Abstract
A novel actinomycete, designated strain NEAU-gq11T, was isolated from wolfberry root (Lycium chinense Mill) and characterized using a polyphasic approach. Strain NEAU-gq11T was observed to form rough-surfaced spores that borne singly on the substrate hyphae but aerial mycelia were not developed. The organism showed closest 16S rRNA gene sequence similarity to Micromonospora saelicesensis Lupac 09T (99.4%), and phylogenetically clustered with M. chokoriensis 2–19/6T (99.3%), ‘M. zeae’ NEAU-gq9T (99.3%), M. violae NEAU-zh8T (99.1%), ‘M. jinlongensis’ NEAU-GRX11T (99.1%), M. zamorensis CR38T (99.0%), M. taraxaci NEAU-P5T (98.9%) and M. lupini Lupac 14NT (98.6%). Phylogenetic analysis based on the gyrB gene sequences also indicated that the isolate clustered with the above type strains except M. violae NEAU-zh8T. The cell-wall peptidoglycan consisted of meso-diaminopimelic acid and glycine. The major menaquinones were MK-9(H8), MK-10(H6) and MK-10(H2). The phospholipid profile contained diphosphatidylglycerol, phosphatidylethanolamine and phosphatidylinositol. The major fatty acid was iso-C16:0. Furthermore, some physiological and biochemical properties and low DNA–DNA relatedness values enabled the strain to be differentiated from members of closely related species. Therefore, it is proposed that strain NEAU-gq11T represents a novel species of the genus Micromonospora, for which the name Micromonospora lycii sp. nov. is proposed. The type strain is NEAU-gq11T (=CGMCC 4.7149T=DSM 46703T).
Similar content being viewed by others
Introduction
Micromonospora is the type genus of the family Micromonosporaceae that was first described by Ørskov1 for actinomycetes that formed single, non-motile spores directly on the substrate mycelium, but no aerial mycelia were produced. Up to now, the genus Micromonospora comprises 65 species including recently described M. taraxaci NEAU-P5T,2 M. costi CS1-12T,3 ‘M. palomenae’ NEAU-CX1T,4 and ‘M. harpali’ NEAU-JC6T.4 Furthermore, the genus Micromonospora has gradually been recognized as an important source of secondary metabolites.5 Many antibiotics, such as gentamicin, netamicin, lomaiviticins A and B, tetrocarcin A, diazepinomicin and lupinacidins A and B have been isolated from this genus.5, 6, 7 Therefore, new species in the genus Micromonospora remains a focus of efficient research for the discovery of new bioactive compounds. During the investigation of exploring potential sources of endophytic actinobacteria with novel natural products in the root of wolfberry (Lycium chinense Mill), a total of 14 isolates of endophytic actinomycetes were isolated from wolfberry root sample. Out of 14 isolates, Micromonospora was the dominant genus (n=8, 57.1% of isolates), followed by Streptomyces spp. (n=6, 42.9%). In this study, we performed polyphasic taxonomy on Micromonospora strain NEAU-gq11T, and proposed that the novel isolate belongs to a new species of the genus, for which the name Micromonospora lycii sp. nov. is proposed.
Materials and Methods
Isolation and cultivation
Strain NEAU-gq11T was isolated from wolfberry root (L. chinense Mill) collected from Harbin, Heilongjiang province, north China (45°45′ N, 126°41′ E). The root sample was processed as described by Wang et al.8 and placed on a plate of humic acid-vitamin agar (HV)9 supplemented with cycloheximide (50 mg l–1) and nalidixic acid (50 mg l–1). After 2 weeks of aerobic incubation at 28 °C, colonies were transferred and purified on International Streptomyces Project (ISP) medium 310 and maintained as glycerol suspensions (20%, v/v) at –80 °C.
