Abstract
A series of olivomycin A derivatives containing different combinations of the acyl residues in the carbohydrate chains was obtained. The formation of complexes of Mg2+-coordinated dimers of these compounds with double-stranded DNA was studied using spectral methods such as absorption, fluorescence and circular dichroism (CD) spectral analyses. There was a good correlation of the values of binding constants of complexes (antibiotic)2Mg2+–DNA, the quantum yields of fluorescence and changes of the induced CD spectra with topoisomerase I inhibition and cytotoxicity. We demonstrate that the presence of the acyl groups in the saccharide residues of olivomycin A derivatives is absolutely necessary for a high cytotoxic potency of these antibiotics. On the basis of the experimental results and quantum chemical calculations, we presume that the acyl residue in the 4-O-position in the A-sugar residue is involved, to the most part, in the antibiotic–antibiotic interactions in the (olivomycin)2Mg2+ dimers, whereas the O-acyl group in E-olivomicose residue largely participates in the formation of the (olivomycin)2Mg2+–DNA complexes.
Similar content being viewed by others
Introduction
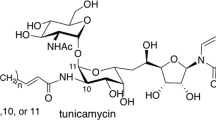
Olivomycin A (1, Scheme 1) is a member of the aureolic acid family of antibiotics. This group of highly potent agents tentatively applicable in the clinic also includes mithramycin and chromomycins (Figure 1).1
The aureolic acid derivatives contain a tricyclic chromophore (aglycon) glycosylated at positions 6 and 2 with di- and trisaccharides, respectively. The structures of olivomycins and chromomycins are very similar; the main difference is the presence of the 7-methyl group in the chromomycin aglycon (Scheme 1, Figure 1). Total synthesis of olivomycin A (1) has been described.2 Mithramycin carries the same aglycon as chromomycin A3 but different sugar moieties (Figure 1).
The cytotoxicity of these compounds for mammalian cells has been ascribed to their remarkably potent inhibitory effects on DNA-dependent enzymatic processes including replication, transcription and control of duplex topology due to the formation of non-intercalative, Mg2+-containing complexes with GC-rich regions in the DNA minor groove.3, 4, 5, 6, 7, 8 In these complexes, the compound acts as a bidentate ligand, coordinating the metal through a phenol (C9-OH) group and the neighboring ketone oxygen (C1-O).5
The carbohydrate moieties are essential for biological properties of these antibiotics.1 It has been demonstrated that the sequential hydrolysis of carbohydrate chains in chromomycin A3 was paralleled by a concomitant decrease of cytotoxicity.9 Moreover, it has been demonstrated that the absence of the D-mycarose residue (E-sugar) of the trisaccharide branch of mithramycin decreased antiproliferative activity of the mithramycin analog by one order of magnitude.10 A combinatorial biosynthesis approach used to generate derivatives of the antitumor drug mithramycin with glycosylation profiles different from that of the parental compound revealed that the presence of all five sugars is not absolutely essential for the antitumor activity of mithramycin.10
Mithramycin lacks the O-acyl substituents in polysaccharide branches, and it has a lower DNA sequence preference than chromomycin A3.11 The acetyl groups in the sugars of chromomycins are critical for the activity. Three tailoring modification steps occur during chromomycin A3 biosynthesis, affecting the sugar moieties: two O-acetylations and one O-methylation.12 Acetylation of the E-sugar moiety is the ultimate step in chromomycin A3 biosynthesis, being important for self-protection of the producing organism, as this step yields a more toxic antibiotic. The conversion of a low-active molecule into a potent one is catalyzed by the enzyme predicted to be located in the bacterial cell wall.12 Moreover, it has been recently reported that deacetylchromomycin A3 obtained from the marine-derived Streptomyces species SNB-005 strain demonstrated a decreased cytotoxicity compared with chromomycin A3.13
Whereas chromomycins and mithramycin have been intensively studied, olivomycin A (1) was investigated mainly in the last century, and structure–activity relationships for 1 are much less established. It has been shown that the structure of the sugar moieties in the saccharides is a major factor of 1–DNA complex formation, as the latter parameter has been directly attributed to the cytotoxicity of 1.14
In the present study, we elucidated the role of the acyl substituents in sugar moieties in anticancer properties of 1, namely, the drug–DNA complex formation, the interference with the catalytic function of the DNA-dependent enzyme topoisomerase I (topo I) and the cytotoxicity for cultured human tumor cells. To this aim we considered three natural aureolic acid antibiotics: olivomycin A (1), olivomycin C (2) and olivomycin B (3),15 and elaborated the methods of selective hydrolysis of E4-isobutyryl group of olivomycins (Scheme 1). It allowed investigating structure–activity relationships for a series of olivomycins 1–5, which have different combinations of the acyl resides in sugar moieties.
