Abstract
Anaerobic nitrate-dependent Fe(II) oxidation is widespread in various environments and is known to be performed by both heterotrophic and autotrophic microorganisms. Although Fe(II) oxidation is predominantly biological under acidic conditions, to date most of the studies on nitrate-dependent Fe(II) oxidation were from environments of circumneutral pH. The present study was conducted in Lake Grosse Fuchskuhle, a moderately acidic ecosystem receiving humic acids from an adjacent bog, with the objective of identifying, characterizing and enumerating the microorganisms responsible for this process. The incubations of sediment under chemolithotrophic nitrate-dependent Fe(II)-oxidizing conditions have shown the enrichment of TM3 group of uncultured Actinobacteria. A time-course experiment done on these Actinobacteria showed a consumption of Fe(II) and nitrate in accordance with the expected stoichiometry (1:0.2) required for nitrate-dependent Fe(II) oxidation. Quantifications done by most probable number showed the presence of 1 × 104 autotrophic and 1 × 107 heterotrophic nitrate-dependent Fe(II) oxidizers per gram fresh weight of sediment. The analysis of microbial community by 16S rRNA gene amplicon pyrosequencing showed that these actinobacterial sequences correspond to ∼0.6% of bacterial 16S rRNA gene sequences. Stable isotope probing using 13CO2 was performed with the lake sediment and showed labeling of these Actinobacteria. This indicated that they might be important autotrophs in this environment. Although these Actinobacteria are not dominant members of the sediment microbial community, they could be of functional significance due to their contribution to the regeneration of Fe(III), which has a critical role as an electron acceptor for anaerobic microorganisms mineralizing sediment organic matter. To the best of our knowledge this is the first study to show the autotrophic nitrate-dependent Fe(II)-oxidizing nature of TM3 group of uncultured Actinobacteria.
Similar content being viewed by others
Introduction
Organic matter degradation in aquatic environments such as wetlands or lake sediments is mediated anaerobically according to the redox stratification of the sediment with methanogenesis being the final process (Ponnamperuma, 1972). Studies have shown that Fe(III) reduction can suppress methanogenesis and could contribute to the mineralization of a large fraction of sediment organic matter (Roden and Wetzel, 1996; Küsel et al., 2008). Due to the high concentration of biologically available iron (Steinmann and Shotyk, 1997) and favorable limnological conditions in humic-rich habitats, Fe(III) reduction could act as a dominant terminal electron-accepting process (Reiche et al., 2008) given a continuous recycling of Fe(III) from Fe(II) in the sediment.
Fe(II) oxidation in natural environments occurs at the oxic-anoxic interface by chemically reacting with atmospheric O2 or by aerobic Fe(II)-oxidizing bacteria (Emerson and Revsbech, 1994; Edwards et al., 2004). In the deeper layers of the sediment, Fe(II) oxidation is known to happen in the rhizosphere of plants by root released O2 (Frenzel et al., 1999; Neubauer et al., 2007; Weiss et al., 2007) or in the sediment by nitrate-dependent Fe(II) oxidation by organisms growing either autotrophically or heterotrophically (Straub et al., 1996; Benz et al., 1998). An increasing body of literature over the last decade has shown that biogenic Fe(II) oxidation could have a more prominent role in iron cycling than considered previously, especially in low-pH environments (Lu et al., 2010; Lüdecke et al., 2010).
Chemolithotrophic nitrate-dependent Fe(II) oxidation was first reported by Straub et al., (1996) and has been reported in various environments since, primarily in lake sediments (Straub and Buchholz-Cleven, 1998; Hauck et al., 2001; Muehe et al., 2009). Although autotrophic nitrate-dependent oxidation of Fe(II) was observed in many natural habitats, to date there are only two pure cultures available: the hyperthermophilic archaeon Ferroglobus placidus (Hafenbradl et al., 1996) and the betaproteobacterium Pseudogulbenkiania sp. strain 2002, which can alternatively grow autotrophically or heterotrophically (Weber et al., 2006). Autotrophic nitrate-dependent Fe(II) oxidation is a poorly understood process but could be of major significance in anoxic cycling of iron in lake sediments; however, due to the difficulty in culturing these organisms and lack of either functional gene or 16S rRNA gene-based primers for this functional guild, the ecology of these microorganisms is difficult to investigate.
Lake Grosse Fuchskuhle is an acidic bog lake with a high concentration of recalcitrant and high molecular weight humic acids (Sachse et al., 2001). The low pH of the sediment (pH 4.5), complexation of Fe(II) by organic matter (Theis and Singer, 1974) and the ability of oxygen to penetrate only the top few millimeters of the sediment could greatly reduce the rate of abiotic Fe(II) oxidation (Stumm and Morgan, 1996). As a result, we hypothesize that microbially-mediated anaerobic nitrate-dependent Fe(II) oxidation in the sediment could be contributing significantly to Fe(II) oxidation. The objective of this study was to identify microbial groups involved in nitrate-dependent Fe(II) oxidation in the littoral sediment of Lake Grosse Fuchskuhle.
Materials and methods
Sampling
Lake sediment samples were collected in April 2010 from acidic dystrophic Lake Grosse Fuchskuhle in the Brandenburg-Mecklenburg Lake District (Germany). Lake Grosse Fuchskuhle is an artificially divided lake with different pH values in the four compartments. A main divergent factor is the inflow of humic acids from an adjacent bog; the southwest (SW) basin is the most acidic and the northeast (NE) is the least (Koschel, 1995). Sediment samples were taken by a gravity corer (Uwitec, Mondsee, Austria) from the profundal and littoral of both the SW and NE basins, but most experimental work in this study focused on the SW littoral samples. The top 10 cm of the sediments were collected using 6 cm diameter plexiglass cores. The sediment had a high concentration of humic acids, which were visible from the dark brown colour of the sediment and overlying water. The collected sediment was composed mainly of coarse particulate organic material (mainly half-decayed leaves and small pieces of wood). The pH of the SW littoral sediment was 4.5. The concentration of Fe(II) and Fe(III) in the sediment was determined using the ferrozine assay (Stookey, 1970) and nitrate concentrations were measured by flow injection analysis (Tecator, Rellingen, Germany).