Morphological and physiological characteristics
Gram staining was carried out by using the standard Gram reaction method. Morphological characteristics were observed by light microscopy (Nikon ECLIPSE E200, Tokyo, Japan) and scanning electron microscopy (Hitachi S-3400N, Tokyo, Japan) after cultivation on ISP 3 medium at 28 °C for 3 weeks. Cultural characteristics were determined after being growth for 3 weeks at 28 °C on SA 1 agar,11 N-Z amine agar,12 Bennett’s agar13 and ISP media 2–7.10 ISCC-NBS color charts Standard Samples No 210614 was used to determine the color of colonies and soluble pigments. The growth temperature range (4, 10, 15, 20, 28, 35, 37, 40 and 45 °C) was determined on ISP 3 medium after culturing for 2 weeks. Tolerance of various pH (3, 4, 5, 6, 7, 8, 9, 10, 11 and 12) and NaCl (0, 1, 2, 3, 4 and 5%, w/v) were determined in GY medium15 in shake flasks (250 r.p.m.) at 28 °C for 7 days. Hydrolysis of Tweens (20, 40 and 80) were tested as described by Smibert and Krieg.16 Reduction of nitrate, degradation of gelatin, cellulose and urea, hydrolysis of starch and aesculin, coagulation of milk and production of catalase and H2S were examined as described by Gordon et al.17 Utilization of sole carbon sources was tested on ISP 9 medium.10 Utilization of amino acids as nitrogen sources was tested as described by Williams et al.18 The reference strains M. saelicesensis Lupac 09T, M. chokoriensis 2–19/6T, M. zamorensis CR38T and M. lupini Lupac 14NT were obtained from Japan Collection of Microorganisms and DSMZ (German Collection of Microorganisms and Cell Cultures), ‘M. zeae’ NEAU-gq9T,19 ‘M. jinlongensis’ NEAU-GRX11T,20 M. violae NEAU-zh8T,21 and M. taraxaci NEAU-P5T,2 were from our laboratory. These strains were cultured under the same conditions for comparative analyses.
Chemotaxonomic analysis
Biomass for chemical studies was obtained by growing the strain in GY medium in shake flasks (250 r.p.m.) at 28 °C for 7 days. Cells were collected by centrifugation, then washed with distilled water and freeze-dried. Phospholipids in cells were extracted and identified using the method of Minnikin et al.22 Whole-cell sugar composition was analyzed according to the method of Lechevalier and Lechevalier.23 For the analysis of the amino acids in peptidoglycan, the cell walls were prepared according to the procedures described by McKerrow et al.24 and analyzed by a HPLC method using Agilent TC-C18 Column (250 × 4.6 mm i.d. 5 μm) with a mobile phase consisting of acetonitrile: 0.05 mol l–1 phosphate buffer pH 7.2 (15:85, v/v) at a flow rate of 0.5 ml min–1. Peak detection was performed by using an Agilent G1321A fluorescence detector (Agilent Technologies, Santa Clara, CA, USA) with a 365 nm excitation and 455 nm longpass emission filters.24 Menaquinones were extracted from freeze-dried biomass and purified according to Collins.25 The extracts were analyzed by a HPLC−UV method using an Agilent Extend-C18 Column (150 × 4.6 mm, i.d. 5 μm), typically at 270 nm. The mobile phase was acetonitrile: 2-propanol (60:40, v/v) and the flow rate was set to 1.0 ml min–1 and the run time was 60 min. The injection volume was 20 μl, and the chromatographic column was controlled at 40 °C.20 Mycolic acids were checked by the acid methanolysis method as described previously.26 Biomass for fatty acids analysis was obtained in Trypticase Soy Broth at 28 °C for 14 days and detected by GC–MS using the method of Gao et al.20
Phylogenetic analysis