Results
The natural antibiotics 1–3 were isolated from the olivomycin complex obtained by fermentation of Streptoverticillum cinnamoneum strain and purified by column chromatography on silica gel and semipreparative HPLC. The obtained compounds 1–3 were totally identical to natural olivomycins A, C and B, respectively, by spectral characteristics.15 Two semisynthetic derivatives, de-E4-isobutyrylolivomycin A (4) and de-E4-isobutyrylolivomycin C (5), were obtained by selective alkaline hydrolysis of E4-O-isobutyryl group in 1 or 2, respectively (Scheme 1). Additional purification of 4 and 5 was performed by semipreparative HPLC. The structures of all the obtained compounds were confirmed by 1H NMR and mass spectral data obtained by MALDI MS (Table 1). The 1H NMR spectra of compounds 1–5 contain all the corresponding signals of the aglycon moiety and carbohydrate residues, and corresponding acyl substituents (if any) (Table 1).
As chemical acylation of the hydroxyl groups of carbohydrate moieties is accompanied by the simultaneous acylations of the hydroxyl groups of the aglycon and the side chain,16 it is not possible to obtain olivomycin derivatives containing various acyl groups in sugar residues starting from (partly) deacylated olivomycin. However, our approach based on chemical hydrolysis of natural olivomycins permitted us to have various combinations of the acyl residues in the carbohydrate chains, and study their role in biological properties of the antibiotic.
Given that binding to the DNA minor groove is the critical parameter of cytotoxicity of aureolic acid derivatives,1, 9 we studied the interaction of 1–5 with dsDNA. The concentrations of 1–5 were calculated using the known molar extinction coefficient at 400 nm (10000 M−1cm−1) and at 405 nm (8800 M−1cm−1).17, 18, 19 To avoid a re-absorption effect during fluorescence experiments, the OD (absorbance) of solutions at 420 nm did not exceed 0.05. Owing to minimal differences between the absorption spectra of free (unbound) antibiotics and their complexes with DNA in the region 410–430 nm, the excitation wavelength 420 nm was used, thereby avoiding an absorbance-related correction in further calculations.
The full overlap of the normalized absorption spectra of (antibiotic)2Mg2+ complexes of 1–5 (Figure 2a) is the evidence of a weak influence of acyl substituents in sugar residues on the aglycon. On the other hand, noticeable differences in normalized uncorrected fluorescence spectra of (antibiotic)2Mg2+ complexes of 1–5 (Figure 2b) suggested that the acyl substituents in di- and trisaccharide moieties of 1–5 could alter the structure of Mg2+ dimers. This hypothesis was supported by significantly different shape of CD spectra of (antibiotic)2Mg2+ complexes (Figures 3a and b).
Normalized absorbance spectra and spectra of fluorescence intensity of 1–5 in Mg2+-coordinated dimers and in complexes with dsDNA. Panels (a,b): 1 – 5 in dimers with Mg2+. (a), (antibiotic)=20 μM; (b), (antibiotic)=0.1 μM. Panels (c,d): Mg2+-coordinated dimers of 1 – 5 in complexes with DNA. (c), (antibiotic)=20 μM, (DNA)=3 mM; (d), (antibiotic)=0.1 μM, (DNA)=3 mM.
Interaction of Mg2+-coordinated dimers of 1–5 with dsDNA induced substantial changes in absorption, fluorescence and CD spectra (Figures 2c and d and Figures 3a and b). The absorption spectra of complexes of 1–3 with DNA showed an apparent long-wavelength shoulder; such an effect was weaker in the complex 4–DNA and was absent in 5–DNA complex (Figure 2c). The long-wavelength band was detectable in normalized fluorescence spectra of complexes of 1–3 with DNA (Figure 2d). For the complexes of 4 and 5 with DNA, the long-wavelength band was less pronounced. One may hypothesize that the structures of the drug–DNA complexes formed by 1–3 are similar, but differ from the structures of the complexes 4–DNA and 5–DNA. The peaks of intensity of the induced CD spectra of drug–DNA complexes were the highest for 1 and 3, but significantly lower for 2, 4 and 5 (Figure 3).