Enrichment of nitrate-dependent Fe(II)-oxidizing bacteria
The enrichment of nitrate-dependent Fe(II)-oxidizing bacteria was done according to the procedure described by Straub et al., (1996) with the exception of using phosphate buffer, a pH of 4.5 and FeCl2 instead of FeSO4 to prevent the growth of sulfate reducing bacteria. The freshwater medium was prepared as follows: NH4Cl (0.3 g), MgSO4.7H2O (0.05 g), MgCl2.6H2O (0.4 g) and CaCl2 (0.1 g) buffered by the addition of 100 ml of 0.5 M KH2PO4 to 900 ml of the above medium (final pH 4.5) added after autoclaving and cooling to room temperature to avoid precipitation. Filter-sterilized vitamins and trace elements (Widdel and Bak, 1992) were added to the medium after autoclaving. Oxygen was depleted using a vacuum manifold and repeatedly flushing the headspace with N2 gas. Twenty milliliters of the medium was dispensed into 120 ml serum bottles under a N2 atmosphere in an anaerobic chamber (Mecaplex, Grenchen, Switzerland). Different combinations of FeCl2 (10 mM), sodium nitrate (4 mM), sodium acetate (2.5 mM) and CO2 (5% headspace) were included in the enrichment medium. The Fe(II)-EDTA stock was prepared by mixing 50 mM EDTA with 100 mM FeCl2. One set of the incubations was done without phosphate buffered medium and the pH adjusted to 4.5 with HCl. A volume of 1.5 ml of Lake Grosse Fuchskuhle SW littoral sediment was used as an inoculum. All incubations were performed in triplicate and incubated in the dark on a shaker (150 r.p.m.) for 10 days.
A time-course experiment was performed to determine the ratio of Fe(II) to nitrate consumed over a period of 14 days. The experimental setup contained the above medium with nitrate and Fe(II) with 1% of the actinobacterial enrichment as inoculum. 5% CO2 was added to the headspace as the sole carbon source. Samples for terminal restriction fragment length polymorphism (T-RFLP) as well as for Fe(II) and nitrate measurements were taken every 24 h. The sediment filtrate was used for ammonia determinations (Kandler and Gerber, 1988). Nitrate measurements were done colorimetrically as described previously (Hart et al., 1994). N2O was measured by gas chromatography (Carlo Erba Instruments, GC 8000, Wigan, UK) using a 63Ni-electron capture detector.
Enumeration of Fe(II)-oxidizing bacteria by most probable number (MPN)
An MPN method was used to enumerate the Fe(II)-oxidizing bacteria according to the procedure described previously (Straub and Buchholz-Cleven, 1998) with the above mentioned modifications. Two sets of MPN tubes were incubated each in triplicate. One set of tubes contained the phosphate buffered freshwater medium (pH 4.5) with Fe(II) and nitrate to enumerate autotrophic Fe(II)-oxidizing bacteria and the other set of tubes contained phosphate buffered freshwater medium, nitrate, Fe(II) and acetate for enumeration of heterotrophic Fe(II)-oxidizing bacteria. The tubes were incubated for 12 weeks in the dark at 25 °C and gently inverted daily. Tubes were scored positive based on the reduction in the amount of Fe(II) and acetate in the respective tubes compared with the uninoculated controls. The Fe(II) estimations were done using the ferrozine assay (Stookey, 1970) and a standard MPN table was used to calculate the cell numbers. DNA was extracted from the positive tubes and (T-RFLP) profiling was performed as described below for the identification of the bacterial groups growing in the tubes.
Stable isotope probing (SIP) experimental setup and gradient fractionation
Time series SIP incubations were performed in duplicate. Equal volumes of sterile deionized water and sediment were mixed and 40 ml was dispensed into 120 ml serum bottles capped with black butyl stoppers. The bottles were made anaerobic by flushing with N2 and 13CO2 (Campro Scientific, Berlin, Germany) was added to the headspace to a final concentration of 5%. Control bottles contained 5% unlabeled CO2. All the bottles were incubated at 25 °C on a shaker (150 r.p.m.) in the dark. The headspaces were renewed after every 4 days. Four microliters of sample were collected from all the incubation bottles after 3, 6 and 12 weeks. CO2 and CH4 measurements were done by gas chromatography fitted with a methanizer and flame-ionizing detector. The ratios of 12C and 13C were determined weekly using gas chromatography isotope ratio mass spectrometer. The collected sediment was centrifuged and the pore water was used for the analysis of cations by ion chromatography and volatile fatty acids by HPLC. The centrifuged sediment was frozen in liquid N2 and stored at −80 °C until the extraction of DNA. The pore water was used for the analysis of volatile fatty acids. Nucleic acids were extracted from the sediment using the Nucleospin soil kit (Macherey-Nagel, Düren, Germany).
Density gradient centrifugation of DNA (5.0 μg) extracted from the incubated samples was performed with a cesium chloride buoyant density of 1.72 g ml−1 subjected to centrifugation at 177 000 g for 36 h at 20 °C (Lueders et al., 2004). Cesium chloride gradients were fractionated from bottom to top by displacing the gradient medium with nuclease-free water using a syringe pump (Kent Scientific, Torrington, CT, USA) at a flow rate of 0.45 ml min−1, generating 12 fractions per density gradient. The density of each fraction was determined by refractometry (Reichert, Depew, NY, USA). DNA was recovered by PEG 6000 precipitation and dissolved in 30 μl of nuclease-free water (Applied Biosystems, Darmstadt, Germany).
The relative abundance of 16S rRNA genes within gradients was determined by real-time PCR using the SYBR Green JumpStart ReadyMix System (Sigma, Taufkirchen, Germany) as described previously (Stubner, 2002). The assays were performed using an iCycler instrument (Bio-Rad, Munich, Germany) and the associated software.
PCR, cloning and sequence analysis
For cloning and sequencing 16S rRNA genes from SIP experiments, PCR amplification was done using the Ba27f and Ba907r primers, which generates a product of ∼900 bp (Lueders et al., 2004). All the PCR reactions were performed in 50 μl volume with the following composition: 1 × PCR buffer (Promega, Mannheim, Germany), 0.2 mM MgCl2, 10 pmol of each primer, 10 μg of BSA (Roche, Mannheim, Germany), 2 U of GoTaq (Promega), 0.2 mM dNTPs (Fermentas, St Leon-Rot, Germany) and 1 μl of template DNA. The PCR was performed on a GeneAmp PCR system 9700 instrument (Applied Biosystems) with the following cycling conditions: 94 °C for 4 min, 35 cycles of 94 °C for 1 min, 52 °C for 40 s and 72 °C for 1 min, and a final extension for 10 min at 72 °C. PCR products were cloned using the pGEM-T Easy Vector System (Promega) and transformed into E. coli JM109 competent cells according to the manufacturer’s instructions. Twenty-one clones each were randomly picked from heavy and light fractions and sequenced. The phylogenetic affiliation of the clones was done using the ARB software package (Ludwig et al., 2004) and trees were constructed using the neighbor-joining method. Sequences were deposited in GenBank with accession numbers KC540872 to KC540892.
