Abstract
The aggregation of patients with Huntington's disease (HD) around Lake Maracaibo, Zulia State, Venezuela, is widely recognized, but the epidemiology of HD in the whole country is relatively unstudied. We have examined 279 individuals from 60 unrelated affected families residing in various areas of Venezuela for the presence of CAG repeats and other features associated with HD. The number of expanded repeats in 139 carriers varied from 35 to 112. Based on our examination of 71 symptomatic individuals, we developed a log-transformed regression equation, y = −0.0238x + 2.6616, to enable the prediction of age of onset in asymptomatic carriers. Intragenic haplotypes were constructed with two VNTRs (variable number of tandem repeats) and two SNPs (single nucleotide polymorphisms) in the promoter region as well as CCG repeat and Δ2642 polymorphisms to assess kinship between families. In 43 of 45 tested families, the haplotype on the mutated chromosome was 1;G;C;7;(A). The other haplotypes observed, 1;G;C;7;(B) and 4;G;C;7;(A), were of Peruvian and French origins, respectively. The geographic source of the first affected ancestor was assessed in 54 families from 15 different states. Residents of the states of Miranda, Lara and Táchira, excluding those of Zulia, had a mutated allele prevalence five- to ninefold higher than that of other areas. A low (approx. 1/200,000) prevalence, a wide-spread distribution with aggregation in some states and a likely remote European Caucasoid origin are defining epidemiologic features of HD in Venezuela.
Similar content being viewed by others
Introduction
Huntington's disease (HD) is a dominantly inherited neurodegenerative disorder caused by a mutation of the HD locus mapped at 1983 at 4p16.3 (Gusella et al. 1983) and later cloned (HDCG 1993). The unmutated gene consists of 6–34 CAG trinucleotides in exon 1; a mutation with extended CAG trinucleotide repeats beyond this normal range has been identified as the cause of HD. This mutated gene encodes a longer polyglutamine tract with deleterious effects on certain neural cells. The clinical features include motor disturbance, progressive cognitive loss and psychiatric manifestations, eventually leading to death. The age of onset of symptoms is quite variable, and the disease shows a tendency to be transmitted predominantly through the paternal line (Snell et al. 1993). Extended mutations of more than 39 CAG trinucleotides show complete penetrance, but the symptoms still appear mostly late in life.
An assessment of the size of the expanded alleles may enable an accurate diagnosis of the carrier condition and the prediction of the approximate age of onset in asymptomatic individuals, since an inverse correlation is present between the number of CAG repeats and the age of onset. One approach to ascertaining the genetic origin of a mutation(s) involved in any inherited disease is to detect concurrent intragenic markers in a population for that disease allele and the geographic distribution of the population.
Huntington's disease shows a striking geographic prevalence and aggregation among residents of the western coast of Lake Maracaibom, Zulia State, Venezuela (Negrette 1962; Avila-Girón 1973; Wexler et al. 2004). This population has the dubious honor of having the highest known frequency of carriers worldwide. However, all other epidemiologic aspects of HD are unknown among other Venezuelan populations. Here, we describe the results of a molecular analysis of the mutated HD gene among 60 families living in Venezuela of which only four (6.7%) have ancestors that came from within the endemic area in Zulia state.
Materials and methods
DNA samples
Between 1988 and 2006, 279 individuals from 60 unrelated families affected with HD were studied for confirmation of the diagnosis and genetic counseling. All of the individuals had given spontaneously consent for the tests or had been referred by neurologists to the Laboratory of Human Genetics at Instituto Venezolano de Investigaciones Científicas (IVIC). The exact place of birth of the grandparents and great-grandparents, a family history and age of onset of symptoms (psychiatric, involuntary movements, ataxia) were recorded in all index cases and in carriers. Once voluntary informed consent had been obtained from each family, in accordance with the Ethics Committee of the institute, we collected 5 mL of peripheral blood in EDTA as an anticoagulant. DNA was extracted by a saline method (Lahiri and Nurnberger 1991) and kept at room temperature (23° ± 1) in 70% ethanol for as long as 2 years in families diagnosed with HD prior to 1995, or amplified within the following 4 weeks after collection.
