Abstract
Tuberculosis (TB) is one of the most devastating chronic infectious diseases, but the role of host genetics in disease development after infection in this disease remains unidentified. Genome-wide association studies (GWASs) in Thais and Japanese were carried out and separately analyzed, attempted replication, then, combined by meta-analysis were not yielding any convincing association evidences; these results suggested that moderate to high effect-size genetic risks are not existed for TB per se. Because of failure in replication attempt of the top 50 single-nucleotide polymorphisms (SNPs) identified form meta-analysis data, we empirically split TB cases into young TB case/control data sets (GWAS-Tyoung=137/295 and GWAS-Jyoung=60/249) and old TB case/control data sets (GWAS-Told=300/295 and GWAS-Jold=123/685), re-analyzed GWAS based on age-stratified data and replicated the significant findings in two independent replication samples (young TB; Rep-Tyoung=155/249, Rep-Jyoung=41/462 and old TB; Rep-Told=212/187, Rep-Jold=71/619). GWAS and replication studies conducted in young TB identified at-risk locus in 20q12. Although the locus is located in inter-genic region, the nearest genes (HSPEP1-MAFB) from this locus are promising candidates for TB susceptibility. This locus was also associated with anti-TNF responsiveness, drug with increased susceptibility for TB. Moreover, eight SNPs in an old TB meta-analysis and six SNPs in young TB meta-analysis provided replication evidences but did not survive genome-wide significance.These findings suggest that host genetic risks for TB are affected by age at onset of TB, and this approach may accelerate the identification of the major host factors that affect TB in human populations.
Similar content being viewed by others
Introduction
Tuberculosis (TB) remains the leading cause of death in developing countries, despite the advanced insights into the mechanisms of disease development from basic science studies with animal models of host immunity and extensive studies of the virulence factors in Mycobacterium tuberculosis. Among the host immune mechanisms postulated from studies of disease development in the animal models, confirmation of their roles in human host response to M. tuberculosis infection can be obtained by identifying functional polymorphisms that affect the variability of disease development after infection in humans. Studies of polymorphisms distributed differently in the human population that affect disease development in TB would provide an unbiased estimate of the role of each immune mechanism in humans.
The conventional candidate gene approach has been applied in the search for genetic risks for leprosy, a related mycobacterial disease caused by Mycobacterium leprae. In addition, linkage analyses1, 2, 3 and a genome-wide association study (GWAS) in leprosy4 identified multiple genetic factors with moderate to large effect sizes in biologically relevant candidate genes, such as LTA, PARK2, HLA-DR-DQ, C13orf31, CCDC122 and RIPK2. For several common infectious diseases, susceptibility loci with moderate to large effect sizes (at-risk odds ratios >1.5) were successfully identified by GWAS. For example, associations have been identified between HbS and malaria5 and between CFH locus and infection by Neisseria meningitides.6 Recently, a GWAS of two African TB populations identified a gene desert in 18q11.2 as a novel candidate locus for TB, but unlike GWASs in other common infectious diseases, GWASs of TB were not successful in identifying any genetic factors with moderate to large effect sizes.7
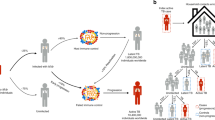
Clinical and epidemiological classifications of TB have been based on the timing of the expression of signs and symptoms after infection. Classically, three groups of TB based on this epidemiological classification were proposed: 1) primary disease after infection; 2) exogenous reinfection; and 3) endogenous reactivation.8 The primary infection and exogenous reinfection are clinical diseases that develop within 5 years after infection, whereas endogenous reactivation is a disease that develops >5 years after infection. Although there are overlapping features of clinical diseases by this classification, a proportion of primary infection TB expresses distinct clinical features, and it is hypothesized that genetic risk factors might have a major role in these TB cases.9 Unfortunately, surrogate clinical biomarkers or definite clinical definitions are not routinely available to classify patients into these categories. We cannot directly use this classification in clinical practice. However, epidemiological studies and mathematical modeling indicate that the majority of old TB cases in developed countries with a comparatively low TB incidence are mainly endogenous reactivation, whereas young TB patients in developing countries are mainly primary TB. These transmission models are typically used for prioritization of limited resources in developing countries to minimize TB incidence. The biological significance of these disease development models for TB has not been clearly defined, but the age at onset of TB may be the best available classifier to identify the subset of TB patients based on the disease development model.