T-RFLP
The PCR amplification of bacterial 16S rRNA genes for T-RFLP analysis was performed as described above, except that the Ba27F primer was labeled with FAM (6-carboxyfluorescein). PCR products were purified using Qiagen PCR Purification Kit (Qiagen, Hilden, Germany). Approximately 100 ng of purified PCR product was used for restriction digestion. Digestions were performed in a reaction volume of 20 μl containing 1 × Tango buffer and 5 U of MspI enzyme (Fermentas); reactions were incubated at 37 °C in an incubator overnight. The reactions were processed using SigmaSpin Post Reaction Clean-Up Columns (Sigma) and 2 μl of the processed fragments were mixed with 11 μl of Hi-Di formamide (Applied Biosystems), 0.3 μl of ROX-labeled MapMarker 1000+30, 40 (BioVentures, Murfreesboro, TN, USA) and incubated at 94 °C for 3 min and cooled on ice. The size separation was performed using 3130 Genetic Analyzer (Applied Biosystems). T-RFs shorter than 50 bp were not considered to avoid the detection of primers and primer-dimers.
Pyrosequencing of bacterial 16S rRNA genes
DNA was extracted from littoral and profundal sediment from both the SW and NE basins of Lake Grosse Fuchskuhle using the Nucleospin soil kit (Macherey-Nagel). Bacterial 16S rRNA gene PCR products were amplified using the primers 343Fmod and 784Rmod as described previously (Köhler et al., 2012). The PCR products were sequenced using a Roche 454 GS Junior instrument. Sequence analysis was performed using the Mothur software v1.25.0 (Schloss et al., 2009). Processing of sequences within Mothur, including denoising and chimera removal, was performed according to the standard operating procedure of the software developer. Briefly, sequences were screened by allowing 1 mismatch to the barcode, 2 mismatches to the primer and a maximum homopolymer length of 8 bases. Sequences shorter than 200 bp were removed. Chimeras were removed using uchime within Mothur. Between 1047 and 3444 high-quality sequences were obtained from each sample. Identification of Actinobacteria was performed using the SILVA taxonomy and the classification was verified by adding these sequences into the SILVA108 reference tree (Pruesse et al., 2007) by parsimony within ARB (Ludwig et al., 2004).
Results
Fe(II), Fe(III), nitrate and methanogenesis potential in Lake Fuchskuhle littoral sediment
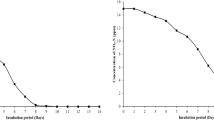
Duplicate sediment cores were collected from the littoral zone of Lake Grosse Fuchskuhle and partitioned into 5-cm sections to a depth of 30 cm. The concentration of Fe(II) and Fe(III) was determined at each depth (Figure 1a). Although insufficient replication of the data was available to calculate the significance, the observed Fe(II) concentration was higher than Fe(III) only between 5 to 20 cm. The concentration of CH4 and CO2 after anaerobic incubation for 7 weeks was determined (Figure 1b) and greatest methanogenesis potential was observed at the 15–20 cm depth. Nitrate was detectable in small amounts in the hypolimnion (4.0 m;<2 μmol l−1) and also in sediment pore water (2 μmol l−1) (results not shown).
Enumeration and enrichment of nitrate-dependent Fe(II)-oxidizing microorganisms
A MPN enrichment assay was performed to estimate the abundance of readily cultivable anaerobic nitrate-dependent Fe(II)-oxidizing microorganisms in the littoral sediment of Lake Grosse Fuchskuhle. The assay was performed with either CO2 or acetate as carbon source to estimate potential autotrophic and heterotrophic microorganisms. The assays indicated the presence of 1 × 104 autotrophic and 1 × 107 heterotrophic nitrate-dependent Fe(II)-oxidizers per gram fresh weight of sediment.
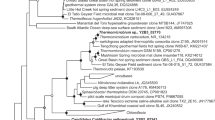
A freshwater medium was used to enrich potential nitrate-dependent Fe(II)-oxidizing microorganisms from Lake Grosse Fuchskuhle littoral sediment. After 10 days of incubation the enrichments were characterized by T-RFLP fingerprinting (Figure 2). The incubation with added Fe(II), nitrate and CO2 (Figure 2b) resulted in an enrichment culture characterized by a dominant T-RF of 146 bp. When Fe(II) or both Fe(II) and nitrate were omitted, enrichments characterized by a dominant T-RF of 429 bp were obtained (Figures 2c and d). When nitrate alone was omitted, the dominant T-RF was 97 bp (Figure 2e). Finally, an enrichment characterized by a 437 bp peak was obtained when acetate was provided. The results of the various enrichments were reproducible across triplicates (results not shown). 16S rRNA gene cloning and sequencing were performed to identify the 146 bp T-RF in the enrichment. A total of 86% of the clones obtained had the 146 bp T-RF and a phylogenetic analysis (Figure 3) indicated they belong to the uncultivated TM3 group Actinobacteria (Rheims et al., 1996). To our knowledge, no isolates from this group have been identified and their physiology is unknown.
T-RFLP fingerprinting of bacterial 16S rRNA genes from different enrichment incubations. The enrichment conditions are described in detail in the Materials and methods section. All enrichments were performed in triplicate and each produced similar T-RFLP profiles, therefore a single representative is shown. (a) T-RFLP from the unincubated sediment; (b) freshwater medium containing added Fe(II) as electron donor, NO3− as terminal electron acceptor and CO2 as carbon source; (c) as (b), but without added Fe(II) and NO3−; (d) as (b), but without added NO3−; (e) as (b), but without added Fe(II); (f) as (b), but with acetate instead of CO2 as carbon source.