DNA analyses
DNA amplification of CAG repeats was achieved with primers F-5′-ATG AAG GCC TTC GAG TCC CTC AAG TCC TTC (HDCG 1993) and R-5′-GGC GGC GGT GGC TGT TG, which were designed to exclude the CCG tract nearby the CAG repeats. A PCR analysis was performed in a volume of 15 μL containing 10 pmol of each primer, 3.2 mM MgCl2and 200 μM dNTPs at 65°C (annealing temperature) with a hot start protocol (80°C for 3 min before thermal cycling). The products were resolved on 8% polyacrylamide gels (14 × 14 cm, thickness 0.8 mm; acrylamide:bisacrylamide, 29:1) in TAE buffer. The gels were stained with silver nitrate (Brandt et al. 1992).
Polymorphisms used to define haplotypes
The polymorphic markers used to ascertain ancestral origins (Arias 1994) are two variable number of tandom repeats (VNTRs) and two single nucleotide polymorphisms (SNPs) located in the promoter region, the CCG repeats in exon 1 and the GAG trinucleotide insertion/deletion in exon 58 (Δ2642), as depicted in Fig. 1. When the affected member of a family was a heterozygote for a number of the polymorphisms, a segregation analysis in the family was carried out to assess the phase. The promoter region (−303 to −1 bp) was PCR amplified with the primers Pf 5′-CAG CGG CTT GCT GTG TGA GG and Pr 5′-AGC TTT TCC AGG GTC GCC AT, which were modified from Coles et al. (1997). The PCR analysis was performed as reported, with the addition of 10% dimethylsulfoxide (DMSO) and annealing at 58°C. The products were resolved on 8% polyacrylamide gels (acrylamide:bisacrylamide, 29:1). The amplified fragments were sequenced for the SNPs at −148 G > A (GenBank: Y07982) and −103 C > T (GenBank: Y07983) by the Macrogen Sequencing Service, Seoul, Korea.
For the two VNTR polymorphisms, a combined system was used for assignment. We observed only four haplotypes associated with the first VNTR, all consisting of a 6-bp tract (one or two copies) and the second VNTR, consisting of a 20-bp stretch (one to three copies), as shown in Fig. 1. Thus, the combined allele 1 was defined as having one and two copies of the respective VNTRs; allele 2, as having two and two copies; allele 3, as having one copy of each VNTR; allele 4, as having one and three copies.
Primers used for the CCG polymorphism were HD419X and HD482X (Andrew et al. 1994). The PCR analysis was performed as reported above, with the addition of 3.5% formamide, using a touchdown protocol for thermal cycling that begins at 68°C and decreases at a rate of 0.5°C/cycle until 65°C. Products were resolved in 10% polyacrylamide gels (19:1). Primers used for the Δ2642 polymorphism were d2642F 5′-CAG TGC CCC GTT TCT GTG and d2642R 5′-TGT TAA AAA TAA AAA GGC CATC, which were designed with the Primer3 program available at http://frodo.wi.mit.edu/. The PCR analysis was performed as above with 5 pmol of each primer at 55°C. The products were resolved on 8% polyacrylamide gels (acrylamide:bisacrylamide, 29:1), and the structure and size of the allele were confirmed by sequencing The assigned alleles correspond to the number of CCG repeats; 'A' indicates deletion and 'B' indicates insertion for the Δ2642 polymorphism (Almqvist et al. 1995; Rubinsztein et al. 1995).
Statistic analyses in CAG repeat expansion
An electrophoreogram in Fig. 2 shows the normal and expanded alleles in an HD-affected family. The number of CAG repeats was calculated by dividing the detected size by three following subtraction of 50 bp that corresponds to the sequence outside the tract of CAG repeats. Thus, a maximum estimation error would be less than two CAG repeats, even in the case of increased retardation during electrophoresis. The statistical analysis was carried out using the Excel software program (Microsoft, Seattle, WA). The χ 2 test was done based on Roff and Bentzen’s methodology (1989). The regression equation graphic was obtained using a SPSS software program (SPSS, Chicago, IL).
