Abstract
Purpose: To evaluate the yield of genetic testing in children with autism spectrum disorders.
Methods: We performed a retrospective chart review of 71 unrelated patients with a diagnosis of an isolated autism spectrum disorder seen in a genetics clinic over a period of 14 months. For most, referrals occurred after evaluation by a developmental pediatrician and/or psychologist to establish the diagnosis. Tiered laboratory testing for the majority of the patients followed a guideline that was developed in collaboration with clinicians at The Autism Center at Children's Hospital, Columbus, OH.
Results: The patients included 57 males and 14 females; 57 met DSM-IV criteria for autism, with the rest being Asperger or pervasive developmental disorder not otherwise specified. Macrocephaly [head circumference (HC) ≥95%] was present in 19 (27%). Two children had visible chromosome abnormalities (47,XYY; 48,XY + 2mar/49,XY + 3mar). Two patients with autism and macrocephaly had heterozygous mutations in the PTEN tumor suppressor gene. Three females had Rett syndrome, each confirmed by DNA sequencing of the MECP2 gene. Extensive metabolic testing produced no positive results, nor did fragile X DNA testing.
Conclusion: The overall diagnostic yield was 10% (7/71). PTEN gene sequencing should be considered in any child with macrocephaly and autism or developmental delay. Metabolic screening may not be warranted in autism spectrum disorders without more specific indications or additional findings.
Similar content being viewed by others
Main
Autism spectrum disorders (ASDs) are a common and heterogeneous group of developmental disorders. Affected infants and children demonstrate persistent impairments in expressive and receptive language and social interaction skills that are evident before age 3, as well as a restricted range of interests and stereotypic behaviors. The prevalence of ASDs [including autism, Asperger syndrome, and pervasive developmental delay not otherwise specified (PDD-NOS)] has recently been estimated by the Centers for Disease Control and Prevention as approximately 5.6 per 1000 children in the United States.1 The increased frequency of diagnosis may be due, at least in part, to broader diagnostic criteria since 1994 and the reporting of autism as a disability since 1992.
Although the causes for most cases of ASDs are not currently known, there is an overwhelming number of scientific studies that demonstrate a strong genetic component to the etiologies of autism.1–5 There is familial aggregation, exemplified by a sibling recurrence rate of 2% to 8% and relative recurrence risk of 3 to 15 times the rate in the general population. Twin studies demonstrate a concordance rate in monozygotic twins of 60% to 90% and in dizygotic twins of 0 to 10%. There is also an increased predilection in males, with a male-to-female ratio of 3–4:1. There are several single gene disorders, such as fragile X syndrome, Rett syndrome, untreated phenylketonuria, and tuberous sclerosis, in which autistic features are prominent and common.6 ASDs are also associated with several chromosome abnormalities, the most common of which is a duplication of 15q11-q13 that includes the region involved in Prader-Willi and Angelman syndromes.7 However, known genetic causes account for 10% or less of ASD cases. It is estimated that 10 to 100 genes, in addition to environmental and, possibly, epigenetic effects, may contribute to disease susceptibility.4,5,8–10
There are several ongoing large collaborative studies in the United States and other countries with goals to identify autism susceptibility genes.11–14 Such research has been hampered, in part, by a lack of comprehensive clinical phenotyping and performing appropriate and complete genetic testing of affected individuals to rule out other known causes. Recently, there have been several publications15,16 as well as active discussion among clinical geneticists, neurologists, and developmental pediatric specialists about the appropriate genetic diagnostic evaluation of children with a suspected or proven ASD, particularly as new genetic etiologies are discovered. In early 2005, we developed a clinical guideline for laboratory testing and referral to a genetics clinic for children diagnosed locally with an ASD. We report here the results of a retrospective chart review of all patients with an ASD seen in genetics clinic within a 14-month period after initiation of this protocol.
