Abstract
A Gram-reaction-positive, non-motile, endophytic actinomycete, designated strain YIM 63235T, was isolated from the surface-sterilized stems of Artemisia annua L., and characterized to determine its taxonomic position. The strain YIM 63235T formed well-differentiated aerial and substrate mycelia on media tested. The phylogenetic tree based on 16S rRNA gene sequences showed that the new isolate formed a distinct lineage within the genus Pseudonocardia, and the strain YIM 63235T was closely related to Pseudonocardia parietis 04-St-002T (99.1%). However, DNA–DNA relatedness demonstrated that strain YIM 63235T was distinct from the closest phylogenetic neighbor. The chemotaxonomic properties of strain YIM 63235T were consistent with those of the genus Pseudonocardia: the diagnostic diamino acid of the cell-wall peptidoglycan was meso-diaminopimelic acid and MK-8(H4) was the predominant menaquinone. The major fatty acids were iso-C16:0 and iso-C16:1 H. The DNA G+C content of strain YIM 63235T was 71.0 mol%. On the basis of the phenotypic and phylogenetic distinctiveness, the novel isolate was identified as representing a novel species of the genus Pseudonocardia, for which the name Pseudonocardia antimicrobica sp. nov. (type strain YIM 63235T =CCTCC AA 208080T=DSM 45303T) is proposed.
Similar content being viewed by others
Introduction
Endophytic bacteria can be defined as those bacteria that colonize the internal tissue of the plant showing no external sign of infection or negative effect on their host. There are more than 300 000 plant species on the earth, and each individual plant is host to one or more endophytes.1 However, only a few of these plants have ever been studied completely relative to their endophytic biology. Consequently, the opportunity to find novel and beneficial endophytic micro-organisms among the diversity of plants in different ecosystems is considerable.2 As part of our long-term study on endophytic actinomcete diversity and bioactive metabolites isolated from tropical rainforest medicinal plants of Xishuangbanna, several novel species have been characterized: Dietzia schimae and Dietzia cercidiphylli,3 Plantactinospora mayteni,4 Pseudonocardia artemisiae5 and Streptomyces artemisiae.6 In this report, the description of the morphological, physiological, chemotaxonomic and phylogenetic characteristics of a Pseudonocardia-like strain YIM 63235T is presented. Phenotypic and genotypic data show that the isolate YIM 63235T represents a novel species of the genus Pseudonocardia, for which the name Pseudonocardia antimicrobica sp. nov. is proposed.
The genus Pseudonocardia within the family Pseudonocardiaceae was first described by Henssen,7 and since then the description of the genus has been revised repeatedly.8, 9, 10, 11, 12 Members of the genus Pseudonocardia displayed vegetative and aerial mycelium with spore chains produced by acropetal budding or fragmentation, type IV cell wall, major menaquinone is MK-8 (H4) or MK-9 and a DNA G+C content of 68–79 mol%. Members of the genus Pseudonocardia have been widely reported and recovered from several ecosystems, such as active sludge soil (including those polluted by industrial chemicals) and plant samples (including stems, leaves, root nodules, tree-bark compost and traditional Chinese medicinal plants). At the time of writing, the genus Pseudonocardia encompasses 46 species with validly published names.13, 14
Materials and methods
Strain and culture conditions
Stem samples of Artemisia annua L. (sweet wormwood) were collected from Xishuangbanna and Kunming City in Yunnan Province, between August 2006 and April 2007. The samples were maintained at 4 °C and transported to the laboratory for immediate analysis. Samples were washed in running water to remove soil particles and sterilized by the established procedure.15 After being surface sterilized, the samples were sliced into pieces, followed by plating on tap water–yeast extract agar plates (containing 0.25 g of yeast extract, 0.5 g of K2HPO4 and 18 g of agar, per liter of tap water, pH 7.2) containing nalidixic acid (25 mg l−1), nystatin (50 mg l−1) and cycloheximide (50 mg l−1) to repress growth of bacteria and fungi. The plates were incubated at 28 °C for 4–8 weeks until the outgrowth of endophytic actinomycetes were discerned. Colonies originating from plant segments were picked up and pure cultures were obtained by repeated streaking on tap water–yeast extract agar plates. The purified strain YIM 63235T was picked and maintained on tryptic soy agar (containing 15 g of tryptone, 5 g of soya peptone, 5 g of NaCl and 15 g of agar, per liter of tap water, pH 7.2) slants at 4 °C and as 20% (w/v) glycerol suspensions at −80 °C.