Polyacrylamide electrophoresis of normal and expanded alleles in a single HD family. (CAG repeats): 1 Index case (22/41), 2 her 91-year-old father (17/36), 3 her mother (22/24), 4 her daughter (16/22), 5 86-bp amplicon (CFTR exon 10) according to Friedman et al. (1990), 6 DNA bp ladder (pBR322MspI digest)
Results
Allele size and its transmission to offspring
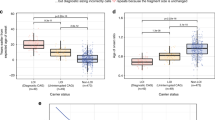
Of the 279 individuals tested, 139 had an expanded allele at HD, of whom 51 (36.7%) were still asymptomatic (ages 7–91) during the study period. The size of the allele in 40 independent control individuals (normal husbands and wives of affected cases) varied between 11 and 31 CAG repeats (Fig. 3), with 17 (21.3%), 18 (16.3%) and 19 (15.0%) CAG repeats appearing the most frequently; heterozygosity was h = 0.877. The number of CAG repeats of all these control subjects therefore fell within the normal size range – i.e. there were no major expansions or contractions. As such, the numbers of CAG repeats found in the control subjects are similar to those reported for other normal populations (Kremer et al. 1994; Masuda et al. 1995; Alonso et al. 1997; Raskin et al. 2000). The expanded alleles varied between 35 and 112 repeats, with 41 (14.4%), 42 (10.1%) and 43 (10.1%) CAG repeats appearing the most frequently (Fig. 3), which is also similar to values reported previously for other populations (Kremer et al. 1994; Masuda et al. 1995; Alonso et al. 1997; Brinkman et al. 1997; Wexler et al. 2004).
We observed eight shorter expanded repeats (35–39 repeats) with incomplete penetrance. Interestingly, some of these shorter expanded repeats showed instability in terms of being passed onto the successive generation. Three cases of this tendency that we observed are: (1) a man with 36 repeats had an affected daughter with 41 repeats (Fig. 2); (2) an asymptomatic 76-year-old father with 39 repeats had an offspring with 46 CAG repeats; (3) a mother had two sons carrying 40 and 43 repeats respectively. These findings contrast those recently reported in 80 Zulian carriers with very stable incomplete penetrant alleles (Wexler et al. 2004) that did not show expansions beyond 40 CAG repeats in their offspring.
The transmission of the mutated allele from parents to offspring was assessed in 55 cases – 30 maternal and 25 paternal transmissions. The repeat numbers changed during the transmission from parents to offspring, as shown in Fig. 4. In paternal transmissions, there were 17 instances (68.0%) of a gain of a further 2–66 repeats; in maternal transmissions, there were 12 instances (40.0%) of expansions of between one and four repeats. Any expansion of more than four repeats was always associated with paternal origin since meiotic instability is greater in fathers (Ranen et al. 1995). In our series, a single father with 46 CAGs had various offspring having between 52 to 112 repeats and sizes in between. As the extent of further expansion was not directly correlated to the expanded repeat size, the haplotype around the mutated HD locus, or even the haplotype on another chromosome, may influence the instability. Further systematic study will be required to clarify the mechanism for transmission instability.
Age of onset
The age of onset of symptoms was ascertained in 75 carriers. The earliest age of onset was, based on family accounts, 3 months – in a young female who carried an expanded allele of 112 repeats and who died at 23 years of age after being severely affected since puberty. In 58.7% of the patients, age of onset occurred between the fourth and fifth decades of life (ages 30–50); 21.3% of the patients had symptoms as early as the third decade, and one 91-year-old father with 36 repeats has not shown symptoms and most probably will not display them during his lifetime, as reported previously (Brinkman et al. 1997; Penney et al. 1997).
A log-transformed regression equation, y = −0.0238x + 2.6616 (Fig. 5), was developed on the basis of the symptomatic cases, i.e. within the range 39–54 CAG repeats (71/75 carriers). The variance in the range of age of onset seems to be lower in our patient cohort for those having more than 46 expansions and those having fewer than 41 expansions (Fig. 5). The obvious variability in the age of onset occurring in patients with 39–46 CAG expansions may reflect the joint action of genetic and environmental factors influencing the HD gene phenotype, as suggested by Wexler et al. (2004). The number of CAG repeats, individual differences in the male meiotic instability and additional, as yet unclear genetic factors seem to influence the age of onset (Rubinsztein et al. 1997; Andresen et al. 2007).
Geographic distribution of the disease
The most frequent geographic origins for the first known affected ancestor were the states of Táchira with seven families (11.7%), Miranda and Lara each with six families (10.0%), followed by Guárico, Zulia and Distrito Capital, each with four families (6.7%) (Fig. 6); in six families the geographic origin was unknown. This distribution conforms to three main geographic foci (circles 55 km in diameter = 2376 km2) (Arias 1994), with centers in Seboruco (Táchira), Curarigua (Lara) and Charallave (Miranda), respectively. The four families from Caracas (Distrito Capital) have an uncertain remote heterogeneous origin and may not constitute an actual focus according to the original definition (Arias 1994). Huntington's disease is present in 15 of 22 states, but it is primarily focal in the states of Táchira, Miranda and Lara. An additional conjoint focus may be present in the southern part of Guárico State and the northern part of Apure State, since there are two families in the San Fernando (Apure) focus.