METHODS
After approval by our hospital Human Subjects Institutional Review Board, a retrospective chart review was performed by one of the authors (G.E.H.) of all patients seen in the genetics clinic at Columbus Children's Hospital between January 1, 2005 and March 7, 2006 with a diagnosis of an ASD. Specifically, after their evaluation of the child, the clinical geneticist listed autism, ASD, Asperger syndrome, PDD-NOS, or Rett syndrome as the primary or one of two secondary diagnoses in our clinical database. A total of 79 patients were ascertained, including two sets of male siblings. Three patients were excluded because they did not have evidence supporting the diagnosis of an ASD on review of the chart. Two additional patients were excluded because they had malformations suggesting the presence of an unknown syndrome in addition to mental retardation and autistic features. Specifically, one girl, age 8, had seizures, mental retardation, and a hypothalamic hamartoma and was thought to have Pallister-Hall syndrome. A 14-year-old boy had autistic features, congenital deafness, cleft palate, and facies resembling those in Treacher Collins syndrome. Finally, for one child, the genetics chart was not available for review, leaving 71 unrelated patients with a diagnosis of an ASD. Forty of these patients (57%) were evaluated by one of the authors (G.E.H., K.L.M.), and the remainder were seen by one of four additional clinical geneticists at our institution. All the physicians used the clinical guideline, although diagnostic testing was performed as clinically indicated at the discretion of the geneticist evaluating the patient.
Sixty-one of the patients were evaluated in genetics for the first time, and 10 were seen for a follow-up appointment. The diagnosis of an ASD was based on DSM-IV criteria and was provided by a referring developmental pediatrician for 49 patients, by a child psychologist or psychiatrist for eight patients, and by a neurologist for four. One patient was referred from his school. For nine patients, the original source of the diagnosis could not be ascertained from the available medical records; however, review of the chart revealed sufficient data to support the diagnosis of an ASD.
Twenty-six of the children participated in psychological testing at The Autism Center at Children's Hospital, Columbus, OH, as part of their evaluation process to help clarify the diagnosis and aid in treatment planning. Although the testing battery varied depending on the age of the child, typical tools included a traditional intellectual assessment (e.g., Wechsler Preschool and Primary Scale of Intelligence–Third Edition, Stanford-Binet Intelligence Scales–Fifth Edition), nonlanguage-based measures of intelligence (e.g., Leiter International Performance Scale–Revised), language screening measures (e.g., Peabody Picture Vocabulary Test–Third Edition, Expressive One Word Picture Vocabulary Test), measures of early academic skills (e.g., Bracken Basic Concepts Scale–Revised), measures of adaptive skills (e.g., Scales of Independent Behavior–Revised), and additional autism specific measures (e.g., Gilliam Autism Rating Scale, Autism Diagnostic Observation Schedule). Five additional children had an intellectual assessment through their school system, and for two, these data were included in the evaluation provided by a developmental pediatrician.
Laboratory testing was performed by Columbus Children's Hospital unless stated otherwise and included peripheral blood karyotype at 550-band resolution or higher, molecular studies for fragile X syndrome (The Ohio State University Molecular Pathology Laboratory), array comparative genomic hybridization (aCGH) (SignatureChip®, Signature Genomic Laboratories), plasma amino acids, urine organic acids, plasma homocysteine, uric acid, plasma and urine guanidinoacetate (GAA), 7-dehydrocholesterol (7DHC), MECP2 gene sequencing (Baylor College of Medicine), PTEN gene sequencing (The Ohio State University Molecular Pathology Laboratory), and DNA methylation analysis for Prader-Willi/Angelman syndromes. A urine purine and pyrimidine panel was performed at Mayo Laboratories and urine sulfocysteine at Duke University Biochemical Genetics Laboratory.
RESULTS
Testing and referral guideline
There have been several recent reports discussing the appropriate genetic evaluation and diagnostic yield of genetic testing in children with ASDs.15–19 Some of the data from the publication by Schaefer and Lutz16 were first presented at the 2005 American College of Medical Genetics meeting. After this presentation, in collaboration with developmental pediatricians and psychologists at The Autism Center at Children's Hospital, Columbus, OH, we developed a “suggested” guideline for laboratory testing (Table 1) and referral to genetics for children with a proven or suspected ASD. This guideline is now used in the clinical evaluation of most children newly diagnosed with an ASD locally: new patients seen at our Autism Center undergo an interdisciplinary evaluation that includes a developmental pediatrician and psychologist, as well as additional support staff (e.g., psychometrician, resource coordinator). The appointment includes a clinical interview, administration of the Autism Diagnostic Observation Schedule, and a brief physical examination, after which the family is provided with verbal feedback regarding diagnostic conceptualization and treatment recommendations. If appropriate, each interdisciplinary evaluation is followed by a linked appointment for more extensive psychology testing to further clarify cognitive functioning, preacademic skills, expressive and receptive language skills, visual-motor integration skills, and fine motor abilities. Unless a specific genetic syndrome is diagnosed, “first-tier” testing is ordered by the developmental pediatrician, including chromosome analysis, DNA for fragile X syndrome in both males and females,20,21 hearing screening, lead level, and metabolic screening. The latter includes plasma amino acids and urine organic acids. A plasma homocysteine level is obtained because we have previously identified two children with inherited defects in B12 metabolism22 and elevated homocysteine levels who presented with only mild mental retardation. Mild elevations in homocysteine may be missed on a routine plasma amino acid analysis.