Genomic DNA of strain NEAU-gq11T was extracted as described previously by Lee et al.27 and PCR amplification of 16S rRNA gene was carried out using the method of Loqman et al.28 The PCR product was purified and cloned into the pMD19-T vector (Takara, Dalian, China) and sequenced using an Applied Biosystems DNA sequencer (model 3730XL, Shanghai, China) and software provide by the manufacturer. Almost full-length 16S rRNA gene sequence (1473 nt) was obtained and aligned with multiple sequences obtained from the GenBank/EMBL/DDBJ databases using Clustal X 1.83. The alignment was manually verified and adjusted before the construction of phylogenetic trees. Phylogenetic trees were generated with the neighbor-joining,29 maximum-parsimony30 and maximum-likelihood31 algorithms using Molecular Evolutionary Genetics Analysis software version 5.05.32 The stability of the clades in the trees was appraised by bootstrap analysis with 1000 replicates.33 A distance matrix was generated using the Kimura’s two-parameter model.34 All positions containing gaps and missing data were eliminated from the data set (complete deletion option). 16S rRNA gene sequence similarities between strains were calculated on the basis of pairwise alignment using the EzTaxon-e server.35 PCR amplification of the gyrB gene was carried out using primers GYF1 and GYR3B.36 Sequencing and phylogenetic analysis was performed as described above. The G+C content of the genomic DNA was determined by the thermal denaturation method as described by Mandel and Marmur,37 with Escherichia coli JM109 as the reference strain. DNA–DNA hybridization was carried out between strains NEAU-gq11T, M. saelicesensis Lupac 09T, M. chokoriensis 2–19/6T, ‘M. zeae’ NEAU-gq9T, ‘M. jinlongensis’ NEAU-GRX11T, M. violae NEAU-zh8T, M. zamorensis CR38T, M. taraxaci NEAU-P5T and M. lupini Lupac 14NT in a model Cary 100 Bio UV/VIS-spectrophotometer equipped with a Peltier-thermostatted 6 × 6 multicell changer and a temperature controller with in situ temperature probe (Varian) as described previously.38, 39 The DNA concentration was adjusted spectrophotometrically at 260 nm to around 1 using 0.1 × saline sodium citrate. The renaturation rates of sheared DNA were determined at 70 °C.
Results and Discussion
Morphological and physiological characteristics
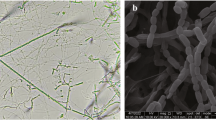
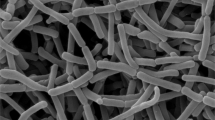
Morphological observation of a 21-day-old culture of strain NEAU-gq11T grown on ISP 3 agar revealed it had the typical characteristics of genus Micromonospora. Substrate mycelium was well-developed without fragmentation. Spores (0.5 × 0.7 μm) were borne singly on the substrate mycelia and the spore surface was rough (Figure 1). Good growth was observed on N-Z amine, Bennett’s, SA 1, ISP 2, ISP 3 and ISP 4 agar and moderate growth was observed on ISP 5, ISP 6 and ISP 7 agar. Aerial mycelia were not detected on any of the tested media. The color of colonies was grayish yellowish brown on N-Z amine agar, dark grayish yellowish brown on Bennett’s agar and SA 1 agar, brilliant orange yellow on ISP 2 agar, dark reddish orange on ISP 3 agar, deep orange yellow on ISP 4 agar and ISP 5 agar, grayish yellow on ISP 6 agar and moderate yellow on ISP 7 agar. No diffusible pigment was detected. Growth of strain NEAU-gq11T occurred in the pH range 6–12 and 0–3% NaCl (w/v), with optimum growth at pH 7.0 and 0% NaCl (w/v). The temperature range for growth was 10–37 °C, with the optimum temperature being 28 °C. Detailed physiological and biochemical properties are presented in the species description.
Chemotaxonomic characteristics
Chemotaxonomic characteristics of strain NEAU-gq11T also supported its classification as a member of the genus Micromonospora. Cells of strain NEAU-gq11T were observed to contain meso-diaminopimelic acid and glycine as diagnostic amino acids. Whole-cell hydrolysates were found to contain rhamnose, xylose and glucose. The menaquinones detected were MK-9(H8) (41.2%), MK-10(H6) (34.8%), MK-10(H2) (13.6%), MK-10(H4) (6.5%) and MK-9(H4) (3.9%). The phopholipid profile was found to consist of diphosphatidylglycerol, phosphatidylethanolamine and phosphatidylinositol (Supplementary Figure S1). The major cellular fatty acid was identified as iso-C16:0 (45.15%) (Supplementary Table S1). Mycolic acids were not detected. The G+C content of the DNA was 71.1±0.25 mol%.