Studies by other researchers and us7, 14, 20 have shown that the formation of complexes of chromomycin A3, mithramycin and 1 with DNA induced changes in CD spectra. Importantly, a correlation between the intensity of the induced CD spectra of drug–DNA complexes and the structure of the drug has been demonstrated.14 The effects of the induced CD of the drug–DNA complexes are determined by the interaction of the ligand molecules with the DNA minor groove. Accordingly, the magnitude of induced CD spectra should increase with the enhancement of drug–DNA interaction. On the other hand, the structure of the DNA–antibiotic complex could influence the magnitude of induced CD spectra. It is likely that the structures of the drug–DNA complexes formed by 1 and 3 are the closest among the investigated compounds.
We next generated the fluorescence-intensity saturation curves and drew Scatchard plots20 to determine the concentrations of bound and free antibiotics, and their binding constants with dsDNA (Figure 4 and Table 2). The values of DNA-binding constants for 1–3 are the biggest. Two binding constants were calculated for compound 2, one of the same order of magnitude as that for 3, whereas the second value was one order of magnitude smaller. These differences between the DNA-binding constants for 2 could be explained by the existence of two DNA-binding sites, with a higher and a weaker affinity to this compound.
Together with the spectral characteristics, the quantum yields of fluorescence can provide information about the influence of DNA groove environment on the aglycons of 1–5. Table 2 shows the values of quantum yields of fluorescence of Mg2+-coordinated dimers of 1–5 in buffer solutions and in complexes with dsDNA. The values of quantum yields of fluorescence of (antibiotic)2Mg2+ dimers (Θlig) differed in the row 1>3>2>4>5, from the biggest for 1 (0.183%) to the lowest for 5 (0.007%). These differences correlated well with the molecular volumes of the Mg2+ dimers of 1–5 calculated by Hartree–Fock 3-21G method (Table 2). These results strongly suggest that the acyl substituents in sugars A and E can reduce the flexibility of (antibiotic)2Mg2+ complexes, in particular, the flexibility of oligosaccharide moieties, thereby affecting the P of the nonradiation processes and, consequently, the quantum yields of fluorescence of 1–5.
The quantum yields of fluorescence of DNA–(antibiotic)2Mg2+ complexes Θcom) were substantially bigger than the respective values of Mg2+-coordinated dimers of 1–5 in buffer (Table 2). This fact suggests that the interaction of the dimeric antibiotic with the DNA minor groove causes a significant decrease in the flexibility of the drug, thereby increasing Θcom in comparison with Θlig. It is noteworthy that the correlation of Θlig for 1–5 is the same as for Θcom, indicating that intermolecular interactions between the nucleotide bases in the groove and Mg2+-coordinated dimers of 1–5 could further limit the flexibility of the compounds.
We failed to carry out quantum chemicals calculation for the dimeric complexes of antibiotics 1–5, with Mg2+ taking into consideration solvation effects (solvent H2O) in the Spartan-08 program using the SM8 algorithm, supposedly, because of the unhandiness of the resulting structures. The attempts of quantum chemical calculation in the Gaussian-03 using classic method for the calculation for the solvated systems PCM also were ineffective. The above mentioned reasons forced us to use only the results of quantum chemical calculations for gaseous phase, although these results allowed to estimate the energy of formation of complexes that had good correlations with the experimental data.
Quantum chemical calculations suggest differential roles of the isobutyryl and acetyl groups: deacetylation of olivomycins results in much easier formation of the (antibiotic)2Mg2+ dimers in comparison with the parent antibiotic, whereas deletion of the isobutyryl group has a lower effect on the dimer formation. Thus, the removal of the isobutyryl group did not significantly change the calculated total energies of the formation of bidentate (antibiotic)2Mg2+ complexes (Figure 5, compare Etot for pairs 1 and 4, or 2 and 5), whereas the removal of the acetyl group substantially decreased total energies of the corresponding dimers (Figure 5, pairs 1 and 2, or 4 and 5).
A facilitated formation of Mg2+-coordinated dimers by 2 and 5 can be associated with an additional free hydroxyl group(s) in deacetylated molecules. This moiety might be in an advantageous position for the formation of an additional hydrogen bond, a negative factor for the interaction of these compact dimers with DNA.