Initially, all new ASD patients were referred for a clinical genetics evaluation. Subsequently, criteria were modified slightly. Patients are now referred if any of the “first-tier” testing is abnormal; if there is a positive family history of an ASD or related disorder; if there is an indication for further genetics evaluation, such as severe developmental delay (IQ <50), dysmorphic features, microcephaly, seizures, or the presence of one or more major malformations and/or the family or referring clinician desires further input from genetics. The genetics clinic visit is typically scheduled 4 to 6 weeks after the initial evaluation, so that the initial laboratory test results are available. If abnormal results are obtained, appropriate genetic counseling of the family can be performed as well as any additional necessary follow-up testing. If no specific genetic diagnosis has been found, the medical history, including records from the developmental pediatrician and psychologist, are reviewed; a three-generation pedigree is constructed by a genetic counselor working with the physician; and a complete physical examination is performed, with special attention to the presence of any dysmorphic features or those suggestive of specific syndromes. Further genetic testing is performed based on specific clinical indications, at the discretion of the clinical geneticist evaluating the child (Table 1) and would often include targeted aCGH (SignatureChip®).23 The SignatureChip detects unbalanced submicroscopic genomic alterations at more than 150 loci, including subtelomeres and known deletions or duplications associated with a clinical phenotype. GAA profiles in plasma and urine, the latter in males only, are obtained to exclude disorders of creatine synthesis. Mutations in an X-linked gene encoding a creatine transporter (SCL6A8) are a recently described cause of mental retardation, seizures, and autism in males.24–26 Plasma sterol profiles are obtained in children suspected to have Smith-Lemli-Opitz syndrome (SLOS) or as a screening test. It has been reported that SLOS may present as autism in rare cases,27,28 although a more recent study did not confirm these initial findings.29
For children who lack speech, DNA methylation analysis is performed to exclude Angelman syndrome, with sequencing of the Angelman gene (UBE3A ligase) in classic cases if a normal pattern of DNA methylation is found.30 Sequencing of the MECP2 gene is performed in girls with features of Rett syndrome as well as in males or females with suspicion of “atypical Rett” syndrome, which can include mild mental retardation or autism.31,32 Included in this group would be girls with autism and preserved speech, normal head circumference, or some use of their hands; girls with no speech and features similar to those found in Angelman syndrome; and girls with autism, mental retardation, and seizures. In addition, MECP2 sequencing would be considered in boys with autism, mental retardation, and ataxia or tremor.
Electroencephalography (EEG) is performed if there is a suspicion of seizures or occasionally with a history of significant regression, whereas brain magnetic resonance imaging (MRI) is indicated for the presence of seizures, suspected brain malformation, or microcephaly. Selected disorders of purine and pyrimidine metabolism can be associated with autism in rare cases.33–35 Abnormal metabolites for these disorders can be detected by specialized testing in a urine sample, whereas plasma uric acid is a surrogate marker to screen for some, but not all, of these disorders. Finally, a recent report identified heterozygous mutations in the PTEN (phosphatase and tensin homologue, deleted on chromosome 10) tumor suppressor gene in three patients with significant macrocephaly and autism.36 To confirm this observation and to help ascertain the frequency of such mutations, we have screened for PTEN mutations by DNA sequence analysis in children with ASDs and absolute or relative macrocephaly.
Clinical features of the patients
A summary of the clinical features of the 71 unrelated patients with ASDs evaluated in the Genetics Clinic during the 14-month period is shown in Table 2. Although our guideline suggests referral to genetics of patients with an ASD and one or more malformations, two children with malformations and syndromic diagnoses were excluded from the analysis (see METHODS) to limit the analysis to those with isolated ASDs and to prevent possible skewing of the results.