Characterization of the Actinobacteria enrichment culture
An enrichment of the TM3 Actinobacteria could be reproducibly obtained by inoculation of the sediment into the enrichment medium. Dilution series done from the enrichment showed the presence of Actinobacteria and the oxidation of Fe(II) until the 10−4 dilution, beyond which neither the growth of Actinobacteria nor Fe(II) oxidation was observed. Modifying the medium, for example by omitting the phosphate buffer to avoid formation of white precipitate (presumably iron phosphates), or the addition of EDTA-chelated Fe(II) to simulate the chelation by humic acids, failed to enrich the TM3 Actinobacteria.
The ratio of Fe(II) oxidized to nitrate reduced by a 10−2 dilution of the enrichment was determined (Figure 4). Both Fe(II) and nitrate were consumed during the incubation with a molar ratio of Fe(II) to nitrate of 1:0.23 at each time point measured. No N2O production was observed in these incubations. Uninoculated and killed controls did not show Fe(II) or nitrate consumption. The T-RFLP profiling done at all time points showed the presence of the single dominant 146 bp T-RF.
Estimation of TM3 Actinobacteria abundance in sediment
Cloning and Sanger sequencing as well as pyrosequencing of 16S rRNA genes from Lake Grosse Fuchskuhle sediment was performed to estimate the abundance of TM3 Actinobacteria in the sediment. Sequencing was performed from four samples taken from four different zones: SW littoral (SWL), SW profundal (SWP), NE littoral (NEL) and NE profundal (NEP). Ninety-six clone sequences from each sample were obtained and none were found to match closely to the TM3 Actinobacteria. Many of the sequences had T-RFs of 146 bp, but these did not belong to Actinobacteria (results not shown). Sequences related to TM3 Actinobacteria were obtained from each sample by pyrosequencing: 11 of 1047 sequences from SWL; 23 of 3444 sequences from SWP; 6 of 2743 sequences from NEL; and 10 of 2369 sequences from NEP sediment. Assuming average 16S rRNA gene copy numbers of these organisms compared with the other community members, this corresponds to an average abundance of 0.57% of bacteria in the sediment. Quantification of total bacterial 16S rRNA genes from SWL, SWP and NEP sediment was performed by real-time PCR and found to be 6.44 × 108 (±2.35 × 108), 1.97 × 108 (±3.99 × 107) and 7.53 × 107 (±5.55 × 107) per gram of fresh sediment, respectively. Assuming the TM3 Actinobacteria sequences are 0.57% of the total, this would correspond to between 4.3 × 105 and 3.7 × 106 TM3 actinobacterial 16S rRNA genes per gram of fresh sediment.
SIP
DNA-based SIP was performed to determine if the TM3 Actinobacteria obtained in the enrichment cultures could be detected by SIP in the sediment using 13CO2. The sediment was incubated with added CO2 in the headspace with no additional supply of nutrients to the sediment samples. A time-course was performed and subsamples of the sediment were taken after 3, 6 and 12 weeks. Volatile fatty acid analysis by HPLC never showed an accumulation of volatile fatty acids and the incubations showed a delayed initiation of methanogenesis (∼10 weeks), which subsequently proceeded at a low rate (results not shown).
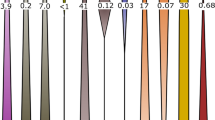
The DNA was isolated from the samples taken at 3-, 6- and 12-weeks and subjected to cesium chloride centrifugation. The quantification of bacterial 16S rRNA genes from the SIP-gradient fractions showed the largest abundance in the light-gradient fractions corresponding to densities between 1.71 and 1.75 g ml−1 in both the labeled and control incubations (Figure 5). Relatively low copy numbers of 16S rRNA genes were observed in the heavy-gradient fractions ranging from 1.8 × 105 to 2.2 × 106 in labeled CO2 incubations and 1.6 × 103 to 5.0 × 104 in the corresponding gradient fractions of the controls. A similar trend was observed at all the time points, indicating that the rate of growth and labeling by the autotrophic microorganisms was very low. T-RFLP fingerprinting was performed and peaks of 69 bp, 119 bp, 146 bp and 487 bp were found in heavy fractions only in the incubations with 13CO2 and unlabeled CO2 (Figure 5). A more diverse T-RFLP profile was observed from the light-gradient fractions. A similar pattern was found after 3-, 6- and 12-weeks (results not shown).
T-RFLP fingerprinting of density resolved bacterial 16S rRNA from the 6-week time point. (a) Incubation with 13CO2 and (b) control with unlabeled CO2. The densities of the fractions are given in right top of each fraction as g ml−1 values. The fractions with densities greater than 1.76 g ml−1 were considered heavy. The sizes of the major T-RFs are indicated.
Cloning and sequencing of 16S rRNA genes were performed from light- and heavy-gradient fractions. Clones from the heavy fraction with T-RF sizes corresponding to the 119 bp, 146 bp and 487 bp fragments observed in the T-RFLP were obtained (Figure 6). The clones corresponding to 119 bp T-RFs were grouped among sequences from Gallionella and Sideroxydans. The clones characterized by a 146 bp T-RFs were clustered with the Actinobacteria TM3 sequences obtained from the enrichment cultures and those characterized by a 487 bp T-RF were grouped among sequences from the genus Thiomonas. A more diverse set of sequences were obtained from the light-gradient fractions, including Planctomyces, Verrucomicrobia and Chloroflexi, which are all previously reported to be abundant in peat bogs (Dedysh et al., 2006; Kulichevskaya et al., 2006, 2007). Several Methylocystis clones were obtained from the light fractions and also had a T-RF length of 146 bp.
A similar T-RFLP analysis of archaeal 16S rRNA gene did not show differences between light and heavy fractions, indicating that Archaea were not labeled during the SIP incubation (results not shown).
Discussion
Previous studies on the profundal sediment of Lake Grosse Fuchskuhle have shown a relatively low rate of methanogenesis and a less diverse methanogenic archaeal community compared with other freshwater lakes (Chan et al., 2002; Casper et al., 2003; Conrad et al., 2010). A similar trend was observed in this study with the littoral sediment showing a relatively poor methanogenic potential (Figure 1). The low-rate of methanogenesis in this sediment might be attributed to favorable conditions for Fe(III) reduction. Indeed, the iron depth-profile of the sediment (Figure 1) showed the presence of biologically available Fe(III) in the top 15 cm of the sediment at concentrations sufficient to suppress methanogenesis (Roden and Wetzel, 1996). A maximum rate of methanogenesis was found at the sediment depth of 15–20 cm, which is similar to an earlier study (Chan et al., 2002) and might be attributed to a relatively high concentration of Fe(II), which is known to be capable of inhibiting Fe(III) reduction (Roden and Zachara, 1996; Roden and Urrutia, 1999). Although our results have suggested that Fe(III) reduction could be having a role in the mineralization of sediment organic matter, the long-term sustainment of Fe(III) reduction in anoxic zones requires a continuous recycling of Fe(III) from Fe(II) by anaerobic Fe(II)-oxidizing bacteria.