Political map of Venezuela showing the number of HD families in the state of birth of the first known affected ancestor. The states of Táchira, Lara and Miranda had the highest number of HD families. Of 60 families, 11 had grandparents born in other countries (Spain, three; Portugal, two; France, two; Colombia, two; Peru, two). Six families had an unknown origin
Ancestors and origin
Two VNTRs in the promoter region – the CCG repeat in exon 1 and the Δ2642 polymorphisms – were ascertained in 90 members of the HD families (75 affected and 15 normal individuals) plus 25 control persons whose ancestors came mainly from the states of Táchira, Lara and Zulia. Our analyses of the partial haplotypes (VNTR;CCG;Δ2642) in both groups (HD and controls) revealed that the most frequent combination of allele 1 was the VNTRs and seven repeats of the CCG stretch (Table 1). The deletion at position 2,642 was found only in one control chromosome of the 50 tested. One affected individual (from Piura, in northern Peru) of the two Peruvian families was homozygous for this marker and thus may have a different genetic origin than those from the known southern foci in Peru (Cuba 1986; Cuba and Torres 1989; Torres et al. 2006). There is a significant (p < 0.001) difference in the frequency of haplotype 1;7;(A) between the control and HD groups. The incidence of the partial haplotype 1;10;(A) was almost sevenfold more frequent in the control group than in the homologous non-mutated chromosome of the HD mutation carriers (Table 1).
Using the SNPs at −148 G > A and −103 C > T, which were ascertained through sequencing an affected member in each Venezuelan family, two Colombian families, two Peruvian families and affected members from European countries, we constructed haplotypes for each family in each state and compared these with those with a Caucasoid origin (Spain, Portugal and France). With the exception of two families (French and Peruvian origins), those tested had the same 1;G;C;7;(A) haplotype in phase with the Huntington mutation (Table 2). The normal chromosomes in HD families had several different haplotypes, although the 1;G;C;7;(A) was also the most frequent (62.2%) (Table 2).
Discussion
Huntington disease is a genetic disorder that occurs worldwide, but the prevalence varies significantly among ethnic groups. The incidence is highest among European Caucasoids [1/11,905 in Canada (Manitoba), 1/13,333 in England (Bedfordshire), 1/18,587 in Spain (Valencia)] and lower among other ethnic groups, such as the Finnish (1/250,000), Chinese (1/270,270, Hong Kong), Japanese (1/138,889, San-in) and Africa Negroids (1/270,270, Cape, South Africa) (Frequency of Inherited Disorders Database, FIDD 2007).
To date, the published prevalence figures from Venezuela (1/143 and 1/23,000) (Negrette 1962; Avila-Girón 1973) deal exclusively with subsets of a singular community inhabiting the western coast of Lake Maracaibo in the state of Zulia ( Gusella et al. 1984; Young et al. 1986; Wexler et al. 1987; Penney et al. 1990; Bonilla 1991). The carriers were postulated originally to be descendants from a single male ancestor (Negrette 1962), although a female origin – neither documented nor empirically supported – has recently been proposed for the mutation (Okun and Thommi 2004; Wexler et al. 2004). However, the extreme high frequency and widespread distribution of the mutation in the state of Zulia within a period of only 100–120 years (Negrette 1962) is an unlikely outcome for any single female founder. Published data on the incidence of HD in areas of Venezuela outside of Zulia are limited to a congress abstract (Arias and Paradisi 1996).
The Genetic Counseling Clinic at IVIC has been – and still is – the single national reference center for the molecular diagnosis of HD in Venezuela. Since HD is a disease that is rarely found outside of Zulia State, it can be hypothesized, without much bias, that it should be possible to assess the geographic distribution and ethnic origin(s) of the mutation(s). However, the prevalence of the HD in the “general population” of different Venezuelan regions cannot yet be estimated accurately due to a definitive incomplete detection, although it does seem to be low, with a frequency of approximately 1/200,000. This incidence is estimated from data gathered during the last two decades, assuming an under-detection rate of 50% since only single families have been found in 30.4% of all the 22 nation states. Intrafocal prevalence in the three main foci described here (Táchira, Lara and Miranda) lies between 1/18,315 and 1/31,803, which is 9.1- to 5.2-fold higher than that for the “general” Venezuelan population.