By history, there were eight nuclear and five extended families with more than a single individual affected with an ASD, eight extended families with psychiatric disease, and four with both (21/71 families; 30%). Data concerning the extended family members need to be confirmed through medical records and/or direct evaluation. Psychiatric disease reported in parents and/or extended family members included bipolar disorder, depression, schizophrenia, obsessive-compulsive disorder, and “other mental health problems.” In addition, several parents reported that they or members of their extended family had learning disabilities, speech delay or dyslexia, or mental retardation. A “broader autism phenotype” (BAP),37–39 in which core components of an ASD may be present in milder form, as well as a range of psychiatric diagnoses,40,41 have been described previously in parents and extended family members in ASD pedigrees. The presence of the BAP was not formally assessed in parents during clinical genetics evaluations.
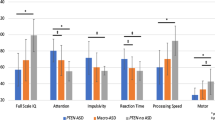
For 30 patients, results of psychological testing included standard scores or equivalents on measures of intellectual functioning. These data demonstrated mental retardation (IQ <70) in 12 of 30 (40%), borderline IQ (70–80) in nine of 30 (30%), and low average or above in nine of 30 (30%). Additionally, standard scores on the Leiter International Performance Scale–Revised, a nonlanguage-based measure of intellectual functioning, ranged from 52 to 105, with two of 11(18%) of the children who were administered the test falling in the mildly impaired range or below, three of 11 (27%) borderline, and six of 11 (55%) low average or above.
Results of genetic testing
As shown in Tables 3 and 4, the diagnostic yield of genetic testing in this population was 10% (7/71). Two children had visible chromosome abnormalities: a male with a karyotype of 47,XYY and another male with several de novo small marker chromosomes. ASDs have been reported in several males with 47,XYY,42–45 although a causal association has not been firmly established.
One male with autism, age 34 months, was noted to have a submicroscopic deletion, detected by aCGH, but not by regular chromosome analysis. The rearrangement, confirmed by fluorescence in situ hybridization (FISH), was found in an interstitial region on chromosome 1 [(ish del(1)(q21.1q21.1(RP11-841D19-)], which is not associated with a defined genetic syndrome. One parent carried the identical genomic rearrangement. The child was not dysmorphic, lacked all speech, and had a nonlanguage IQ of 69 (2nd percentile) using the Leiter International Performance Scale–Revised. Additional testing, including DNA methylation analysis for Angelman syndrome, was normal. Deletions involving 1q21.1 have been identified in several probands with congenital heart defects,46 but may also be a normal copy number variant.23,47 For this reason, the identification of the genomic rearrangement in this patient was not included in the overall positive yield.
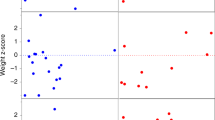
PTEN gene sequencing was performed in 16 patients, 12 with absolute macrocephaly (HC ≥95% for age) and four with relative macrocephaly (75%–98% for age) compared with their weight and/or height. Heterozygous PTEN mutations were identified in two patients with classic autism. A girl, age 2 years, had a de novo single base insertion, 530insT, creating a frame shift and premature termination, and a boy, age 4 years, carried a nonsense mutation, R130X, inherited from his asymptomatic father. Both patients had marked macrocephaly, with HCs of 55.5 cm and 56 cm (+5.2 and +2.6 SD), respectively. Clinical features of these patients are more fully described elsewhere.48 Two females with classic Rett syndrome, identified by developmental pediatricians based on their clinical features, had heterozygous nonsense mutations R255X and R168X, respectively, in the MECP2 gene. A third female, age 2 years, with the heterozygous MECP2 mutation R255X, had some delays and typical hand wringing but a head circumference in the 50th percentile.
Interestingly, metabolic screening produced no positive testing results (Table 3). This included screening for disorders of purine and pyrimidine metabolism using plasma uric acid in 26 patients, a urine purine and pyrimidine panel in three, and both in five. One child with autism had a slightly low plasma uric acid (1.6 mL/dL; normal, 2–7 mg/dL); however, repeat uric acid was 2.0 mg/dL and urine sulfocysteine was normal.
Finally, neuroimaging had been performed previously in 16 patients (computed tomography in four; MRI in 12), with normal results in all. One of nine patients in whom an EEG was previously performed was found to have subclinical focal epileptiform discharges, a finding that was considered nonspecific and nondiagnostic.