Biomass for chemical and molecular studies was obtained by cultivation in shaken flasks (about 200 r.p.m.) using tryptic soy broth (containing 15 g of tryptone, 5 g of soya peptone and 5 g of NaCl, per liter of tap water, pH 7.2) medium at 28 °C for 1 week.
Phenotypic characterization
Morphological, cultural, physiological and biochemical characterization of the strain YIM 63235T was studied by following the guidelines of the International Streptomyces Project.16 The morphological characteristics were observed by light microscopy (BH2; Olympus, Japan) and scanning electron microscopy (Quanta 200; FEI, USA) using cultures grown on ISP 2 medium at 28 °C for 7–14 days. Cultural characteristics were recorded on ISP media (International Streptomyces Project), Czapek’s agar, potato–glucose agar and nutrient agar prepared as described by Dong and Cai.17 Cell motility, Gram staining, growth parameter (temperature range, pH range and NaCl tolerance), starch hydrolysis, nitrate reduction and oxidase activity were determined.18 The antimicrobial activities of strain YIM 63235T were investigated by using media containing Aspergillus niger, Bacillus subtilis and Escherichia coli.19
Chemotaxonomy
The isomer of diaminopimelic acid and sugar analysis of whole-cell hydrolysates were performed according to the procedures described by Hasegawa et al.,20 Lechevalier and Lechevalier21 and Tang et al.22 Phospholipids were extracted, examined by two-dimensional thin layer chromatography and identified using previously described procedures.23, 24 Mycolic acids were extracted and analyzed by one-dimensional thin layer chromatography described by Minnikin et al.25 Menaquinones were isolated according to Collins et al.26 and separated by HPLC.27 Cellular fatty acids were extracted, methylated and analyzed by using the Sherlock Microbial Identification System (MIDI) according to the manufacturer’s instructions. The fatty acid methyl esters were analyzed by using the Microbial Identification software package (Sherlock Version 4.0; MIDI database: TSBA40, MIDI Company, USA). The G+C content of the genomic DNA was determined by using the HPLC method28 with E. coli JM-109 as the reference strain.
Molecular analysis
Extraction of genomic DNA, PCR amplification and sequencing of the 16S rRNA gene were performed as described by Li et al.29 The phylogenetic neighbors were identified and pairwise16S rRNA gene sequence similarities were calculated using the EzTaxon-e server Database30 (http://eztaxon-e.ezbiocloud.net/). The almost-complete 16S rRNA gene sequence determined in this study was aligned with reference sequences of the genus Pseudonocardia by using the CLUSTAL_X program.31 The phylogenetic trees were constructed by the neighbor-joining,32 maximum-parsimony,33 minimum-evolution34 and maximum-likelihood35 tree-making algorithms by using the software packages MEGA version 4.036 and PHYML.37 The topologies of the phylogenetic trees were evaluated by using the bootstrap resampling method of Felsenstein38 with 1000 replicates. The strain Kutneria kofuensis NRRL B-24061T (AF114801) was used as the outgroup. DNA–DNA relatedness was studied according to the fluorometric micro-well method,39, 40, 41 and the hybridizations were performed with six replications.
The 16S rRNA gene sequence of strain YIM 63235T has been deposited in GenBank under the accession number FJ817380.