The complete haplotype that we constructed using six intragenic polymorphisms, five of which lie within a stretch of less than 500 bases, was identical in almost all of the chromosomes carrying the mutation. This haplotype was observed in the families inside the states (within a focus) and in all others with which it was compared, including the affected Spanish, Portuguese, Colombian and one Peruvian family, suggesting a common and very old Caucasoid origin. According to published data, the 7/10 CCG polymorphism enables affected Mongoloid (Japanese, Chinese) and Negroid populations to be differentiated from European Caucasoid (including Finnish) individuals (Squitieri et al. 1994); the frequency of ten CCGs is lower than 6% in the latter group compared to 61% among the former two ethnic groups (Squitieri et al. 1994). All of our HD chromosomes tested had seven CCG repeats, which again supports a plausible European Caucasoid common source for the mutation in Venezuela. In contrast, the study of the CCG polymorphism in other populations, such as the Colombian and the Spanish ones (Valencia), clearly revealed at least two different origins for their mutations (Alfonso et al. 1996; García-Planells et al. 2005).
The Δ2642 deletion has been reported to be in allelic disequilibrium with some Huntington chromosomes, showing marked ethnic differences in its frequency (Almqvist et al. 1995). It is absent in Negroid, Chinese or Japanese chromosomes but has a frequency between 0.103 and 0.32 in European Caucasoid HD chromosomes (Squitieri et al. 1994; Almqvist et al. 1995). No Δ2642 deletion was found in 47 out of 48 independent HD families tested. The homologous non-mutated chromosomes of the Venezuelan HD patients also showed a lower haplotype diversity than the chromosomes of the control population (four versus seven different VNTR;CCG;Δ2642 haplotypes). This lower diversity may reflect a very remote common ancestor within the foci with the same ethnic origin in the HD families for both chromosomes but a different ethnic origin for the homologous (out-of-phase) ones in the control population.
There have been no published reports on the frequencies of the haplotype with six intragenic markers that was used in this study; therefore, any comparison is impossible at this time. Of the four promoter region polymorphisms for which frequencies are available (Coles et al. 1997; Kartsaki et al. 2006), the 1;G;C haplotype appears to be the most frequent in all studied populations, including our sample, although with interethnic differences. The 4;G;C haplotype is almost absent in normal Caucasoids, but its frequency in control Japanese chromosomes is around 10% (Coles et al. 1997); in our control sample, it is 12%. While this may indicate an Amerindian (Mongoloid) origin for the control chromosomes (Kolman et al. 1996; Tokunaga et al. 2001), it rules out this possibility for the homologous chromosome in carriers from the Venezuelan populations.
The wide distribution of affected families throughout Venezuela may suggest more than one founder phenomenon for the HD subset of European Caucasoid chromosomes. Conversely, the haplotype findings do indicate a common origin for at least the families from the states of Zulia and Táchira, and it is possible that the same interpretation could be extended to other foci. There has been demographic exchange between the southern Zulia and northern Táchira populations from the eighteenth century onwards (Osorio 1996); consequently, the ancestral identity of the mutation in these two nearby regions must be considered likely for the HD gene, which has a low mutation rate within the normal size range (Rubinsztein et al. 1995). The fact that the same identical haplotype appears in all mutated chromosomes in states far away from Zulia also points toward a probable Caucasoid origin of the mutation. A recent report on the Zulian cluster of HD carriers (Andresen et al. 2007) described different alleles in the mutated chromosomes; there were two (CCG)6 homozygotes (0.65%), and some homozygotes of the 4% of carriers in their sample had the Δ2642 marker. These findings prove the heterogeneity of origin in at least 4% of a subset of the Zulian kindred. Thus, the single HD founder chromosome hypothesis of either sex (Negrette 1962; Okun and Thommi 2004; Wexler et al. 2004) seems to be untenable for all the carriers, although it might still be admissible for the remaining 96% of the whole subset.
A multiple ethnic origin for some of the mutated chromosomes in our sample cannot be entirely excluded, but it does not seem to be likely. The results of haplotype analyses suggest that the mutation has a common ethnic descent (although not necessarily a single origin) in all Venezuelan states with HD families. This origin would appear to be European Caucasoid in Venezuela, including Zulia state, and this may be the case in some population subsets of other IberoAmerican countries.