DISCUSSION
Genetic evaluation of children with an ASD is becoming increasingly important as new single genes with major effects and small genomic rearrangements detected by aCGH are identified in subsets of affected individuals. The identification of a single gene or chromosomal etiology within an individual and family is extremely important for estimating recurrence risks and, possibly, prognosis, including later medical complications. Excluding known causes as completely as possible is also important for research efforts to identify autism susceptibility loci. Indeed, the overall disappointing results in genome-wide linkage and association studies to date may be due, in part, to heterogeneity resulting from inclusion of such individuals.
The yield of genetic testing in our sample is similar to that reported by Abdul-Rahman and Hudgins15 (8.3%), but much lower than that reported by Schaefer and Lutz16 (>40%). This discrepancy likely relates to differences in the referral patterns at each institution and the inclusion of patients with syndromic diagnoses, such as tuberous sclerosis, neurofibromatosis, and Sotos syndrome, in the latter study. The majority of the patients in our series were referred from our Autism Center rather than primary care providers and, hence, represented children with what has been called “isolated” or “nonsyndromic” ASD.6 To avoid skewing of our results, we excluded two children with apparent “syndromic” ASDs from our analysis.
We were initially surprised that we did not detect any patients in our series with fragile X syndrome or the large, SNRPN-positive maternally derived 15q11-q13 duplication. Indeed, review of our clinical genetics database determined that a boy, age 2 years, was referred for genetics counseling within the 14-month time period with a diagnosis of fragile X syndrome after molecular testing by his pediatrician for developmental delay and a positive maternal family history. Another boy, age 22 months, was referred for genetic counseling after a karyotype demonstrating a marker chromosome with an inv dup15q11-13 that was SNRPN positive by FISH. He presented with hypotonia and developmental delay, which are almost universal features in the pathologic maternally derived 15q duplication.49–51 Thus, at our institution, patients with these diagnoses may be referred by their primary care provider or another physician before an ASD is apparent or suspected.
Vorstman et al.52 recently cataloged all the known cytogenetic abnormalities reported in individuals with ASDs, including the common duplication in 15q11-q13 and recurrent deletions at 2q37, 5p15, 22q11.2 (the DiGeorge critical region), 22q13.3, and Xp22-p23. The Xp deletions include neuroligin 4; mutations in this gene are a rare cause of autism in males.53,54 The technology of aCGH has extended the ability to detect submicroscopic genomic rearrangements in patients with nonsyndromic mental retardation and/or autism. Currently, commercial arrays target selected regions of the genome, including the subtelomeres and known deletions and duplications that are associated with genetic syndromes.23,55,56 With the advent of arrays with genome-wide coverage, it is likely that additional small cryptic genomic rearrangements will be identified in some ASD patients. Indeed, Jacquemot et al.57 identified “clinically relevant” genomic rearrangements in 8/29 (27.5%) patients with “syndromic” ASDs using a large insert clone (BAC or PAC) array developed at the Wellcome Trust Sanger Institute with a resolution of 1 Mb. All the patients studied met DSM-IV criteria for the diagnosis of an ASD and had a normal high-resolution peripheral blood karyotype. The rearrangements were all unique and included two duplications and six deletions, ranging in size from 1.4 to 16 Mb. The eight patients with a cytogenetic abnormality were all thought to be dysmorphic and had mild to moderate mental retardation.
A single patient in our series had a submicroscopic deletion (1q21.1) detected by targeted BAC array. The clinical significance of this rearrangement is unknown due to its occurrence in one of the child's parents and in some normal individuals.23,47 It is likely that this deletion is unrelated to his ASD and is a normal variant, and, for this reason, we did not include the result in our positive findings. It is also possible that it is associated with disease susceptibility, and additional susceptibility loci are present in the affected child but not in the parent with the same finding. Finally, it is possible that a second disease-causing event, such as a specific gene mutation, is present on the remaining chromosome in the affected child, but not in the parent.
Given the fact that no confirmed pathogenic genomic rearrangements were identified in the 38 patients with “isolated” ASDs examined here, it may be prudent to reserve aCGH for those children with “syndromic” autism, moderate or severe mental retardation (nonverbal IQ <50), and/or dysmorphic features as in the study of Jacquemot et al.57 However, because our study employed targeted arrays, it remains possible that oligonucleotide or tiling arrays with genome-wide coverage will detect pathogenic genomic rearrangements in some patients with nonsyndromic ASDs.