Results and Discussion
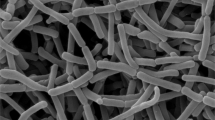
Strain YIM 63235T formed visible colonies within 5 days on ISP 2 incubated at 28 °C. Good growth occurred at 20–28 °C, which formed extensively branched substrate mycelia (orange–yellow/yellow–brown) and aerial mycelia (white) (Supplementary Table S1). Morphological observation of 7 day-old cultures of strain YIM 63235T revealed that both aerial and vegetative hyphae were abundant, well developed and fragmented into rod-shaped elements (Supplementary Figure S1). Yellowish-brown soluble pigment is produced on the potato–glucose agar. Cells were aerobic, non-motile, Gram-stain-positive and catalase-positive. Temperature range for growth is 10–42 °C, with optimal growth occurring at 20–28 °C. The pH range for growth is 5.0–9.0 (optimum, pH 7.0–8.0). The NaCl concentration range for growth is 0–10% (optimum, 0–5% NaCl, w/v). Other physiological and biochemical characteristics are summarized in the species description. Strain YIM 63235T was differentiated from Pseudonocardia parietis 04-St-002T by the oxidase reaction, degradation of Tween 20, Tween 80 and starch, utilization of D-cellobiose, lactose, D-raffinose, ribose, xylose, hypoxanthine and L-phenylalanine and the range of temperature/pH/NaCl tolerance of growth (Table 1). Strain YIM 63235T produced a substance that inhibited the growth of B. subtilis and E. coli.
Strain YIM 63235T was examined for chemical markers considered to be characteristic of Pseudonocardia strains. Strain YIM 63235T contained meso-diaminopimelic acid (meso-DAP) as the diagnostic diamino acid and the whole-cell hydrolysates were rich in glucose, arabinose, galactose and mannose (type IV cell wall). The quinone system of YIM 63235T was composed of menaquinones MK-8(H4) (94.5%) and MK-8(H2) (5.5%). Mycolic acids were absent. The predominant polar lipids contained of diphosphatidylglycerol, phosphatidylglycerol, phosphatidylmethylethanolamine, phosphatidylcholine, phosphatidylinositol, phosphatidylinositol mannoside and four unknown polar lipids (type PIII phospholipid, Supplementary Figure S2). The fatty acid profile of strain YIM 63235T contained saturated, unsaturated, 10-methyl and hydroxyl components, with the major fatty acids being iso-C16:0 (60.9%), iso-C16:1 H (22.7%) and 10-methyl-C17:0 (2.3%). The detailed cellular fatty acid profiles of strain YIM 63235T and P. parietis are given in Supplementary Table S2. Strain YIM 63235T could be distinguished easily from the type strain of P. parietis 04-St-002T based on the presence/absence and amount of C16:0, C18:0, C18:1ω9c, iso-C16:0, iso-C16:1 H, C16:0 10-methyl, C17:0 10-methyl and C16:1ω7c and C15:0 iso 2-OH. The DNA G+C content of strain YIM 65235T was 71.0 mol%.
An almost-complete 16S rRNA gene sequence of strain YIM 63235T (1399 nucleotide) was determined. Comparison of the sequence with those stored in GenBank indicated that strain YIM 63235T was a member of the genus Pseudonocardia, with which it shared 94.8–99.1% 16S rRNA gene sequence similarity. Strain YIM 63235T shared a 16S rRNA similarity of 99.1%, 97.9%, 97.9%, 97.9%, 97.8% and 97.8% with the type strains of P. parietis 04-St-002T, P. carboxydivorans Y8T, P. tropica YIM 61452T, P. ammonioxydans H9T, P. antarctica DVS 5a1T and P. alni DSM 44104T, respectively. Levels of the 16S rRNA gene sequence similarity between strain YIM 63235T and the other Pseudonocardia species were <97.0%. The phylogenetic tree constructed with 16S rRNA gene sequence data by neighbor-joining method (Figure 1 and Supplementary Figure S3) showed that strain YIM 63235T formed a monophyletic clade with P. parietis 04-St-002T, and which was supported by the maximum-parsimony (Supplementary Figure S4), minimum-evolution (Supplementary Figure S5) and maximum-likelihood method (Supplementary Figure S6). However, strain YIM 63235T did not form a cluster with strains P. carboxydivorans Y8T, P. tropica YIM 61452T, P. ammonioxydans H9T, P. antarctica DVS 5a1T and P. alni DSM 44104T in any of the four tree-making algorithms.
Neighbor-joining tree of Pseudonocardia antimicrobica YIM 63235T sp. nov. and related species based on 16S rRNA gene sequences. Bar, 0.002 substitutions per nucleotide position. Asterisks indicate branches that were also recovered using the maximum-parsimony, minimum-evolution and maximum-likelihood methods.