Change history
01 April 2008
In Table 2, a column heading was inadvertently omitted. Corrected table is as follows:
References
Alfonso HS, Daza J, Carbonell C, Brokate F, Caiffa H (1996) Caracterización de las secuencias polimórficas de tripletes CAG y CCG del gen de la enfermedad de Huntington en familias colombianas. Acta Méd Colomb 12:70–75
Almqvist E, Spence N, Nichol K, Andrew SE, Vesa J, Peltonen L, Anvret M, Goto J, Kanazawa I, Goldberg YP, Hayden MR (1995) Ancestral differences in the distribution of the delta 2642 glutamic acid polymorphism is associated with varying CAG repeat lengths on normal chromosomes: insights into the genetic evolution of Huntington disease. Hum Mol Genet 4:207–214
Alonso ME, Yescas P, Cisneros B, Martínez C, Silva G, Ochoa A, Montanez C (1997) Analysis of the (CAG)n repeat causing Huntington’s disease in a Mexican population. Clin Genet 51:225–230
Andresen JM, Gayán J, Cherny SS, Brocklebank D, Alkorta-Aranburu G, Addis EA, The US–Venezuelan Collaborative Research Group, Cardon LR, Housman DE, Wexler NS (2007) Replication of twelve association studies for Huntington’s disease residual age of onset in large Venezuelan kindreds. J Med Genet 44:44–50
Andrew S, Goldberg YP, Theilmann J, Zeisler J, Hayden MR (1994) A CCG repeat polymorphism adjacent to the CAG repeat in the Huntington disease gene: implications for diagnostic accuracy and predictive testing. Hum Mol Genet 3:65–67
Arias S (1994) Selective detection among the high risk populations, instead of mass screening, proposed for countries with various homogeneous origins, and a stratified distribution of abnormal genes. In: Farriaux JP, Dhondt L (eds) New horizons in neonatal screening. Excerpta Med Int Cong Ser 1041:97–100
Arias S, Paradisi I (1996) Supratipos (=haplotipos) con marcadores de IDUA (intrón 2 y exón 3) y D4S43, para identificar alelos anormales en IT15 (Corea de Huntington) y en IDUA (mucopolisacaridosis IH). Acta Cient Venez 47[Suppl 1]:60
Avila-Girón R (1973) Medical and social aspects of Huntington’s chorea in the state of Zulia, Venezuela. Adv Neurol 1:261–266
Bonilla E (1991) ¿Por qué es importante el estudio de la Enfermedad de Huntington en Venezuela? Invest Clin 32:107–108
Brandt B, Greger V, Yandell D, Passarge E, Horsthemke B (1992) A simple and nonradioactive method for detecting the Rb1.20 DNA polymorphism in the retinoblastoma gene. Am J Hum Genet 51:1450–1451
Brinkman R, Mezei M, Theilmann J, Almqvist E, Hayden M (1997) The likelihood of being affected with Huntington disease by a particular age, for a specific CAG size. Am J Hum Genet 60:1202–1210
Coles R, Leggo J, Rubinsztein DC (1997) Analysis of the 5′ upstream sequence of the Huntington’s disease (HD) gene shows six new rare alleles which are unrelated to the age at onset of HD. J Med Genet 34:371–374
Cuba JM (1986) Un foyer de Chorée de Huntington au Pérou. Rev Neurol (Paris) 142:151–153
Cuba JM, Torres L (1989) Huit arbres généalogiques de Chorée de Huntington à Cañete (Pérou). Rev Neurol (Paris) 145:482–484
FIDD (Frequency of Inherited Disorders Database) (2007) Available at: http://www.uwcm.ac.uk/uwcm/mg/fidd
Friedman KJ, Highsmith WE, Prior TW, Perry T, Silverman LM (1990) Cystic fibrosis deletion mutation detected by PCR-mediated site-directed mutagenesis. Clin Chem 36:695–696
García-Planells JG, Burguera JA, Solís P, Millán JM, Ginestar D, Palau F, Espinos C (2005) Ancient origin of the CAG expansion causing Huntington disease in a Spanish population. Hum Mutat 25:453–459