As described in more detail elsewhere,48 we identified heterozygous PTEN mutations in two children with autism and macrocephaly, confirming the earlier report of Butler et al.36 Subsequently, we identified PTEN mutations in two additional families, including one with identical twins both of whom have autism. All the affected individuals had HCs much greater than the 98 percentile for age, except for one of the identical twins whose head circumference was 95% to 98% for age. We believe that PTEN mutations are a recently recognized causative gene in a subset of children with autism and macrocephaly. PTEN gene sequencing should be encouraged in all individuals with developmental delay and/or autism with concomitant macrocephaly.
Finally, like others,15,16 we did not detect the presence of a metabolic disorder in any of the patients screened. Routine metabolic testing may not be warranted in children with an ASD without another indication, such as recurrent illness, failure to thrive, seizures, or hypotonia. However, given the relatively low cost of such screening and the potential for specific therapeutic intervention, at least in some cases, we have continued to obtain plasma amino acids, urine organic acids, and plasma homocysteine in ASD patients seen at our institution.
REFERENCES
MMWR. Mental health in the United States: parental report of diagnosed autism in children aged 4-17 years–United States, 2003–2004. MMWR Morb Mortal Wkly Rep 2006; 55: 481–486.
Muhle R, Trentacoste SV, Rapin I . The genetics of autism. Pediatrics 2004; 113: e472–486.
Spence SJ . The genetics of autism. Semin Pediatr Neurol 2004; 11: 196–204.
Wassink TH, Brzustowicz LM, Bartlett CW, Szatmari P . The search for autism disease genes. Ment Retard Dev Disabil Res Rev 2004; 10: 272–283.
Coon H . Current perspectives on the genetic analysis of autism. Am J Med Genet C Semin Med Genet 2006; 142: 24–32.
Cohen D, Pichard N, Tordjman S, Baumann C, et al. Specific genetic disorders and autism: clinical contribution towards their identification. J Autism Dev Disord 2005; 35: 103–116.
Dykens EM, Sutcliffe JS, Levitt P . Autism and 15q11-q13 disorders: behavioral, genetic, and pathophysiological issues. Ment Retard Dev Disabil Res Rev 2004; 10: 284–291.
Jiang YH, Bressler J, Beaudet AL . Epigenetics and human disease. Annu Rev Genomics Hum Genet 2004; 5: 479–510.
Schanen NC . Epigenetics of autism spectrum disorders. Hum Mol Genet 2006; 15: R138–150.
Lawler CP, Croen LA, Grether JK, Van de Water J . Identifying environmental contributions to autism: provocative clues and false leads. Ment Retard Dev Disabil Res Rev 2004; 10: 292–302.
IMGSAC. A genomewide screen for autism: strong evidence for linkage to chromosomes 2q, 7q, and 16p. Am J Hum Genet 2001; 69: 570–581.
Ylisaukko-Oja T, Alarcon M, Cantor RM, Auranen M, et al. Search for autism loci by combined analysis of Autism Genetic Resource Exchange and Finnish families. Ann Neurol 2006; 59: 145–155.
Hu-Lince D, Craig DW, Huentelman MJ, Stephan DA . The Autism Genome Project: goals and strategies. Am J Pharmacogenomics 2005; 5: 233–246.
Schellenberg GD, Dawson G, Sung YJ, Estes A, et al. Evidence for multiple loci from a genome scan of autism kindreds. Mol Psychiatry 2006; 11: 1049–1060.
Abdul-Rahman OA, Hudgins L . The diagnostic utility of a genetics evaluation in children with pervasive developmental disorders. Genet Med 2006; 8: 50–54.
Schaefer GB, Lutz RE . Diagnostic yield in the clinical genetic evaluation of autism spectrum disorders. Genet Med 2006; 8: 549–556.
Battaglia A, Bonaglia MC . The yield of subtelomeric FISH analysis in the evaluation of autistic spectrum disorders. Am J Med Genet C Semin Med Genet 2006; 142: 8–12.
Challman TD, Barbaresi WJ, Katusic SK, Weaver A . The yield of the medical evaluation of children with pervasive developmental disorders. J Autism Dev Disord 2003; 33: 187–192.
Battaglia A, Carey JC . Etiologic yield of autistic spectrum disorders: a prospective study. Am J Med Genet C Semin Med Genet 2006; 142: 3–7.
