Chen CZ . An unsolved mystery: the target-recognizing RNA species of microRNA genes. Biochimie 2013; 95: 1663–1676.
Fragoso R, Mao T, Wang S, Schaffert S, Gong X, Yue S et al. Modulating the strength and threshold of NOTCH oncogenic signals by mir-181a-1/b-1. PLoS Genet 2012; 8: e1002855.
Freud AG, Yokohama A, Becknell B, Lee MT, Mao HC, Ferketich AK et al. Evidence for discrete stages of human natural killer cell differentiation in vivo. J Exp Med 2006; 203: 1033–1043.
Slezak-Prochazka I, Kluiver J, de Jong D, Kortman G, Halsema N, Poppema S et al. Cellular localization and processing of primary transcripts of exonic microRNAs. PLoS ONE 2013; 8: e76647.
Lan ZJ, Xu X, Chung AC, Cooney AJ . Extra-germ cell expression of mouse nuclear receptor subfamily 6, group A, member 1 (NR6A1). Biol Reprod 2009; 80: 905–912.
Giresi PG, Kim J, McDaniell RM, Iyer VR, Lieb JD . FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res 2007; 17: 877–885.
Ernst J, Kellis M . ChromHMM: automating chromatin-state discovery and characterization. Nat Methods 2012; 9: 215–216.
Shin KH, Bae SD, Hong HS, Kim RH, Kang MK, Park NH . miR-181a shows tumor suppressive effect against oral squamous cell carcinoma cells by downregulating K-ras. Biochem Biophys Res Commun 2011; 404: 896–902.
Lo K, Smale ST . Generality of a functional initiator consensus sequence. Gene 1996; 182: 13–22.
Liang R, Bates DJ, Wang E . Epigenetic control of microRNA expression and aging. Curr Genomics 2009; 10: 184–193.
Massague J . TGFβ signalling in context. Nat Rev Mol Cell Biol 2012; 13: 616–630.
Chen C-Z, Li L, Lodish HF, Bartel DP . MicroRNAs modulate hematopoietic lineage differentiation. Science 2004; 303: 83–86.
Bachanova V, McCullar V, Lenvik T, Wangen R, Peterson KA, Ankarlo DE et al. Activated notch supports development of cytokine producing NK cells which are hyporesponsive and fail to acquire NK cell effector functions. Biol Blood Marrow Transplant 2009; 15: 183–194.
Neel JC, Lebrun JJ . Activin and TGFβ regulate expression of the microRNA-181 family to promote cell migration and invasion in breast cancer cells. Cell Signal 2013; 25: 1556–1566.
Taylor MA, Sossey-Alaoui K, Thompson CL, Danielpour D, Schiemann WP . TGF-beta upregulates miR-181a expression to promote breast cancer metastasis. J Clin Invest 2013; 123: 150–163.
Kirigin FF, Lindstedt K, Sellars M, Ciofani M, Low SL, Jones L et al. Dynamic microRNA gene transcription and processing during T cell development. J Immunol 2012; 188: 3257–3267.
Hickey CJ, Schwind S, Radomska HS, Dorrance AM, Santhanam R, Mishra A et al. Lenalidomide-mediated enhanced translation of C/EBPα-p30 protein up-regulates expression of the antileukemic microRNA-181a in acute myeloid leukemia. Blood 2013; 121: 159–169.
Marsico A, Huska MR, Lasserre J, Hu H, Vucicevic D, Musahl A et al. PROmiRNA: a new miRNA promoter recognition method uncovers the complex regulation of intronic miRNAs. Genome Biol 2013; 14: R84.
Barski A, Jothi R, Cuddapah S, Cui K, Roh TY, Schones DE et al. Chromatin poises miRNA- and protein-coding genes for expression. Genome Res 2009; 19: 1742–1751.
Bert AG, Burrows J, Osborne CS, Cockerill PN . Generation of an improved luciferase reporter gene plasmid that employs a novel mechanism for high-copy replication. Plasmid 2000; 44: 173–182.
Presnell SR, Zhang L, Ramilo CA, Chan H-W, Lutz CT . Functional redundancy of transcription factor-binding sites in the killer cell Ig-like receptor (KIR) gene promoter. Int Immunol 2006; 18: 1221–1232.
Presnell SR, Zhang L, Chlebowy CN, Al-Attar A, Lutz CT . Differential transcription factor use by the KIR2DL4 promoter under constitutive and IL-2/15-treated conditions. J Immunol 2012; 188: 4394–4404.
Acknowledgements
We thank Jeffrey Ebersole, Yelena Alimova, Peter Nelson and Luke Bradley for use of equipment, Martha L Peterson, Peter T Nelson, Wangxia Wang and Francesc Marti for advice, Brett Spear and Francesc Marti for cell lines, Teresa Woodruff for plasmids, Dennis Williams and the Kentucky Blood Center for help with blood products and the National Cancer Institute for IL-2. This work was supported by grants from the National Institutes of Health R01 AI56506 to CTL, R01 55417 to JSM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Rights and permissions
About this article
Cite this article
Presnell, S., Al-Attar, A., Cichocki, F. et al. Human natural killer cell microRNA: differential expression of MIR181A1B1 and MIR181A2B2 genes encoding identical mature microRNAs. Genes Immun 16, 89–98 (2015). https://doi.org/10.1038/gene.2014.65
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2014.65
This article is cited by
-
Restoration of natural killer cell cytotoxicity in the suppressive tumor microenvironment: novel approaches to treat AML
Journal of Hematopathology (2017)