Gusella JF, Wexler NS, Conneally PM, Naylor SL, Anderson MA, Tanzi RE, Watkins PC, Otttina K, Wallace MR, Sakaguchi AY, Young AB, Shoulson I, Bonilla E, Martin JB (1983) A polymorphic marker genetically linked to Huntington’s disease. Nature 306:234–238
Gusella JF, Tanzi R, Anderson MA, Hobbs W, Gibbons K, Raschtchian R, Gilliam TC, Wallace MR, Wexler NS, Conneally PM (1984) DNA markers for nervous system diseases. Science 225:1320–1326
HDCG (1993) A novel gene containing a trinucleotide repeat that is unstable in Huntington’s disease chromosomes. The Huntington’s Disease Collaborative Research Group. Cell 72:971–983
Kartsaki E, Spanaki C, Tzagournissakis M, Petsakou A, Moschonas N, MacDonald M, Plaitakis A (2006) Late-onset typical Huntington disease families from Crete have distinct genetic origins. Int J Mol Med 17:335–346
Kolman CJ, Sambuughin N, Bermingham E (1996) Mitochondrial DNA analysis of Mongolian populations and implications for the origin of New World founders. Genetics 142:1321–1334
Kremer B, Goldberg P, Andrew S, Theilmann J, Telenius H, Zeisler J, Squitieri F, Lin B, Basset A, Almqvist E, Bird T, Hayden M (1994) A worldwide study of the Huntington’s disease mutation. N Engl J Med 330:1401–1406
Lahiri D, Nurnberger J (1991) A rapid and non-enzymatic method for the preparation of HMW DNA from blood for RFLP studies. Nucleic Acids Res 19:5444
Masuda N, Goto J, Murayama N, Watanabe M, Kondo I, Kanazawa I (1995) Analysis of triplet repeats in the huntingtin gene in Japanese families affected with Huntington’s disease. J Med Genet 32:701–705
Negrette A (1962) Corea de Huntington. Facultad de Medicina, Universidad del Zulia, Maracaibo
Okun MS, Thommi N (2004) Americo Negrette (1924 to 2003) diagnosing Huntington disease in Venezuela. Neurology 63:340–343
Osorio FE (1996) Los Andes venezolanos. Proceso social y estructura demográfica (1800–1873). Universidad de los Andes, Cons. Publ., Mérida
Penney JB, Young AB, Shoulson I, Starosta-Rubinstein S, Snodgrass SR, Sánchez-Ramos J, Ramos-Arroyo M, Gómez F, Penchaszadeh G, Alvir J (1990) Huntington’s disease in Venezuela: 7 years of follow-up on symptomatic and asymptomatic individuals. Mov Disord 5:93–99
Penney JB, Vonsattel JP, MacDonald ME., Gusella JF, Myers RH (1997) CAG repeat number governs the development rate of pathology in Huntington’s disease. Ann Neurol 41:689–692
Ranen NG, Stine OC, Abbott MH, Sherr M, Codori AM, Chao NI, Chung AS, Pleasant N, Callahan C (1995) Anticipation and instability of IT-15 (CAG)n repeats in parent–offspring pairs with Huntington disease. Am J Hum Genet 57:593–602
Raskin S, Allan N, Teive H, Cardoso F, Santoro M, Levi G, Boy R, Lerena J, Santos V, Janzen-Dück M, Bannach L, Fellander F, Franco A (2000) Huntington disease: DNA analysis in Brazilian population. Arq Neuropsiquiatr 58:977–985
Roff DA, Bentzen P (1989) The statistical analysis of mitochondrial DNA polymorphisms: χ2 and the problem of small samples. Mol Biol Evol 6:539–545
Rubinsztein DC, Leggo J, Goodburn S, Barton DE, Ferguson-Smith MA (1995) Haplotype analysis of the Δ2642 and (CAG)n polymorphisms in the Huntington´s disease (HD) gene provides an explanation for an apparent ‘founder’ HD haplotype. Hum Mol Genet 4:203–206
Rubinsztein DC, Leggo J, Chiano M, Dodge A, Norbury G, Rosser E, Craufurd D (1997) Genotypes at the GluR6 kainate receptor locus are associated with variation in the age of onset of Huntington disease. Proc Natl Acad Sci USA 94:3872–3876
Snell R, MacMillan J, Cheadle J, Fenton I, Lazarou LP, Davies P, MacDonald ME, Gusella JF, Harper PS, Shaw DJ (1993) Relationship between trinucleotide repeat expansion and phenotypic variation in Huntington’s disease. Nat Genet 4:393–397