In order to isolate and characterize the chemolithotrophic Fe(II)-oxidizing bacteria in Lake Grosse Fuchskuhle, we performed incubations according to the procedure described by Straub and Buchholz-Cleven (1998), which resulted in the enrichment of TM3 group of uncultured Actinobacteria, phylogenetically close to the sequences previously reported from the same basin of this lake (Glöckner et al., 2000). Previous culture-independent studies have revealed the presence of deeply branching phylogenetic groups of Actinobacteria from various terrestrial and marine environments (Liesack and Stackebrandt, 1992; Colquhoun et al., 1998; Wohl and McArthur, 1998). These actinobacterial groups from soil have been designated as TM, forming three major clusters TM1, TM2 and TM3 (Rheims et al., 1996). Actinobacteria belonging to groups TM2 and TM3 in particular have been reported from various environments worldwide and with a greater abundance in low-pH environments such as peat bogs (Rheims et al., 1999). Due to their ubiquitous distribution, they are believed to be contributing to ecologically important processes (Felske et al., 1997; Rheims et al., 1999). Studies have shown that these Actinobacteria are metabolically active (Felske et al., 1997), slow-growing and easily overgrown under enrichment conditions (Rheims et al., 1999). Cultivation attempts have led to the isolation of representatives related to the TM2 group, namely Ferrithrix thermotolerans (Johnson et al., 2009), Ferrimicrobium acidiphilum (Johnson et al., 2009) and Acidimicrobium ferrooxidans (Clark and Norris, 1996), all of which are autotrophic iron-oxidizers; however, until now the TM3 Actinobacteria lacked cultivated representatives.
Characterization of the TM3 group Actinobacteria by incubations with different combinations of Fe(II), nitrate, acetate and CO2 showed that a combination of Fe(II), nitrate and CO2 were required for growth. Repeated attempts to isolate the Actinobacteria in pure culture by serial dilution of the enrichment neither showed growth nor a reduction in Fe(II) in the dilutions greater than 10−4. Other attempts to modify the medium like using EDTA-chelated Fe(II) or removal of phosphate buffer, to prevent the formation of white precipitate, did not lead to an enrichment of the Actinobacteria. The role of ferrous phosphates on the growth of Fe(II)-oxidizing bacteria is currently unknown, however, a similar phenomenon was reported earlier (Straub et al., 2004).
A time-course of Fe(II) oxidation and nitrate consumption performed with the actinobacterial enrichment was in good agreement with the expected stoichiometry required for nitrate-dependent Fe(II) oxidation (Figure 4). The molar ratio of Fe(II) to nitrate consumed was 1.00:0.23 at all time points compared with the ideal ratio of 1.00:0.20. This slightly higher ratio of Fe(II) to nitrate consumed could be due to utilization of some nitrate for growth. Abiotic Fe(II) oxidation with nitrate, nitrite or nitrous oxide would not be possible under the incubation conditions due to acidic pH and absence of copper in high concentrations (Buresh and Moraghan, 1976; Langmuir, 1997). Denitrification and abiotic oxidation of Fe(II) by nitrate was unlikely because of the lack of any nitrous oxide in the incubations. No accumulation of ammonia was observed indicating no reduction of nitrate to ammonia. These results indicate that the TM3 actinobacterial enrichment was capable of Fe(II) oxidation coupled to nitrate reduction.
The quantification of nitrate-dependent Fe(II)-oxidizing microorganisms in the sediment could not be done by molecular methods due to the absence of a suitable functional marker gene or 16S rRNA primers for this group. For this reason, an MPN method was used and the results showed the presence of 1 × 104 autotrophic and 1 × 107 heterotrophic nitrate-dependent Fe(II) oxidizers per gram fresh weight of sediment. These results are in accordance with previous findings that the heterotrophic outnumber the autotrophic Fe(II)-oxidizers by several orders of magnitude in lake sediments (Straub and Buchholz-Cleven, 1998b; Hauck et al., 2001; Muehe et al., 2009); however, the quantification by this approach will be an underestimation due to the selective nature of enrichment media and incubation conditions. The different microbial community composition in both autotrophic and heterotrophic MPN tubes suggest that these processes are mediated by different groups of organisms.
In addition to MPN assays, we used a combination of real-time PCR and amplicon pyrosequencing to quantify TM3 Actinobacteria in Lake Grosse Fuchskuhle sediment. This indicated TM3 group Actinobacteria 16S rRNA genes could account for ∼0.6% of total bacterial sequences in the lake sediment. On the basis of the comparison of these results with the bacterial quantifications by real-time PCR, the number of these Actinobacteria would correspond to 106 cells per gram of wet sediment. According to the rate of nitrate-dependent Fe(II)-oxidation per cell estimated by Melton et al. (2012), these organisms could be contributing significantly to the regeneration of Fe(III) in the sediment.
SIP under anoxic conditions using 13CO2 was performed in order to investigate the potential importance of TM3 Actinobacteria in situ. The same Actinobacteria TM3 organisms that were obtained in the enrichment cultures were labeled in the 13CO2 SIP incubations, indicating that these are important autotrophic organisms in this sediment. In addition, organisms related to Gallionella and Sideroxydans species as well as Thiomonas were labeled. The fact that the T-RFLP profiles in the heavy fractions did not change over the time-course suggests that cross-feeding of the 13C did not occur, which is a common phenomenon observed in many SIP studies involving long incubation times (Lueders et al., 2004; Qiu et al., 2009). Probably, the relatively low abundance of labeled organisms was insufficient to provide carbon to the community and observe cross-feeding of heterotrophic organisms. The quantification of bacterial 16S rRNA genes from the SIP-gradient fractions showed a large abundance in the light fractions compared with the heavy-gradient fractions. A similar trend was observed in all the time points, indicating the labeling of a relatively small fraction of the sediment microbial community and a slow rate of growth of these organisms.