Phylogenetic characteristics
The almost complete 16S rRNA gene sequence (1473 nt) of strain NEAU-gq11T was determined and deposited as KC193249 in the GenBank/EMBL/DDBJ databases. Comparative 16S rRNA gene sequence analysis showed that strain NEAU-gq11T was phylogenetically related to members of the genus Micromonospora. Phylogenetic analysis based on the 16S rRNA gene sequence indicated that strain NEAU-gq11T clustered with M. saelicesensis Lupac 09T (99.4%), ‘M. zeae’ NEAU-gq9T (99.3%), M. chokoriensis 2–19/6T (99.3%), ‘M. jinlongensis’ NEAU-GRX11T (99.1%), M. violae NEAU-zh8T (99.1%), M. zamorensis CR38T (99.0%), M. taraxaci NEAU-P5T (98.9%) and M. lupini Lupac 14NT (98.6%), which was supported by a bootstrap value of 69% in the neighbour-joining tree (Figure 2) and also recovered with the maximum-parsimony and maximum-likelihood algorithms (Supplementary Figures S2 and S3). Partial sequence of gyrB gene was obtained (1162 nt) and deposited as KJ609000 in the GenBank/EMBL/DDBJ databases. The similarities of gyrB nucleotide sequence between strains NEAU-gq11T, M. saelicesensis Lupac 09T, ‘M. zeae’ NEAU-gq9T, M. chokoriensis 2–19/6T, ‘M. jinlongensis’ NEAU-GRX11T, M. violae NEAU-zh8T, M. zamorensis CR38T, M. taraxaci NEAU-P5T and M. lupini Lupac 14NT were 97.1, 96.0, 94.9, 95.7, 92.3, 95.4 and 94.2%, respectively. Phylogenetic analysis of gyrB nucleotide sequence supported that strain NEAU-gq11T was placed in the genus Micromonospora, near to M. saelicesensis Lupac 09T, ‘M. zeae’ NEAU-gq9T, M. chokoriensis 2–19/6T, ‘M. jinlongensis’ NEAU-GRX11T, M. zamorensis CR38T, M. taraxaci NEAU-P5T and M. lupini Lupac 14NT by a 68% bootstrap value but far away from M. violae NEAU-zh8T (Supplementary Figure S4). Other studies have also reported similar results that the topology of phylogenetic trees based on gyrB gene sequences differed from those based on 16S rRNA gene sequence data.40, 41, 42
Neighbour-joining tree based on 16S rRNA gene sequence (1354 nt) showing relationship between strain NEAU-gq11T and members of the genus Micromonospora. The out-group used was Catellatospora citrea subsp. citrea DSM 44097T. Only bootstrap values above 50% (percentages of 1000 replications) are indicated. Asterisks indicate branches also recovered in the maximum-parsimony and maximum-likelihood trees. Bar, 0.005 nucleotide substitutions per site.
DNA–DNA hybridization was employed to further clarify the relatedness between strains NEAU-gq11T, M. saelicesensis Lupac 09T, ‘M. zeae’ NEAU-gq9T, M. chokoriensis 2–19/6T, ‘M. jinlongensis’ NEAU-GRX11T, M. violae NEAU-zh8T, M. zamorensis CR38T, M. taraxaci NEAU-P5T and M. lupini Lupac 14NT; the levels of DNA–DNA relatedness between them were 65.4±1.4, 64.0±2.7, 43.5±1.3, 57.5±2.4, 57.4±1.2, 62.4±1.8, 45.9±2.4 and 41.3±2.0%, respectively. These values were below the threshold value of 70% recommended by Wayne et al.43 for assigning strains to the same genomic species.