DNA–DNA relatedness value between strain YIM 63235T and the most closely type strain P. parietis 04-St-002T was determined using the fluorometric micro-well method under optimal hybridization conditions. The DNA–DNA relatedness study was not carried out between strain YIM 63235T and other phylogenetic relatives with 16S rRNA gene sequence similarities that were <98.0%. Strain YIM 63235T exhibited relatively low levels of DNA–DNA relatedness with respect to P. parietis 04-St-002T (32.3±2.0%), which is well below the 70% cutoff point recommended for the assignment of bacterial strains to the same genomic species.42 These data suggest that strain YIM 63235T represent a novel species of the genus Pseudonocardia.
The phenotypic properties of strain YIM 63235T and the 16S rRNA gene sequence comparison supported the classification of the isolate in the genus Pseudonocardia. Differentiating characteristics (Table 1), phylogenetic analysis of the 16S rRNA gene sequence and DNA–DNA relatedness distinguished strain YIM 63235T from other members of the genus Pseudonocardia. Therefore, strain YIM 63235T is proposed to represent a hitherto unrecognized species of the genus Pseudonocardia, with the name Pseudonocardia antimicrobica sp. nov.
Description of Pseudonocardia antimicrobica sp. nov.
Pseudonocardia antimicrobica (an.ti.mi.cro′bi.ca. Gr. prep. anti against; N.L. n. microbium microbe; L. adj. suff.-cus -a -um suffix used with various meanings; N.L.fem. adj. antimicrobica antimicrobial) is an aerobic, non-motile, Gram-positive actinomycete that forms extensively branched substrate mycelia (orange–yellow/yellow–brown) and aerial mycelia (white). It produces yellowish-brown soluble pigment on the potato–glucose agar. The temperature range for growth is 10–42 °C, with optimal growth occurring at 20–28 °C, and the pH range for growth is 5.0–9.0 (optimum, pH 7.0–8.0). The NaCl concentration range for growth is 0–10% (optimum, 0–5% NaCl, w/v). It is positive for catalase, milk coagulation and milk peptonization, but negative for nitrate reduction, oxidase, urease, gelatin liquefaction, cellulose and starch hydrolysis and H2S production. Tweens 20, 40 and 80 are hydrolyzed by it and it utilizes L-arabinose, D-cellobiose, D-fructose, D-galactose, glucose, maltose, D-mannitol, D-mannose, D-raffinose, L-rhamnose, ribose, D-sorbitol, sucrose and xylose as the sole carbon sources, whereas Dulcitol, glycerol, lactose, myo-inositol and sodium acetate are not utilized. L-alanine, L-arginine, L-asparagine, glycine, L-hydroxyproline, hypoxanthine, L-phenylalanine, L-serine, L-tyrosine, L-valine and xanthine can be used as sole nitrogen sources, but not L-lysine. Acid is produced from D-galactose and glucose and it shows antimicrobial activities against B. subtilis and E. coli. The cell wall of strain YIM 63235T contains meso-DAP. The whole-cell sugar pattern consists of glucose, arabinose, galactose and mannose (type IV cell wall). MK-8(H4) is the predominant menaquinone. Mycolic acids are absent. The phospholipids consist of diphosphatidylglycerol, phosphatidylglycerol, phosphatidylmethylethanolamine, phosphatidylcholine, phosphatidylinositol, phosphatidylinositol mannoside and four unknown polar lipids (type PIII phospholipid). The major fatty acids are iso-C16:0 (60.9%) and iso-C16:1 H (22.7%). The G+C content of genomic DNA is 71.0 mol%.
The type strain, YIM 63235T (=CCTCC AA 208080T=DSM 45303T), was isolated from surface-sterilized stems of Artemisia annua L. collected from Yunnan province, Southwest China.
Accession codes
References
Strobel, G., Daisy, B., Castillo, U. & Harper, J. Natural products from endophytic microorganisms. J. Nat. Prod. 67, 257–268 (2004).