Gallionella were the first microorganisms shown to be Fe(II) oxidizers (Ehrenberg, 1836). They are known to be prevalent in groundwater systems and mineral springs (Hanert, 2006) and are capable of both autotrophic and mixotrophic growth (Hallbeck and Pedersen, 1991). Sideroxydans spp. are prevalent in iron-rich environments and prefer microaerophilic conditions (Emerson and Moyer, 1997; Weiss et al., 2007); furthermore, they have been detected in acidic peatlands indicating that some species are acidophilic or acid-tolerant (Lüdecke et al., 2010). A previous study indicated that Gallionella ferruginea could perform autotrophic nitrate-dependent Fe(II) oxidation (Gouy et al., 1984), but no further studies with Gallionella have investigated this capability; however, studies have shown a complex distribution of Gallionella in relationship to the redox zonation in wetland soils (Wang et al., 2009), which is consistent with the existence of anaerobic phylotypes.
Members of genus Thiomonas are metabolically versatile with autotrophic, mixotrophic and heterotrophic physiologies and are capable of deriving energy by oxidation of reduced inorganic sulfur compounds or As(III) (Battaglia-Brunet et al., 2002; Duquesne et al., 2007). They are common inhabitants of extreme environments such as acid mine drainage, which have a low pH and high concentrations of sulfur and metals, such as iron and arsenite (Bruneel et al., 2003; Duquesne et al., 2008). The labeling of Thiomonas in our SIP experiment indicates that they also might be important anaerobic autotrophs in Lake Grosse Fuchskuhle sediment.
The fast growth of TM3 Actinobacteria under enrichment conditions compared with the SIP incubations could be due to the higher concentration of nitrate provided in the enrichments compared with that naturally present in the sediment. Our measurements have shown the presence of a low concentration of nitrate in the sediment, but the source of nitrate is not clear. Studies have indicated a reduced rate of denitrification (Nagele and Conrad, 1990; Šimek and Cooper, 2002) and a longer retention of nitrate (Müller et al., 1980) in low-pH environments. Moreover, ammonia oxidation within the rhizosphere of macrophytes could be a continuous source in the top 10 cm of the sediment (Herrmann et al., 2009). Due to these factors, we assume that there could be a continuous supply of nitrate in low concentrations to facilitate nitrate-dependent Fe(II) oxidation in Lake Grosse Fuchskuhle.
In summary, our results have indicated the chemolithotrophic nitrate-dependent Fe(II)-oxidizing nature of the TM3 group of uncultured soil Actinobacteria, which are widely distributed and whose function was previously unknown. The labeling of species related to the genera Gallionella, Sideroxydans and Thiomonas, which are capable of Fe(II) oxidation, suggests that Fe(II) oxidation is an important process in Lake Grosse Fuchskuhle sediment and may explain the relatively low-methanogenic potential of this lake. Although TM3 Actinobacteria may represent less than 1% of the bacterial community in the sediment, they could be of great functional significance in this environment due to their contribution to the regeneration of Fe(III) that is essential for iron-reducing microorganisms, which have an important role in the mineralization of sediment organic matter.
References
Battaglia-Brunet F, Dictor MC, Garrido F, Crouzet C, Morin D, Dekeyser K . (2002). An arsenic(III)-oxidizing bacterial population: selection, characterization, and performance in reactors. J Appl Microbiol 93: 656–667.
Benz M, Brune A, Schink B . (1998). Anaerobic and aerobic oxidation of ferrous iron at neutral pH by chemoheterotrophic nitrate-reducing bacteria. Arch Microbiol 169: 159–165.
Bruneel O, Personne JC, Casiot C, Leblanc M, Elbaz-Poulichet F, Mahler BJ . (2003). Mediation of arsenic oxidation by Thiomonas sp. in acid-mine drainage (Carnoulès, France). J Appl Microbiol 95: 492–499.
Buresh RJ, Moraghan JT . (1976). Chemical reduction of nitrate by ferrous iron. J Environ Quality 5: 320–325.
Casper P, Chan OC, Furtado ALS, Adams DD . (2003). Methane in an acidic bog lake: the influence of peat in the catchment on the biogeochemistry of methane. Aquat Sci 65: 36–46.
Chan OC, Wolf M, Hepperle D, Casper P . (2002). Methanogenic archaeal community in the sediment of an artificially partitioned acidic bog lake. FEMS Microbiol Ecol 42: 119–129.
Clark DA, Norris PR . (1996). Acidimicrobium ferrooxidans gen. nov. sp. nov.: mixed culture ferrous iron oxidation with Sulfobacillus species. Microbiology 141: 785–790.
Colquhoun JA, Heald SC, Li L, Tamaoka J, Kato C, Horikoshi K et al (1998). Taxonomy and biotransformation activities of some deep-sea actinomycetes. Extremophiles 2: 269–277.
Conrad R, Claus P, Casper P . (2010). Stable isotope fractionation during the methanogenic degradation of organic matter in the sediment of an acidic bog lake, Lake Grosse Fuchskuhle. Limnol Oceanogr 55: 1932–1942.
Dedysh SN, Pankratov TA, Belova SE, Kulichevskaya IS, Liesack W . (2006). Phylogenetic analysis and in situ identification of Bacteria community composition in an acidic Sphagnum peat bog. Appl Environ Microbiol 72: 2110–2117.
Duquesne K, Lieutaud A, Ratouchniak J, Muller D, Lett MC, Bonnefoy V . (2008). Arsenite oxidation by a chemoautotrophic moderately acidophilic Thiomonas sp. from the strain isolation to the gene study. Environ Microbiol 10: 228–237.
Duquesne K, Lieutaud A, Ratouchniak J, Yarzabal A, Bonnefoy V . (2007). Mechanisms of arsenite elimination by Thiomonas sp. isolated from Carnoulès acid mine drainage. Eur J Soil Biol 43: 351–355.
Edwards KJ, Bach W, McCollom TM, Rogers DR . (2004). Neutrophilic iron-oxidizing bacteria in the ocean: their habitats, diversity, and roles in mineral deposition, rock alteration, and biomass production in the deep-sea. Geomicrobiol J 21: 393–404.
Ehrenberg CG . (1836). Vorläufige Mitteilungen über das wirkliche Vorkommen fossiler Infusorien und ihre große Verbreitung. Poggendorff’s Ann Phys Chem 38: 213–227.
Emerson D, Moyer C . (1997). Isolation and characterization of novel iron-oxidizing bacteria that grow at circumneutral pH. Appl Environ Microbiol 63: 4784–4792.