Philofsky A, Hepburn SL, Hayes A, Hagerman R, et al. Linguistic and cognitive functioning and autism symptoms in young children with fragile X syndrome. Am J Ment Retard 2004; 109: 208–218.
Hagerman RJ, Ono MY, Hagerman PJ . Recent advances in fragile X: a model for autism and neurodegeneration. Curr Opin Psychiatry 2005; 18: 490–496.
Rosenblatt DS, Fenton WA . Inherited disorders of folate and cobalamin transport and metabolism. In: Scriver CR, Beaudet AL, Sly WS, Valle D, editors, The metabolic and molecular bases of inherited disease. New York: McGraw-Hill, 2001; 3897–3933.
Shaffer LG, Kashork CD, Saleki R, Rorem E, et al. Targeted genomic microarray analysis for identification of chromosome abnormalities in 1500 consecutive clinical cases. J Pediatr 2006; 149: 98–102.
Verhoeven NM, Salomons GS, Jakobs C . Laboratory diagnosis of defects of creatine biosynthesis and transport. Clin Chim Acta 2005; 361: 1–9.
Clark AJ, Rosenberg EH, Almeida LS, Wood TC, et al. X-linked creatine transporter (SLC6A8) mutations in about 1% of males with mental retardation of unknown etiology. Hum Genet 2006; 119: 604–610.
Poo-Arguelles P, Arias A, Vilaseca MA, Ribes A, et al. X-Linked creatine transporter deficiency in two patients with severe mental retardation and autism. J Inherit Metab Dis 2006; 29: 220–223.
Tierney E, Nwokoro NA, Porter FD, Freund LS, et al. Behavior phenotype in the RSH/Smith-Lemli-Opitz syndrome. Am J Med Genet 2001; 98: 191–200.
Kelley RI, Herman GE . Inborn errors of sterol biosynthesis. Annu Rev Genomics Hum Genet 2001; 2: 299–341.
Tierney E, Bukelis I, Thompson RE, Ahmed K, et al. Abnormalities of cholesterol metabolism in autism spectrum disorders. Am J Med Genet B Neuropsychiatr Genet 2006; 141: 666–668.
Williams CA, Beaudet AL, Clayton-Smith J, Knoll JH, et al. Angelman syndrome 2005: updated consensus for diagnostic criteria. Am J Med Genet A 2006; 140: 413–418.
Moretti P, Zoghbi HY . MeCP2 dysfunction in Rett syndrome and related disorders. Curr Opin Genet Dev 2006; 16: 276–281.
Ham AL, Kumar A, Deeter R, Schanen NC . Does genotype predict phenotype in Rett syndrome?. J Child Neurol 2005; 20: 768–778.
Simmonds HA, Duley JA, Fairbanks LD, McBride MB . When to investigate for purine and pyrimidine disorders. Introduction and review of clinical and laboratory indications. J Inherit Metab Dis 1997; 20: 214–226.
Spiegel EK, Colman RF, Patterson D . Adenylosuccinate lyase deficiency. Mol Genet Metab 2006; 89: 19–31.
Nyhan WL . Disorders of purine and pyrimidine metabolism. Mol Genet Metab 2005; 86: 25–33.
Butler MG, Dasouki MJ, Zhou XP, Talebizadeh Z, et al. Subset of individuals with autism spectrum disorders and extreme macrocephaly associated with germline PTEN tumour suppressor gene mutations. J Med Genet 2005; 42: 318–321.
Piven J, Palmer P, Jacobi D, Childress D, et al. Broader autism phenotype: evidence from a family history study of multiple-incidence autism families. Am J Psychiatry 1997; 154: 185–190.
Constantino JN, Todd RD . Autistic traits in the general population: a twin study. Arch Gen Psychiatry 2003; 60: 524–530.
Bishop DV, Maybery M, Wong D, Maley A, et al. Characteristics of the broader phenotype in autism: a study of siblings using the children's communication checklist-2. Am J Med Genet B Neuropsychiatr Genet 2006; 141: 117–122.
Piven J, Palmer P . Psychiatric disorder and the broad autism phenotype: evidence from a family study of multiple-incidence autism families. Am J Psychiatry 1999; 156: 557–563.
Larsson HJ, Eaton WW, Madsen KM, Vestergaard M, et al. Risk factors for autism: perinatal factors, parental psychiatric history, and socioeconomic status. Am J Epidemiol 2005; 161: 916–925.