Besides the genotypic evidence above, the strain could also be distinguished from the seven related Micromonospora type strains by morphological, physiological and biochemical characteristics as summarized in Table 1, such as the clearly different colony colors on Bennett’s, SA 1, ISP 2 and ISP 3 media after being incubated at 28 °C for 3 weeks (Supplementary Figure S5) and the differences in production of urease, hydrolysis of starch, decomposition of cellulose and patterns of carbon and nitrogen utilization (Table 1). In addition, the fatty acid composition of strain NEAU-gq11T was similar to those of their closely related species such as the presence of anteiso-C15:0, iso-C16:0, C17:0, C17:1 ω7c and C18:0, but differed based on the presence or proportions of 10-methyl C16:0, C17:1 ω5c, iso-C18:0, C18:1 ω5c and C18:1 ω7c (Supplementary Table S1). Therefore, strain NEAU-gq11T represents a novel species of the genus Micromonospora, for which the name Micromonospora lycii sp. nov. is proposed. It is quite interesting that several Micromonospora species isolated from different endophytic root origins are clustering together after 16S rRNA and gyrB gene sequencing and worthy of further research.
Description of M. lycii sp. nov.
M. lycii (ly'ci.i. L. n. Lycium, name of a plant, and also a botanical generic name (Lycium); L. gen. n. lycii of Lycium, referring to the isolation of the organism from L. chinense Mill).
Gram staining positive and aerobic actinomycete that forms well-developed and branched substrate mycelium but no aerial hypha. Good growth is observed on N-Z amine, Bennett’s, SA 1, ISP 2, ISP 3 and ISP 4 agar; moderate growth is observed on ISP 5, ISP 6 and ISP 7 agar. Colonies are orange yellow color series. No diffusible pigment is detected on any of the tested media. Spores (0.5 × 0.7 μm) are single, oval with rough surface. Growth occurs at pH values between 6 and 12, the optimum being pH 7.0. Tolerates up to 3% NaCl and grows at temperatures between 10 and 37 °C, with an optimum temperature of 28 °C. Positive for liquefaction of gelatin, hydrolysis of aesculin, cellulose and Tween 40, production of catalase and urease and negative for reduction of nitrate, coagulation of milk, hydrolysis of starch, Tweens (20 and 80) and production of H2S. l-arabinose, d-fructose, d-galactose, d-glucose, inositol, lactose, d-maltose, d-mannitol, d-mannose, l-rhamnose, d-raffinose, d-ribose, d-sucrose and d-xylose are used as sole carbon sources but d-sorbitol is not. l-alanine, l-arginine, l-asparagine, l-aspartic acid, creatine, l-glutamine, l-glutamic acid, l-serine, l-threonine and l-tyrosine are used as sole nitrogen sources but glycine is not. Cell wall contains meso-diaminopimelic acid and glycine, and the characteristic whole-cell sugars are rhamnose, xylose and glucose. The phopholipids include diphosphatidylglycerol, phosphatidylethanolamine and phosphatidylinositol. The major menaquinones are MK-9(H8), MK-10(H6) and MK-10(H2). The major cellular fatty acid is iso-C16:0. Mycolic acids are absent. The G+C content of the DNA is 71.1±0.25 mol%. The type strain, NEAU-gq11T (=CGMCC 4.7149 T=DSM 46703 T), was isolated from a root of wolfberry (L. chinense Mill) collected from Harbin, Northeast China. The GenBank/EMBL/DDBJ accession numbers for the 16S rRNA gene and gyrB gene sequences of strain NEAU-gq11T are KC193249 and KJ609000, respectively.
References
Ørskov, J. Investigations into the Morphology of the Ray Fungi, Levin and Munksgaard, Enhagen, (1923).
Zhao, J. W. et al. Micromonospora taraxaci sp. nov., a novel endophytic actinomycete isolated from dandelion root (Taraxacum mongolicum Hand. -Mazz.). Antonie van Leeuwenhoek 106, 667–674 (2014).
Thawai, C. Micromonospora costi sp. nov., isolated from a leaf of Costus speciosus. Int. J. Syst. Evol. Microbiol. 65, 1456–1461 (2015).