Emerson D, Revsbech NP . (1994). Investigation of an iron-oxidizing microbial mat community located near Aarhus, Denmark—laboratory studies. Appl Environ Microbiol 60: 4032–4038.
Felske A, Rheims H, Wolterink A, Stackebrandt E, Akkermans ADL . (1997). Ribosome analysis reveals prominent activity of an uncultured member of the class Actinobacteria in grassland soils. Microbiology 143: 2983–2989.
Frenzel P, Bosse U, Janssen PH . (1999). Rice roots and methanogenesis in a paddy soil: ferric iron as an alternative electron acceptor in the rooted soil. Soil Biol Biochem 31: 421–430.
Glöckner FO, Zaichikov E, Belkova N, Denissova L, Pernthaler J, Pernthaler A et al (2000). Comparative 16S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria. Appl Environ Microbiol 66: 5053–5065.
Gouy J-L, Bergé P, Labroue L . (1984). Gallionella ferruginea, facteur de dénitrification dans les eaux pauvres en matière organique. C R Acad Sci Paris 298: 153–156.
Hafenbradl D, Keller M, Dirmeier R, Rachel R, Rossnagel P, Burggraf S et al (1996). Ferroglobus placidus gen. nov., sp. nov., a novel hyperthermophilic archaeum that oxidizes Fe2+ at neutral pH under anoxic conditions. Arch Microbiol 166: 308–314.
Hallbeck L, Pedersen K . (1991). Autotrophic and mixotrophic growth of Gallionella ferruginea. J Gen Microbiol 137: 2657–2661.
Hanert HH . (2006). The genus Gallionella. The Prokaryotes 7: 990–995.
Hart SC, Stark JM, Davidson EA, Firestone MK . (1994). Nitrogen mineralization, immobilization, and nitrification. In Weaver RW, Angle S, Bottomley P, Bezdiecek D, Smith S, Tabatabai A, Wollum A, (eds) Methods of Soil Analysis: Part 2. Microbiological and Biochemical Properties. Soil Science Society of America: Madison, WI, USA, Book series no. 5 pp 985–1018.
Hauck S, Benz M, Brune A, Schink B . (2001). Ferrous iron oxidation by denitrifying bacteria in profundal sediments of a deep lake (Lake Constance). FEMS Microbiol Ecol 37: 127–134.
Herrmann M, Saunders AM, Schramm A . (2009). Effect of lake trophic status and rooted macrophytes on community composition and abundance of ammonia-oxidizing prokaryotes in freshwater sediments. Appl Environ Microbiol 75: 3127–3136.
Johnson DB, Bacelar-Nicolau P, Okibe N, Thomas A, Hallberg KB . (2009). Ferrimicrobium acidiphilum gen. nov., sp. nov. and Ferrithrix thermotolerans gen. nov., sp. nov.: heterotrophic, iron-oxidizing, extremely acidophilic actinobacteria. Int J Syst Evol Microbiol 59: 1082–1089.
Kandeler E, Gerber H . (1988). Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol Fertil Soils 6: 68–72.
Koschel R . (1995). Manipulation of whole-lake ecosystems and long-term limnological observations in the Brandenburg—Mecklenburg lake district, Germany. Int Rev Ges Hydrobiol 80: 507–518.
Kulichevskaya IS, Belova SE, Kevbrin VV, Dedysh SN, Zavarzin GA . (2007). Analysis of the bacterial community developing in the course of Sphagnum moss decomposition. Microbiology 76: 621–629.
Kulichevskaya IS, Pankratov TA, Dedysh SN . (2006). Detection of representatives of the Planctomycetes in Sphagnum peat bogs by molecular and cultivation approaches. Microbiology 75: 329–335.
Köhler T, Dietrich C, Scheffrahn RH, Brune A . (2012). High-resolution analysis of gut environment and bacterial microbiota reveals functional compartmentation of the gut in wood-feeding higher termites (Nasutitermes spp.). Appl Environ Microbiol 78: 4691–46701.
Küsel K, Blothe M, Schulz D, Reiche M, Drake HL . (2008). Microbial reduction of iron and porewater biogeochemistry in acidic peatlands. Biogeosciences 5: 1537–1549.
Langmuir D . (1997) Aqueous Environmental Geochemistry. Prentice-Hall: Englewood Cliffs, NJ, USA.
Liesack W, Stackebrandt E . (1992). Occurrence of novel groups of the domain Bacteria as revealed by analysis of genetic material isolated from an Australian terrestrial environment. J Bacteriol 75: 5072–5078.
Lu S, Gischkat S, Reiche M, Akob DM, Hallberg KB, Küsel K . (2010). Ecophysiology of Fe-cycling bacteria in acidic sediments. Appl Environ Microbiol 76: 8174–8183.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Lueders T, Wagner B, Claus P, Friedrich MW . (2004). Stable isotope probing of rRNA and DNA reveals a dynamic methylotroph community and trophic interactions with fungi and protozoa in oxic rice field soil. Environ Microbiol 6: 60–72.
Lüdecke C, Reiche M, Eusterhues K, Nietzsche S, Küsel K . (2010). Acid-tolerant microaerophilic Fe(II)-oxidizing bacteria promote Fe(III)-accumulation in a fen. Environ Microbiol 12: 2814–2825.
Melton ED, Schmidt C, Kappler A . (2012). Microbial Fe(II) oxidation in littoral freshwater lake sediment: the potential for competition between phototrophic vs. nitrate-dependent iron(II)-oxidizers. Front Microbiol 3: 1–12.
Muehe EM, Gerhardt S, Schink B, Kappler A . (2009). Ecophysiology and the energetic benefit of mixotrophic Fe(II) oxidation by various strains of nitrate-reducing bacteria. FEMS Microbiol Ecol 70: 335–343.
Müller MM, Sundman V, Skujins J . (1980). Denitrification in low pH spodosols and peats determined with the acetylene inhibition method. Appl Environ Microbiol 40: 235–239.
Nagele W, Conrad R . (1990). Influence of soil-pH on the nitrate-reducing microbial populations and their potential to reduce nitrate to NO and N2O. FEMS Microbiol Ecol 74: 49–57.
Neubauer SC, Toledo-Duran GE, Emerson D, Megonigal JP . (2007). Returning to their roots: iron-oxidizing bacteria enhance short-term plaque formation in the wetland-plant rhizosphere. Geomicrobiol J 24: 65–73.