Altogether, our data demonstrated that the acyl substituents in the sugar residues can seriously influence the formation of drug–DNA complexes, as well as the structure and flexibility of derivatives of 1. The higher affinity drug–DNA complexes were characterized by more pronounced changes in the absorption and fluorescence spectra, and by higher values of the induced CD. Important is the correlation between the values of drug–DNA-binding constants and Θcom. The bigger DNA-binding constants imply a decrease of intermolecular flexibility of drug molecules and an increased Θcom. It is possible that weak complexes of 4 and 5, and the second-type complexes of 2 are localized not as deeply in the DNA minor groove as the complexes of 1 and 3 whose intermolecular movements in the groove are limited.
Given that (1) the abilities of 1-5 to form complexes with dsDNA varied dramatically (see above), and (2) the antitumor properties of 1 have been mechanistically attributed to the formation of complexes with DNA;14 it is plausible to expect that 1-5 exert differential effects on the two parameters related to the antitumor drug potential, that is, an attenuation of topo I-mediated unwinding of supercoiled DNA20 and the cytotoxic potency for the human HCT116 colon carcinoma cell line.16 Data in Figure 6 showed almost complete inhibition of plasmid relaxation at 1 μM of 1 (rendered as +++ in Table 2), partial inhibition at 10 μM of 2 and complete inhibition at 10 μM of 3 (+ and ++ in Table 2, respectively). No inhibitory effect was detectable for 4 and 5 even at 10 μM (− in Table 2).
Attenuation of topo I-mediated plasmid DNA relaxation by 1–5. Lanes: DNAsc, supercoiled plasmid. T1, topo I+plasmid. Other lanes: topo I incubated with indicated concentrations of 1–5. See text and Tevyashova et al.20 for details.
Comparison of growth inhibitory activities revealed that 1 and 3 were extremely potent against the HCT116 colon carcinoma cell line (Table 2). These compounds killed tumor cells at nanomolar concentrations; 3 was somewhat less active than 1. In striking contrast, 2 was significantly less cytotoxic than 1 or 3, whereas 4 and 5 exerted virtually no effect on cell viability even at concentrations as high as 50 μM (Table 2).
Our results point at a good correlation between the values of DNA-binding constants, and inhibitory potencies of 1–5 on topo I function and tumor cell survival. The compounds with two acyl residues in carbohydrate chains, that is, 1 and olivomycin B (3) showed big values of DNA-binding constants and induced CD, as well as potent cytotoxic and topo I inhibitory activities (Table 2, Figure 6). The removal of the acyl residues in carbohydrate chains (compounds 2, 4 and 5) led to a decrease of the affinity to DNA paralleled by a lower cytotoxicity and topo I inhibition (Table 2 and Figure 6). Removal of O-acyl group from monosaccharide residues A (A-oliose) or E (E-olivomicose) attenuated the biological properties to various degrees. In compound 2 devoid of the O-acyl group in A-oliose the DNA-binding constant, topo I inhibitory potency and cytotoxicity decreased substantially but still remained detectable, whereas de-O-acylation of E-olivomicose residue (compound 4) as well as the complete de-O-acylation (compound 5) abrogated the affinity to DNA, the ability to inhibit topo I catalytic activity and the cytotoxicity (Table 2 and Figure 6). Interestingly, despite the limitations and assumptions of quantum chemical calculations implicated in the Hartree–Fock 3-21G method (for example, the role of a solvent was not taken into consideration) there was a good correlation between experimental and theoretical results regarding the role of the acetyl and isobutyryl groups.
Discussion
Our data clearly demonstrate that the acyl substituents in A and E sugars of 1 seriously influence the structure of (antibiotic)2Mg2+ dimers and the interaction of these dimers with DNA. The presence of the acyl groups in saccharide residues of 1 is absolutely required for valuable antitumor characteristics of this antibiotic, that is, topo I inhibition and tumor cell killing.
Nevertheless, both experimental data and quantum chemical calculations indicate somewhat different roles of the A4-acetyl group in A-monosaccharide and E4-isobutyryl group in E-monosaccharide in the biological properties of olivomycins. The absence of the isobutyryl substituent in E4-monosaccharide did not seriously influence the calculated total energies of the formation of (antibiotic)2Mg2+ dimers. In striking contrast, this modification significantly changed the fluorescence and induced CD spectra of the drug–DNA complexes, and totally abrogated the affinity of 4 and 5 to DNA. Accordingly, the cytotoxic potencies of 4 and 5, and their ability to inhibit topo I function decreased dramatically.