Squitieri F, Andrew SE, Goldberg YP, Kremer B, Spence N, Zeisler J, Nichol K, Theilmann J, Greeberg J, Goto J, Kanazawa I, Vesa J, Peltonen L, Almqvist E, Anvret M, Telenius H, Lin B, Napolitano G, Morgan K, Hayden MR (1994) DNA haplotype analysis of Huntington disease reveals clues to the origins and mechanisms of CAG expansion and reasons for geographic variations of prevalence. Hum Mol Genet 3:2103–2114
Tokunaga K, Ohashi J, Bannai M, Juji T (2001) Genetic link between Asians and native Americans: evidence from HLA genes and haplotypes. Hum Immunol 62:1001–1008
Torres L, Mori N, Mendoza M, Montoya J, Cuentas M, Domínguez J, Mendoza H, Pérez D, Mazzetti P (2006) Estudio clínico molecular de la enfermedad de Huntington en pacientes del Valle de Cañete-Perú. Diagnóstico 45:102–108
Wexler N, Young A, Tanzi R, Travers H, Starosta-Rubinstein S, Penney J, Snodgrass R, Shoulson I, Gómez F, Ramos Arroyo M, Penchaszadeh G, Moreno H, Gibbons K, Faryniarz A, Hobbs W, Anderson M, Bonilla E, Conneally P, Gusella J (1987) Homozygotes for Huntington’s disease. Nature 326:194–197
Wexler NS, Lorimer J, Porter J, Gómez F, Moskowitz C, Shackell E, Marder K, Penchaszadeh G, Roberts SA, Gayán J, Brocklebank D, Cherny SS, Cardon LR, Gray J, Dlouhy SR, Wiktorski S, Hodes ME, Conneally PM, Penney JB, Gusella J, Cha JH, Irizarry M, Rosas D, Hersch S, Hollingsworth Z, MacDonald M, Young AB, Andresen JM, Housman DE, De Young MM, Bonilla E, Stillings T, Negrette A, Snodgrass SR, Martinez-Jaurrieta MD, Ramos-Arroyo MA, Bickham J, Ramos JS, Marshall F, Shoulson I, Rey GJ, Feigin A, Arnheim N, Acevedo-Cruz A, Acosta L, Alvir J, Fischbeck K, Thompson LM, Young A, Dure L, O’Brien CJ, Paulsen J, Brickman A, Krch D, Peery S, Hogarth P, Higgins DS Jr, Landwehrmeyer B, U.S.-Venezuela Collaborative Research Project (2004) Venezuelan kindreds reveal that genetic and environmental factors modulate Huntington’s disease age of onset. Proc Natl Acad Sci USA 101:3498–3503
Young AB, Shoulson I, Penney JB, Starosta-Rubinstein S, Gómez F, Travers H, Ramos-Arroyo MA, Snodgrass SR, Bonilla E, Moreno H (1986) Huntington’s disease in Venezuela: neurological features and functional decline. Neurology 36:244–249
Acknowledgments
We are grateful to all of the families who generously gave their consent, time and patience during the long process of convening these data, and to the various physicians who sent us their patients, in particular Dr. Roberto Weiser from Caracas and the Venezuelan Association for Huntington Disease (AVEHUN). Our thanks are extended to Greta Rodríguez-Arroyo for her assistance with the SPSS program and to Miss Elena Mylonás, who efficiently processed three of the polymorphisms.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Paradisi, I., Hernández, A. & Arias, S. Huntington disease mutation in Venezuela: age of onset, haplotype analyses and geographic aggregation. J Hum Genet 53, 127–135 (2008). https://doi.org/10.1007/s10038-007-0227-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-007-0227-1
Keywords
This article is cited by
-
A deep population reference panel of tandem repeat variation
Nature Communications (2023)
-
The known burden of Huntington disease in the North of Scotland: prevalence of manifest and identified pre-symptomatic gene expansion carriers in the molecular era
Journal of Neurology (2021)
-
The targetable A1 Huntington disease haplotype has distinct Amerindian and European origins in Latin America
European Journal of Human Genetics (2017)
-
Regional research priorities in brain and nervous system disorders
Nature (2015)
-
Huntington disease-like 2 (HDL2) in Venezuela: frequency and ethnic origin
Journal of Human Genetics (2013)