Ponnamperuma FN . (1972). The chemistry of submerged soils. Adv Agron 24: 29–96.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J et al (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Qiu Q, Conrad R, Lu Y . (2009). Cross-feeding of methane carbon among bacteria on rice roots revealed by DNA-stable isotope probing. Environ Microbiol Rep 1: 355–361.
Reiche M, Torburg G, Küsel K . (2008). Competition of Fe(III) reduction and methanogenesis in an acidic fen. FEMS Microbiol Ecol 65: 88–101.
Rheims H, Felske A, Seufert S, Stackebrandt E . (1999). Molecular monitoring of an uncultured group of the class Actinobacteria in two terrestrial environments. J Microbiol Methods 36: 65–75.
Rheims H, Sproer C, Rainey FA, Stackebrandt E . (1996). Molecular biological evidence for the occurrence of uncultured members of the actinomycete line of descent in different environments and geographical locations. Microbiology 142: 2863–2870.
Roden EE, Urrutia MM . (1999). Ferrous iron removal promotes microbial reduction of crystalline iron(III) oxides. Environ Sci Technol 33: 1847–1853.
Roden EE, Wetzel RG . (1996). Organic carbon oxidation and suppression of methane production by microbial Fe(III) oxide reduction in vegetated and unvegetated freshwater wetland sediments. Limnol Oceanogr 41: 1733–1748.
Roden EE, Zachara JM . (1996). Microbial reduction of crystalline iron(III) oxides: influence of oxide surface area and potential for cell growth. Environ Sci Technol 30: 1618–1628.
Sachse A, Babenzien D, Ginzel G, Gelbrecht J, Steinberg CEW . (2001). Characterization of dissolved organic carbon (DOC) of a dystrophic lake and an adjacent fen. Biogeochem 54: 279–296.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Steinmann P, Shotyk W . (1997). Chemical composition, pH, and redox state of sulfur and iron in complete vertical porewater profiles from two Sphagnum peat bogs, Jura Mountains, Switzerland. Geochim Cosmochim Acta 61: 1143–1163.
Stookey LL . (1970). Ferrozine—A new spectrophotometric reagent for iron. Anal Chem 42: 779–781.
Straub KL, Benz M, Schink B, Widdel F . (1996). Anaerobic, nitrate-dependent microbial oxidation of ferrous iron. Appl Environ Microbiol 62: 1458–1460.
Straub KL, Buchholz-Cleven BEE . (1998). Enumeration and detection of anaerobic ferrous iron-oxidizing, nitrate-reducing bacteria from diverse European sediments. Appl Environ Microbiol 64: 4846–4856.
Straub KL, Schonhuber WA, Buchholz-Cleven BEE, Schink B . (2004). Diversity of ferrous iron-oxidizing, nitrate-reducing bacteria and their involvement in oxygen-independent iron cycling. Geomicrobiol J 21: 371–378.
Stubner S . (2002). Enumeration of 16S rDNA of Desulfotomaculum lineage 1 in rice field soil by real-time PCR with SybrGreen detection. J Microbiol Methods 50: 155–164.
Stumm W, Morgan JJ . (1996) Aquatic Chemistry: Chemical Equilibria and Rates in Natural Waters. John Wiley & Sons: New York, NY, USA.
Theis TL, Singer PC . (1974). Complexation of iron(II) by organic matter and its effect on iron(II) oxygenation. Environ Sci Technol 8: 569–573.
Wang J, Muyzer G, Bodelier PLE, Laanbroek HJ . (2009). Diversity of iron oxidizers in wetland soils revealed by novel 16S rRNA primers targeting Gallionella-related bacteria. ISME J 3: 715–725.
Weber KA, Pollock J, Cole KA, O’Connor SM, Achenbach LA, Coates JD . (2006). Anaerobic nitrate-dependent iron(II) bio-oxidation by a novel lithoautotrophic betaproteobacterium, strain 2002. Appl Environ Microbiol 72: 686–694.
Weiss JV, Rentz JA, Plaia T, Neubauer SC, Merrill-Floyd M, Lilburn et al (2007). Characterization of neutrophilic Fe(II)-oxidizing bacteria isolated from the rhizosphere of wetland plants and description of Ferritrophicum radicicola gen. nov. sp. nov., and Sideroxydans paludicola sp. nov. Geomicrobiol J 24: 559–570.
Widdel F, Bak F . (1992). Gram-negative mesophilic sulfate reducing bacteria. In Balows A, Trüper HG, Dworkin M, Harder W, Schleifer K-H, (eds) In The Prokaryotes 2nd edn Springer: New York, pp 3352–3378.
Wohl DL, McArthur JV . (1998). Actinomycete-flora associated with submersed freshwater macrophytes. FEMS Microbiol Ecol 26: 135–140.
Šimek M, Cooper JE . (2002). The influence of soil pH on denitrification: progress towards the understanding of this interaction over the last 50 years. Eur J Soil Sci 53: 345–354.
Acknowledgements
This study was funded by the Max Planck Society (DK, BP and MGD) and a fellowship from the International Max Planck Research School for Environmental, Cellular and Molecular Microbiology (DK). The authors gratefully acknowledge Professor Ralf Conrad for his advice throughout the work and for critical reading of the manuscript. We also gratefully acknowledge Dr Werner Liesack for helpful discussions and Peter Claus, Markus Drescher, Melanie Klose, Michael Sachtleben and Roman Degebrodt for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kanaparthi, D., Pommerenke, B., Casper, P. et al. Chemolithotrophic nitrate-dependent Fe(II)-oxidizing nature of actinobacterial subdivision lineage TM3. ISME J 7, 1582–1594 (2013). https://doi.org/10.1038/ismej.2013.38
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.38
Keywords
This article is cited by
-
The reproduction process of Gram-positive protocells
Scientific Reports (2024)
-
Inter-phylum negative interactions affect soil bacterial community dynamics and functions during soybean development under long-term nitrogen fertilization
Stress Biology (2021)
-
Influence of pH, EDTA/Fe(II) ratio, and microbial culture on Fe(II)-mediated autotrophic denitrification
Environmental Science and Pollution Research (2017)
-
Characteristics of the cultivable bacteria from sediments associated with two deep-sea hydrothermal vents in Okinawa Trough
World Journal of Microbiology and Biotechnology (2015)