One may suggest that the O-acyl group in the E-olivomicose residue is critical for the formation of (olivomycin)2Mg2+ complexes with DNA. Hydrophobicity of the E4-acyl residue is preferable for drug–DNA complex formation: a more hydrophobic compound 1 (Table 1, log P data) that carries the E4-isobutyryl group was more potent than 3 that carries the E4-acetyl group (Table 2). This suggestion is in agreement with the demonstration that the binding of mithramycin and chromomycin A3 analogs to DNA is entropically driven, dominated by the hydrophobic transfer of these compounds from the solution into the DNA minor groove.21
The lack of the A4-acetyl group (compound 2) led to less pronounced changes in the fluorescence and induced CD spectra of the drug–DNA complexes in comparison with the absence of the E4-isobutyryl group (compounds 4 and 5; Figure 2d). Deletion of O-acetyl in the A-oliose residue (2 vs 1) resulted in a significant decrease (but not full loss) of the DNA-binding constant, topo I inhibitory potency and cytotoxicity. Based on our experiments, quantum chemical calculations and information about strong intermolecular interactions in the complex of (chromomycin A3)2Mg2+–DNA (including the hydrogen bonds between O4 of D-sugar and O1 of A-sugar, and between O4 of D-sugar and O4 of A-sugar)4 one may assume that besides drug–DNA binding, the residue in the position 4-O of the A-monosaccharide residue also mediates the interactions within the (olivomycin)2Mg2+ dimer. The quantum chemical calculations of the dynamic interactions of olivomycins 1–5 with GC-rich oligonucleotides using the program for molecular modeling ICM-Pro (MolSoft LLC, San Diego, CA, USA) are in progress and will be published elsewhere.
Conclusion
We provide evidence that the residue in the 4-O-position in the A-sugar residue is largely involved in the antibiotic–antibiotic interactions in the (olivomycin)2Mg2+ dimer, whereas the O-acyl group in the E-olivomicose residue is the most important for the formation of the (olivomycin)2Mg2+ complexes with DNA. Together with the modifications at the side chain of the aglycon in 1,16 the elucidation of the roles of individual chemical moieties of this molecule contributes to rational design of drug candidates on the basis of highly potent antibiotic 1. It suggests that the development of biochemical and fermentative methods of selective acylation of hydroxyl groups of the sugar moieties of olivomycin may lead to compounds with higher affinity to DNA, and more selective antiproliferative activity.
Experimental procedure
The olivomycin complex was produced at the pilot plant of Gause Institute of New Antibiotics. TLC was performed on precoated Merck (Whitehouse Station, NJ, USA) G60F254 plates in the system CHCl3–acetone–MeOH, 11:1:1. The spots were detected by sight and with a UV lamp at 254 nm. Analytical HPLC and measurement of log P were performed on a Shimadzu LC-20AD instrument (Kyoto, Japan) on a Kromasil-100-5-C18 (Akzo-Nobel, Sweden) column 4.6 × 250 mm at 20 °C. Detection was carried out using the Photo Diode Array detector Shimadzu SPD-M20A set at 274 nm at 20 °c. The mobile phase consisted of 0.01 N H3PO4 pH 2.6 and MeCN, using MeCN gradient from 33 to 45% during 0–5 min, and from 45 to 50% for 5–25 min at a flow rate of 1.0 ml min−1 (System A). The injection volume was 20 μl. The sample was dissolved in the mixture of 0.01 N H3PO4 pH 2.6 and MeCN (1:1) to a concentration of 0.2–0.5 mg ml−1. For measurement of log P by HPLC, the mobile phases consisted of MeCN–H2O (43:57 v/v) with isocratic elution, the flow rate was 1 ml min−1 (System B). Semipreparative HPLC was performed on a Diasorb-130-C16T column (16 × 250 mm, BioChemMack, Moscow, Russia) at a flow rate of 2.5–3 ml min−1 in H2O–MeCN mixture at 20 °C. The injection volume was 100 μl; the sample concentration was <20 mg ml−1. 1H NMR spectra were recorded on a Unity +400 spectrometer (Varian, Palo Alto, CA, USA) at 400 MHz in CDCl3, as a solvent using TMS as an external standard. Mass spectra determined by Matrix Assisted Laser Desorption/Ionization (MALDI) were recorded on Brucker BIFLEX III (Bruker Daltonics GmbH, Bremen, Germany). Data for predominant monoisotope peak are presented. All solutions were dried over sodium sulfate and evaporated at reduced pressure on a Buchi (Flawil, Switzerland) rotary evaporator at the temperature below 35 °C.