Fang, B. Z. et al. Two new species of the genus Micromonospora Micromonospora palomenae sp. nov. and Micromonospora harpali sp. nov. isolated from the insects. Antonie van Leeuwenhoek 108, 141–150 (2015).
Bérdy, J. Bioactive microbial metabolites. J. Antibiot. 58, 1–26 (2005).
Charan, R. D. et al. Diazepinomicin, a new antimicrobial alkaloid from a marine Micromonospora sp. J. Nat. Prod. 67, 1431–1433 (2004).
Igarashi, Y. et al. Antitumor anthraquinones from an endophytic actinomycete Micromonospora lupini sp. nov. Bioorg. Med. Chem. Lett. 17, 3702–3705 (2007).
Wang, X. J. et al. Nonomuraea solani sp. nov., a novel actinomycete isolated from eggplant root (Solanum melongena L.). Int. J. Syst. Evol. Microbiol. 63, 2418–2423 (2013).
Hayakawa, M. & Nonomura, H. Humic acid-vitamin agar, a new medium for selective isolation of soil actinomycetes. J. Ferment. Technol. 65, 501–509 (1987).
Shirling, E. B. & Gottlieb, D. Methods for characterization of Streptomyces species. Int. Syst. Bacteriol. 16, 313–340 (1966).
Trujillo, M. E. et al. Micromonospora mirobrigensis sp. nov. Int. J. Syst. Evol. Microbiol. 55, 877–880 (2005).
Trujillo, M. E., Kroppenstedt, R. M., Fernández-Molinero, C., Schumann, P. & Martínez-Molina, E. Micromonospora lupini sp. nov. and Micromonospora saelicesensis sp. nov., isolated from root nodules of Lupinus angustifolius. Int. J. Syst. Evol. Microbiol. 57, 2799–2804 (2007).
Jones, K. L. Fresh isolates of actinomycetes in which the presence of sporogenous aerial mycelia is a fluctuating characteristic. J. Bacteriol. 57, 141–145 (1949).
Kelly, K. L. Inter-society color council-national bureau of standards color-name charts illustrated with centroid colors, US Government Printing Office, Washington, DC, (1964).
Jia, F. Y. et al. Wangella harbinensis gen. nov., sp. nov., a new member of the family Micromonosporaceae. Antonie van Leeuwenhoek 103, 399–408 (2013).
Smibert, R. M. & Krieg, N. R. in Phenotypic characterization (eds Gerhardt, P., Murray, R. G. E., Wood, W. A. & Krieg, N. R.). Methods for general and molecular bacteriology 607−654 (American Society for Microbiology, Washington, DC, 1994)..
Gordon, R. E., Barnett, D. A., Handerhan, J. E. & Pang, C. Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int. J. Syst. Bacteriol 24, 54–63 (1974)..
Williams, S. T. et al. Numerical classification of Streptomyces and related genera. J. Gen. Microbiol. 129, 1743–1813 (1983).
Shen, Y. et al. Micromonospora zeae sp. nov., a novel endophytic actinomycete isolated from corn root (Zea mays L.). J. Antibiot. 67, 739–743 (2014).
Gao, R. X. et al. Micromonospora jinlongensis sp. nov., isolated from muddy soil in China and emended description of the genus Micromonospora. Antonie van Leeuwenhoek 105, 307–315 (2014).
Zhang, Y. J. et al. Micromonospora violae sp. nov., isolated from a root of Viola philippica Car. Antonie van Leeuwenhoek 106, 219–225 (2014).
Minnikin, D. E. et al. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J. Microbiol. Methods 2, 233–241 (1984).
Lechevalier, M. P. & Lechevalier, H. A. in The chemotaxonomy of actinomycetes Vol. 6 (eds Dietz, A. & Thayer, D. W.). Actinomycete taxonomy special publication, 227–291 (Society of Industrial Microbiology, Arlington, 1980).
McKerrow, J. et al. A simple HPLC method for analysing diaminopimelic acid diastereomers in cell walls of Gram-positive bacteria. Lett. Appl. Microbiol. 30, 178–182 (2000).