Ryan, R. P., Germaine, K., Franks, A., Ryan, D. J. & Dowling, D. N. Bacterial endophytes: recent developments and applications. FEMS Microbiol. Lett. 278, 1–9 (2008).
Li, J. et al. Dietzia schimae sp. nov. and Dietzia cercidiphylli sp. nov., from surface-sterilized plant tissues. Int. J. Syst. Evol. Microbiol. 58, 2549–2554 (2008).
Qin, S. et al. Plantactinospora mayteni gen. nov., sp. nov., a member of the family Micromonosporaceae. Int. J. Syst. Evol. Microbiol. 59, 2527–2533 (2009).
Zhao, G. Z. et al. Pseudonocardia artemisiae sp. nov., a novel actinobacterium isolated from surface-sterilized Artemisia annua L. Int. J. Syst. Evol. Microbiol. 61, 1061–1065 (2011).
Zhao, G. Z. et al. Streptomyces artemisiae sp. nov., isolated from surface-sterilized tissue of Artemisia annua L. Int. J. Syst. Evol. Microbiol. 60, 27–32 (2010).
Henssen, A. Beiträge zur Morphologie und Systematic der thermophilen Actinomyceten. Arch. Mikrobiol. 26, 377–414 (1957).
Warwick, S., Bowen, T., McVeigh, H. P. & Embley, T. M. A phylogenetic analysis of the family Pseudonocardiaceae and the genera Actinokineospora and Saccharothrix with 16S rRNA sequences and a proposal to combine the genera Amycolata and Pseudonocardia in an emended genus Pseudonocardia. Int. J. Syst. Bacteriol. 44, 293–299 (1994).
McVeigh, H. P., Munro, J. & Embley, T. M. The phylogenetic position of Pseudoamycolata halophobica (Akimov et al. 1989) and a proposal to reclassify it as Pseudonocardia halophobica. Int. J. Syst. Bacteriol. 44, 300–302 (1994).
Reichert, K., Lipski, A., Pradella, S., Stackebrandt, E. & Altendorf, K. Pseudonocardia asaccharolytica sp. nov. and Pseudonocardiasulfidoxydans sp. nov., two new dimethyl disulfide-degrading actinomycetes and emended description of the genus Pseudonocardia. Int. J. Syst. Bacteriol. 48, 441–449 (1998).
Huang, Y. et al. Proposal to combine the genera Actinobispora and Pseudonocardia in an emended genus Pseudonocardia, and description of Pseudonocardia zijingensis sp. nov.,. Int. J. Syst. Evol. Microbiol. 52, 977–982 (2002).
Park, S. W., Park, S. T., Lee, J. E. & Kim, Y. M. Pseudonocardia carboxydivorans sp. nov., a carbon monoxide-oxidizing actinomycete, and an emended description of the genus Pseudonocardia. Int. J. Syst. Evol. Microbiol. 58, 2475–2478 (2008).
Nie, G. X. et al. Pseudonocardia yuanmoensis sp. nov., a novel actinobacterium isolated from soil in Yunnan, south-west China. Antonie van Leeuwenhoek 101, 753–760 (2012).
Tian, X. P. et al. Pseudonocardia antitumorica sp. nov., a new deoxynyboquinone producing actinomycete isolated from a deep-sea sedimental sample in South China Sea. Int. J. Syst. Evol. Microbiol. doi:10.1099/ijs.0.037135-0.
Li, J. et al. Antitumour and antimicrobial activities of endophytic streptomycetes from pharmaceutical plants in rainforest. Lett. Appl. Microbiol. 47, 574–580 (2008).
Shirling, E. B. & Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 16, 313–340 (1966).
Dong, X. Z. & Cai, M. Y. Manual of Systematics and Identification of General Bacteria, Science Press: Beijing, (2001).
Zhao, G. Z. et al. Pseudonocardia bannaensis sp. nov., a novel actinomycete isolated from the surface-sterilized roots of Artemisia annua L. Antonie van Leeuwenhoek 100, 35–42 (2011).
Hugo, W. B. & Russell, A. D. Pharmaceutical Microbiology. 3rd edn. (Blackwell: Oxford, 1983).