Based on the epidemiological classification of TB and our findings that subset-ordered linkage analysis in TB sibpairs identified chromosome 17p and chromosome 20p as candidate loci that were enriched in young TB sibpairs10 and the results are quite different from traditional linkage analyses of TB,11, 12 we hypothesized that subdividing (or classifying) patients based on their disease-development model would enrich genetic heritability in each subset and reduce genetic heterogeneity by reducing misclassification bias. In the present study, genome-wide association analyses of TB were conducted in two East Asian populations (Japanese and Thai). The effect of age at onset of TB was examined to determine the effect on each subset of cases based on this covariate in a meta-analysis of TB GWASs, and the meta-analysis results were replicated in two independent replication data sets from Thai and Japanese populations.
Materials and methods
Japanese and Thai genome-wide association samples
The Thai samples for genome-wide genotyping included 433 TB patients and 295 healthy control samples. The patients were recruited from Chiang Rai, Lampang and Bangkok provinces. The control samples were recruited from the blood donors in Chiang Rai province. Populations in these regions were highly similar to each other and also close to the HapMap Chinese population.13 All cases were human immunodeficiency virus-seronegative when TB was diagnosed, and the diagnosis of TB was confirmed by microscopic identification or mycobacterial culture.
Japanese TB patients were recruited from the Japan BioBank projects and included 188 TB cases and 934 healthy controls (934 healthy volunteer blood donors recruited for the JSNP project14). TB was diagnosed by identification of bacteriologically confirmed M. tuberculosis. Screening for human immunodeficiency virus infection was not routinely done in Japan, but the incidence of human immunodeficiency virus in Japan was negligible at the time of the study.
Japanese and Thai replication samples
The replication Japanese samples included 112 cases of microbiologically confirmed TB and 1089 controls, and age and sex-matched controls were selected from the existing control populations into the replication analyses. The replication samples in Thailand included 369 TB cases from Chiang Rai provinces, the Chest Disease Hospital in Bangkok and the Payao Hospital in Northern Thailand. The 439 controls were recruited through the blood donors in Chiang Rai province and the healthy hypertension patients followed-up at Chiang Rai regional hospital. The control populations were screened for a history of TB and a family history of TB; all were free of diabetes mellitus. This study was approved by the Ethics Review Committee of the Ministry of Public Health in Thailand and Institutional Review Board of the Center for Genomic Medicine, RIKEN.
Genome-wide genotyping and quality control for genotyping
The genome-wide genotyping was accomplished with Illumina Hapmap 610 (Illumina, San diego, CA, USA) in the Thai GWAS samples and Illumina Hapmap 550 in the Japanese GWAS samples. Meta-analysis was carried out on the overlapping 533 252 single-nucleotide polymorphisms (SNPs) between the two platforms. Quality controls for genotypes from genome-wide genotyping were carried out using the following criteria: Hardy-Weinburg equilibrium (HWE) cut off was at 10−5 and the minimum allele frequency of polymorphism in the further analysis was 0.05. The genome-wide significance level was determined with Bonferroni’s correction taking into account the 533 252 SNPs that passed quality control (P<9.37 × 10−8). Multidimensional scaling of pairwise identity by state statistics was carried out in the GenABEL package (www.genabel.org) and indicated three outlier samples in the Thai GWAS samples; these outlier samples were later excluded. The genomic inflation factors (λ) were calculated from trend test P-values in the Thai TB GWAS (λ=1.02) and the Japanese TB GWAS (λ=0.98). These levels of stratification were acceptable. SNPs with significance levels <10−5 were checked for allelic discrimination by visual inspection of intensity cluster plots of these SNPs.