The absorption, fluorescence and CD spectra of drug dimers and drug–DNA complexes were detected using Shimadzu UV-3101 PC spectrophotometer, Shimadzu RF–5301 PC spectrofluorophotometer and SKD-2M dichrometer (Moscow, Russia), respectively. The double-stranded (ds) DNA from calf thymus was purchased from Sigma-Aldrich (St Louis, MI, USA).

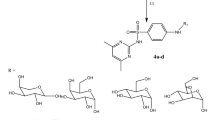
Structures and synthesis of olivomycins 1–5.
Olivomycin A (1), olivomycin C (2) and olivomycin B (3)
Compounds 1–3 were isolated by column chromatography on silica gel (CHCl3-MeOH-CH3COCH3, 20:1:1 for 1, 10:1:1 for 2 and 15:1:1 for 3) from the olivomycin complex produced by Streptoverticillum cinnamoneum strain. Olivomycin A (1) was isolated in pure state, compounds 2 and 3 were additionally purified by semipreparative HPLC in the conditions described above at a flow rate of 3 ml min−1, using 40% of MeCN in the mixture MeCN:H2O for compound 2, and 45% of MeCN for compound 3, respectively.
Compound 1: Rf 0.5, retention time (Rt) (System A) 24.0 min, Rt (System B) 24.8 min.
Compound 2: Rf 0.2, Rt (System A) 14.4 min, Rt (System B) 9.3 min.
Compound 3: Rf 0.45, Rt (System A) 14.7 min, Rt (System B) 9.8 min.
1H NMR spectra of olivomycins 1 and 3 fully corresponded to the 1H NMR spectra (CDCl3 and DMSO-d6) of the antibiotics 1 and 3, respectively, described in the literature.22, 23 13C NMR spectra of olivomycin A (1) (CDCl3 and DMSO-d6) fully corresponded to the 13C NMR spectra (CDCl3 and DMSO-d6) of the antibiotic described in the literature.22, 23 1H NMR spectrum (DMSO-d6) of 2 was virtually identical to the 1H NMR spectrum of olivomycin A22, except the absence of the signal of the acetyl group (Table 1).
De-E4-isobutyrylolivomycin A (4)
Olivomycin A (1) (40 mg, 0.03 mmol) was dissolved in MeOH (1 ml), then 0.1 N CH3ONa (1.5 ml) was added. The reaction mixture was kept at 4 °C for 3 h, diluted with H2O (15 ml) and then acidified with 0.5 N HCl to pH 6. Methanol was evaporated and water fraction was extracted with ethyl acetate (3 × 15 ml). The pooled organic fractions were evaporated to dryness, yielding crude 4 that was further purified by semipreparative HPLC in the conditions described above at a flow rate 2.5 ml min−1, using 42% of MeCN in the mixture MeCN:H2O. Rf 0.2, Rt (System A) 10.1 min, Rt (System B) 4.2 min. 1H NMR spectrum (DMSO-d6) of 4 was virtually identical to the 1H NMR spectrum of olivomycin A22, except the absence of the signals of isobutyryl group (Table 1).
De-E4-isobutyrylolivomycin C (5)
Olivomycin C (2) (110 mg, 0.1 mmol) was dissolved in MeOH (3 ml), then 0.1 N CH3ONa (0.5 ml) was added. The reaction mixture was kept at 4 °C for 5 h, diluted with H2O (35 ml) and acidified with 0.5 N HCl to pH 6. Methanol was evaporated and water fraction was extracted with ethyl acetate (3 × 25 ml). The combined organic fractions were evaporated to dryness, yielding 80 mg of crude 5 (purity 65% by HPLC). Reprecipitation from ethyl acetate-petroleum ether gave 50 mg of 5 (purity 85%). Further purification was carried out by semipreparative HPLC in the conditions described above, at a flow rate of 2.5 ml min−1 using 30% of MeCN in the mixture MeCN:H2O. Rf 0.1, Rt (System A) 6.9 min, Rt (System B) 2.7 min. 1H NMR spectrum (DMSO-d6) of 5 was virtually identical the 1H NMR spectrum of olivomycin A,22 except the absence of the signal of the acetyl and isobutyryl groups (Table 1).
Mass-spectral data and 1H NMR signals of acyl (if any), methyl and methoxy groups of 1–5 are presented in Table 1.