Nicolson R, Bhalerao S, Sloman L . 47,XYY karyotypes and pervasive developmental disorders. Can J Psychiatry 1998; 43: 619–622.
Weidmer-Mikhail E, Sheldon S, Ghaziuddin M . Chromosomes in autism and related pervasive developmental disorders: a cytogenetic study. J Intellect Disabil Res 1998; 42: 8–12.
Geerts M, Steyaert J, Fryns JP . The XYY syndrome: a follow-up study on 38 boys. Genet Couns 2003; 14: 267–279.
Kielinen M, Rantala H, Timonen E, Linna SL, et al. Associated medical disorders and disabilities in children with autistic disorder: a population-based study. Autism 2004; 8: 49–60.
Christiansen J, Dyck JD, Elyas BG, Lilley M, et al. Chromosome 1q21.1 contiguous gene deletion is associated with congenital heart disease. Circ Res 2004; 94: 1429–1435.
Sharp AJ, Hansen S, Selzer RR, Cheng Z, et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 2006; 38: 1038–1042.
Herman GE, Butter E, Enrile B, Pastore M, et al. Increasing knowledge of PTEN germline mutations: two additional patients with autism and macrocephaly. Am J Med Genet A 2007; 143: 589–593.
Battaglia A . The inv dup(15) or idic(15) syndrome: a clinically recognisable neurogenetic disorder. Brain Dev 2005; 27: 365–369.
Eggermann K, Mau UA, Bujdoso G, Koltai E, et al. Supernumerary marker chromosomes derived from chromosome 15: analysis of 32 new cases. Clin Genet 2002; 62: 89–93.
Dennis NR, Veltman MW, Thompson R, Craig E, et al. Clinical findings in 33 subjects with large supernumerary marker(15) chromosomes and 3 subjects with triplication of 15q11-q13. Am J Med Genet A 2006; 140: 434–441.
Vorstman JA, Staal WG, van Daalen E, van Engeland H, et al. Identification of novel autism candidate regions through analysis of reported cytogenetic abnormalities associated with autism. Mol Psychiatry 2006; 11: 1, 18–28.
Laumonnier F, Bonnet-Brilhault F, Gomot M, Blanc R, et al. X-linked mental retardation and autism are associated with a mutation in the NLGN4 gene, a member of the neuroligin family. Am J Hum Genet 2004; 74: 552–557.
Blasi F, Bacchelli E, Pesaresi G, Carone S, et al. Absence of coding mutations in the X-linked genes neuroligin 3 and neuroligin 4 in individuals with autism from the IMGSAC collection. Am J Med Genet B Neuropsychiatr Genet 2006; 141: 220–221.
Tyson C, Harvard C, Locker R, Friedman JM, et al. Submicroscopic deletions and duplications in individuals with intellectual disability detected by array-CGH. Am J Med Genet A 2005; 139: 173–185.
Cheung SW, Shaw CA, Yu W, Li J, et al. Development and validation of a CGH microarray for clinical cytogenetic diagnosis. Genet Med 2005; 7: 422–432.
Jacquemont ML, Sanlaville D, Redon R, Raoul O, et al. Array-based comparative genomic hybridisation identifies high frequency of cryptic chromosomal rearrangements in patients with syndromic autism spectrum disorders. J Med Genet 2006; 43: 843–849.
Acknowledgements
The authors thank Erica Mazzone for help with the preparation of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Herman, G., Henninger, N., Ratliff-Schaub, K. et al. Genetic testing in autism: how much is enough?. Genet Med 9, 268–274 (2007). https://doi.org/10.1097/GIM.0b013e31804d683b
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e31804d683b
Keywords
This article is cited by
-
Detection of autism spectrum disorder-related pathogenic trio variants by a novel structure-based approach
Molecular Autism (2024)
-
A family with PTEN mutations with malignancy and an unusually high number of offspring with autism spectrum disorder: a case report
Journal of Medical Case Reports (2018)
-
Identification of mutations in the PI3K-AKT-mTOR signalling pathway in patients with macrocephaly and developmental delay and/or autism
Molecular Autism (2017)
-
From molecules to neural morphology: understanding neuroinflammation in autism spectrum condition
Molecular Autism (2016)
-
The role of glutamate and its receptors in autism and the use of glutamate receptor antagonists in treatment
Journal of Neural Transmission (2014)