Hasegawa, T., Takizawa, M. & Tanida, S. A rapid analysis for chemical grouping of aerobic actinomycetes. J. Gen. Microbiol. 29, 319–322 (1983).
Lechevalier, M. P. & Lechevalier, H. Chemical composition as a criterion in the classification of aerobic actinomycetes. Int. J. Syst. Bacteriol. 20, 435–443 (1970).
Tang, S. K. et al. Zhihengliuella alba sp. nov., and emended description of the genus Zhihengliuella. Int. J. Syst. Evol. Microbiol. 59, 2025–2032 (2009).
Minnikin, D. E., Collins, M. D. & Goodfellow, M. Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J. Appl. Bacteriol. 47, 87–95 (1979).
Collins, M. D. & Jones, D. Lipids in the classification and identification of coryneform bacteria containing peptidoglycan based on 2, 4-diaminobutyric acid. J. Appl. Bacteriol. 48, 459–470 (1980).
Minnikin, D. E., Hutchinson, I. G., Caldicott, A. B. & Goodfellow, M. Thin layer chromatography of methanolysates of mycolic acid-containing bacteria. J. Chromatogr. 188, 221–233 (1980).
Collins, M. D., Pirouz, T., Goodfellow, M. & Minnikin, D. E. Distribution of menaquinones in actinomycetes and corynebacteria. J. Gen. Microbiol. 100, 221–230 (1977).
Tamaoka, J., Katayama-Fujimura, Y. & Kuraishi, H. Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J. Appl. Bacteriol. 300, 31–36 (1983).
Mesbah, M., Premachandran, U. & Whitman, W. B. Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int. J. Syst. Bacteriol. 39, 159–167 (1989).
Li, W. J. et al. Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China) and emended description of the genus Georgenia. Int. J. Syst. Evol. Microbiol. 57, 1424–1428 (2007).
Kim, O. S. et al. Introducing EzTaxon-e: a prokaryotic 16S rRNA Gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 62, 716–721 (2012).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic tree. Mol. Biol. Evol. 4, 406–425 (1987).
Fitch, W. M. Toward defining the course of evolution: minimum change for a specific tree topology. Syst. Zool 20, 406–416 (1971).
Rzhetsky, A. & Nei, M. A simple method for estimating and testing minimum evolution trees. Mol. Biol. Evol. 9, 945–967 (1992).
Felsenstein, J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007).
Guindon, S. & Gascuel, O. A simple, fast, and accurae algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–789 (1985).
Ezaki, T., Hashimoto, Y. & Yabuuchi, E. Fluorometric deoxyribonucleic acid-deoxyribonucleic acid hybridization in microdilution wells as an alternative to membrane filter hybridization in which radioisotopes are used to determine genetic relatedness among bacterial strains. Int. J. Syst. Bacteriol. 39, 224–229 (1989).
Christensen, H., Angen, O., Mutters, R., Olsen, J. E. & Bisgaard, M. DNA–DNA hybridization determined in micro-wells using covalent attachment of DNA. Int. J. Syst. Evol. Microbiol. 50, 1095–1102 (2000).
He, L. et al. Streptomyces jietaisiensis sp. nov., isolated from soil in northern China. Int. J. Syst. Evol. Microbiol. 55, 1939–1944 (2005).
Stackebrandt, E. & Goebel, B. M. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Bacteriol. 44, 846–849 (1994).
Acknowledgements
This research was supported by the National Basic Research Program of China (No. 2010CB833801), the National Natural Science Foundation of China (No. U0932601) and the Yunnan Provincial Sciences and Technology Department (Nos 2011FB034 and 14051453). W-J Li was also supported by ‘Hundred Talents Program’ of the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhao, GZ., Li, J., Qin, YL. et al. Pseudonocardia antimicrobica sp. nov., a novel endophytic actinomycete associated with Artemisia annua L. (sweet wormwood). J Antibiot 65, 469–472 (2012). https://doi.org/10.1038/ja.2012.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2012.56
Keywords
This article is cited by
-
Continuing hunt for endophytic actinomycetes as a source of novel biologically active metabolites
World Journal of Microbiology and Biotechnology (2015)