Determination of logP by HPLC
Methanol was used (λmax 200 nm) for measurement of the column dead time (t0=2.12 min). Each chromatographic run was repeated three times. The Rt were measured at room temperature. The capacity factors (k) were calculated using the following equation: k=(Rt−t0)/t0.24 As 1–5 belong to the same group of chemical compounds, log k=log P. The obtained data are presented in Table 1.
Spectral methods
The absorption, fluorescence and CD measurements were performed in 10 mM Tris-HCl buffer containing 50 mM MgCl2, pH 7.0 at room temperature. Before the addition of DNA compounds, 1–5 were incubated in the buffer for 1 h to form (antibiotic)2Mg2+ dimers. The quantum yields of fluorescence of 1–5 in the buffer and in drug–DNA complexes were determined using chromomycin A3 as a reference compound (quantum yield of fluorescence 5%).25 The absorption spectra of (antibiotic)2Mg2+ dimers with optical density of 0.2 (concentration of 1–5 was 20 μM) were normalized to 1 in order to reveal differences in the absorption band shape. The absorption spectra of DNA:(antibiotic)2Mg2+ complexes, and the fluorescence spectra of dimers (0.1 μM antibiotic)2Mg2+ and complexes DNA:(0.1 μM antibiotic)2Mg2+ were normalized similarly. Drug–DNA complex formation was determined upon the addition of increased amounts of DNA (1–300 μM) to the solution of (antibiotic)2Mg2+ in the buffer (final concentration of antibiotics in the solution was ∼100 nm). The absence of clear isosbestic point could be due to different DNA-binding site. The constants of drug–DNA-binding equilibrium were calculated from the fluorescence spectra using Scatchard plots, that is, the ratio r/C1 as a function of r, where r is the concentration of the bound drug per base pair, and C1 is the concentration of free (unbound) DNA.
Quantum chemical calculations
Calculations of geometry, energetic parameters and charges on the molecules of olivomycins 1–5 and their complexes with Mg2+ were carried out using Spartan-08 package by the RHF/3-21G method, with the full optimization of the geometric parameters. Structures of olivomycins 1–5, which have been preliminary calculated by the molecular mechanics MMFF method were used as initial approximations for further quantum chemical calculations.
Effects of 1–5 on cell viability and catalytic activity of topo I
The cytotoxic and topo I poisoning potencies of 1–5 were determined as described by us.16, 20 The values of IC50 were defined as the concentration of compound that reduced the number of viable cells by 50%.
References
Lombo, F., Menendez, N., Salas, J. A. & Mendez, C. The aureolic acid family of antitumor compounds: structure, mode of action, biosynthesis, and novel derivatives. Appl. Microbiol. Biotechnol. 73, 1–14 (2006).
Roush, W. R., Hartz, R. A. & Gustin, D. J. Total synthesis of olivomycin A. J. Am. Chem. Soc. 121, 1990–1991 (1999).
Tagashira, M. et al. Mithramycin represses MDR1 gene expression in vitro, modulating multidrug resistance. Biol. Pharm. Bull. 23, 926–929 (2000).
Remsing, L. L. et al. Inhibition of c-src transcription by mithramycin: structure-activity relationships of biosynthetically produced mithramycin analogues using the c-src promoter as target. Biochemistry 42, 8313–8324 (2003).
Hou, M. H., Robinson, H., Gao, Y. G. & Wang, A. H. Crystal structure of the [Mg2+-(chromomycin A3)2]-d(TTGGCCAA)2 complex reveals GGCC binding specificity of the drug dimer chelated by a metal ion. Nucleic Acids Res. 32, 2214–2222 (2004).
Albertini, V. et al. Novel GC-rich DNA-binding compound produced by a genetically engineered mutant of the mithramycin producer Streptomyces argillaceus exhibits improved transcriptional repressor activity: implications for cancer therapy. Nucleic Acids Res. 34, 1721–1734 (2006).
Cheglakov, I. B. et al. Altered transcription and replication are the mechanisms of cytotoxicity of antitumor antibiotic olivomycin A. Dokl. Biochem. Biophys. 435, 320–322 (2010).
Previdi, S. et al. Inhibition of Sp1-dependent transcription and antitumor activity of the new aureolic acid analogues mithramycin SDK and SK in human ovarian cancer xenografts. Gynecol. Oncol. 118, 182–188 (2010).
Hayasaka, T. & Inoue, Y. Chromomycin A3 studies in aqueous solutions. Spectrophotometric evidence for aggregation and interaction with herring sperm deoxyribonucleic acid. Biochemistry 8, 2342–2347 (1969).