Collins, M. D. in Chemical Methods in Bacterial Systematics (eds Goodfellow, M. & Minnikin, D. E .) Isoprenoid quinone analyses in bacterial classification and identification, 267–284 Academic Press, London, (1985).
Minnikin, D. E., Hutchinson, I. G., Caldicott, A. B. & Goodfellow, M. Thin-layer chromatography of methanolysates of mycolic acid-containing bacteria. J. Chromatogr. 188, 221–233 (1980).
Lee, Y. K., Kim, H. W., Liu, C. L. & Lee, H. K. A simple method for DNA extraction from marine bacteria that produce extracellular materials. J. Microbiol. Methods 52, 245–250 (2003).
Loqman, S., Barka, E. A., Clément, C. & Ouhdouch, Y. Antagonistic actinomycetes from Moroccan soil to control the grapevine gray mold. World. J. Microbiol. Biotechnol. 25, 81–91 (2009).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Kluge, A. G. & Farris, F. S. Quantitative phyletics and the evolution of anurans. Syst. Zool. 18, 1–32 (1969).
Felsenstein, J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Tamura, K. et al. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Kim, O. S. et al. Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 62, 716–721 (2012).
Garcia, L. C., Martínez-Molina, E. & Trujillo, M. E. Micromonospora pisi sp. nov., isolated from root nodules of Pisum sativum. Int. J. Syst. Evol. Microbiol. 60, 331–337 (2010).
Mandel, M. & Marmur, J. Use of ultraviolet absorbance temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol. 12B, 195–206 (1968).
De Ley, J., Cattoir, H. & Reynaerts, A. The quantitative measurement of DNA hybridization from renaturation rates. Eur. J. Biochem. 12, 133–142 (1970).
Huss, V. A. R., Festl, H. & Schleifer, K. H. Studies on the spectrometric determination of DNA hybridisation from renaturation rates. Syst. Appl. Microbiol. 4, 184–192 (1983).
Wang, C. et al. Micromonospora rhizosphaerae sp. nov., isolated from mangrove rhizosphere soil. Int. J. Syst. Evol. Microbiol. 61, 320–324 (2011).
Carro, L., Pukall, R., Spröer, C., Kroppenstedt, R. M. & Trujillo, M. E. Micromonospora cremea sp. nov. and Micromonospora zamorensis sp. nov., isolated from the rhizosphere of Pisum sativum. Int. J. Syst. Evol. Microbiol. 62, 2971–2977 (2012).
Li, C. et al. Micromonospora maoerensis sp. nov., isolated from a Chinese pine forest soil. Antonie van Leeuwenhoek 105, 451–459 (2014).
Wayne, L. G. et al. International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int. J. Syst. Bacteriol. 37, 463–464 (1987).
Ara, I. & Kudo, T. Two new species of the genus Micromonospora Micromonospora chokoriensis sp. nov. and Micromonospora coxensis sp. nov., isolated from sandy soil. J. Gen. Appl. Microbiol. 53, 29–37 (2007).
Acknowledgements
This work was supported in part by grants from the National Outstanding Youth Foundation (No. 31225024), the National Natural Science Foundation of China (No. 31471832, 31171913 and 31372006), the National Key Technology R&D Program (No. 2012BAD19B06), the Program for New Century Excellent Talents in University (NCET-11-0953), the Outstanding Youth Foundation of Heilongjiang Province (JC201201) and Chang Jiang Scholar Candidates Program for Provincial Universities in Heilongjiang (CSCP).
Quotation marks indicate these strains are invalidly described species.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhao, J., Guo, L., Liu, C. et al. Micromonospora lycii sp. nov., a novel endophytic actinomycete isolated from wolfberry root (Lycium chinense Mill). J Antibiot 69, 153–158 (2016). https://doi.org/10.1038/ja.2015.109
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2015.109
This article is cited by
-
Infection with Micromonospora strain SB3 promotes in vitro growth of Lolium multiflorum plantlets
Plant Cell, Tissue and Organ Culture (PCTOC) (2018)