Pérez, M. et al. Generation of new derivatives of the antitumor antibiotic mithramycin by altering the glycosylation pattern through combinatorial biosynthesis. Chem. Bio. Chem. 9, 2295–2304 (2008).
Chakrabarti, S., Bhattacharya, D. & Dasgupta, D. Structural basis of DNA recognition by anticancer antibiotics, chromomycin A3, and mithramycin: roles of minor groove width and ligand flexibility. Biopolymers 56, 85–95 (2001).
Menéndez, N. et al. Tailoring modification of deoxysugars during biosynthesis of the antitumor drug chromomycin A by Streptomyces griseus ssp griseus. Mol. Microbiol. 53, 903–915 (2004).
Hu, Y. et al. Chromomycin SA analogs from a marine-derived Streptomyces sp. Bioorg. Med. Chem. 19, 5183–5189 (2011).
Andreeva, E. V. et al. Studies of complex formation of olivomycin A and its derivatives with DNA. Dokl. Biochem. Biophys. 435, 334–338 (2010).
Berlin, Y. A., Esipov, O. A., Kiseleva, O. A. & Kolosov, M. N. Olivomycin and related antibiotics. X. Isolation and acid degradation of olivomycins A, B, C and D. Chemistry of natural compounds 280–285 Springer New York Consultants Bureau, (1967).
Tevyashova, A. N. et al. Modification of olivomycin A at the side chain of the aglycone yields the derivative with perspective antitumor characteristics. Bioorg. Med. Chem. 19, 7387–7393 (2011).
Aich, P., Sen, R. & Dasgupta, D. Role of magnesium ion in the interaction between chromomycin A3 and DNA: binding of chromomycin A3-Mg2+ complexes with DNA. Biochemistry 31, 2988–2997 (1992).
Aich, P. & Dasgupta, D. Role of magnesium ion in mithramycin-DNA interaction: binding of mithramycin-Mg2+ complexes with DNA. Biochemistry 34, 1376–1385 (1995).
Mir, M. & Dasgupta, D. Association of the anticancer antibiotic chromomycin A3 with the nucleosome: role of core histone tail domains in the binding process. Biochemistry 40, 11578–11585 (2001).
Tevyashova, A. N. et al. Modification of the antibiotic olivomycin I at the 2′-keto group of the side chain. Novel derivatives, antitumor and topoisomerase I poisoning activity. J. Antibiot. 62, 37–41 (2009).
Barcel, F. et al. DNA Binding characteristics of mithramycin and chromomycin analogues obtained by combinatorial biosynthesis. Biochemistry 49, 10543–10552 (2010).
Yoshimura, Y., Koenuma, M., Matsumoto, K., Tori, K. & Terui, Y. NMR studies of chromomycins, olivomycins and their derivatives. J. Antibiot. 41, 53–67 (1988).
Thiem, J. & Meyer, B. Studies on the structure of olivomycin A and mithramycin by 1H and 13C nuclear magnetic resonance spectroscopy. Tetrahedron 37, 551–558 (1981).
Giaginis, C. & Tsantili-Kakoulidou, A. Current state of the art in HPLC methodology for lipophilicity assessment of basic drugs. A review. J. Liquid Chromatogr. Rel. Technol. 31, 79–96 (2008).
Langlois, R. G. & Jensen, R. H. Interactions between pairs of DNA-specific fluorescent stains bound to mammalian cells. J. Histochem. Cytochem. 27, 72–79 (1979).
Acknowledgements
We are grateful to K Turchin for NMR studies, A Korolev for MS and N Malyutina for HPLC analyses (all from Gause Institute of New Antibiotics). The study was supported by the grant of the Russian Foundation for Basic Research 10-03-00210-a and the program ‘Basic Science for Medicine’ of the Presidium of the Russian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tevyashova, A., Durandin, N., Vinogradov, A. et al. Role of the acyl groups in carbohydrate chains in cytotoxic properties of olivomycin A. J Antibiot 66, 523–530 (2013). https://doi.org/10.1038/ja.2013.39
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2013.39
Keywords
This article is cited by
-
Olivomycin A – an Antitumor Antibiotic of the Aureolic Acid Group (Review)
Pharmaceutical Chemistry Journal (2016)
-
Intracellular targets of antitumor therapy: an increased complexity in the rational drug design
Russian Chemical Bulletin (2015)