Meta-analysis of genome-wide association studies in Thai and Japanese populations
Meta-analysis was carried out with the Mantel–Haenzel test under the fixed effect model. The genotype counts of cases and controls in the Thai and Japanese TB patients were used for meta-analysis. First, 25 SNPs ordered by their significance levels (P2M-H<10−5) from the meta-analysis were subsequently replicated in the replication samples of Thai and Japanese populations by invader genotyping assay.15 Analyses of the replication samples were done, and meta-analyses of four data sets of Thai and Japanese populations were carried out, but the top at-risk 25 SNPs from the meta-analyses were not replicated in the replication samples. The meta-analysis was modified by stratifying the TB cases into two groups: a group with age at onset <45 years (young TB; GWAS-Tyoung =137/295 and GWAS-Jyoung=60/249) and another group with age at onset ⩾45 years (old TB; GWAS-Told=300/295 and GWAS-Jold=123/685), re-analyzed GWAS based on age-stratified data and replicated the significant findings in two independent replication samples (Rep-Tyoung=155/249, Rep-Jyoung=41/462, Rep-Told=212/187 and Rep-Jold=71/619). Then, the genotypes of the top 50 from meta-analyses for young TB and old TB were carried out in replication samples and presented. In the GWAS-Tyoung and GWAS-Told data sets, only age of TB cases were utilized in stratification for association analysis, but for the rest of the analysis in GWAS-Jyoung and GWAS-Jold and all replication data sets in Thais and Japanese, controls were selected from age-matched control populations.
Results
Meta-analysis of GWASs in Thais and Japanese and replication results
Genome-wide association analyses and a meta-analysis of two GWASs in Thais and Japanese were performed, and initial attempts at replication of the top 25 findings (P2M-H<10−5) carried out in second groups of the Thai and Japanese replication samples failed to identify evidence of replication. A total of 25 SNPs with P2M-H<10−5 were selected for the replication analysis in the Thai and Japanese replication sets. None of the association analyses of the top 25 SNPs passed the genome-wide significance criteria, and the meta-analysis significance levels from all four data sets (P4M-H) were higher than the original meta-analysis (P2M-H) using only the GWAS data, suggesting non-replicable results from genome-wide association of TB in Thai and Japanese populations
QQ plots of various age cut-offs in the Thai GWAS
Genome-wide association analysis using various age cut-offs other than 45 years were plotted for the age range 30–70 years by 5-year intervals. The Q-Q plots of the GWAS in younger TB tended to demonstrate highly differentiated GWAS results, with observed P-values deviating from the expected P-value. This deviation increased with younger age cut-offs despite the lower number of cases in each subset (see Supplementary Figure 1).
Meta-analysis of age-stratified GWASs in Thais and Japanese and replication results
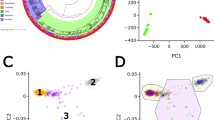
Meta-analyses of young and old TB GWASs in Thais and Japanese and their replication results are shown in Supplementary Tables 1 and 2, respectively. Selected locus that passed genome-wide significance after the meta-analysis is shown in Table 1 and Figure 1. After meta-analysis of all the data sets, there was only one locus, HSPEP1-MAFB on chromosome 20q12, that was significantly associated with young TB by genome-wide significance criteria. In addition, six additional loci provided replication evidence with lower significance levels on meta-analyses of the four data sets. For the old TB group, there were eight additional SNPs with lower P-values in the four data sets meta-analyses.
Plot of −log10 (P-value) against the physical location of the HSPEP1-MAFB locus. Blue diamond represents the PM-H of rs6071980 against its location. White, yellow, orange and red diamonds represent −log10 (P trend) from each SNP genotyped in this region from genome-wide association analysis in young Thai TB; the red diamond represents rs6071980 and white, yellow and orange colors represent the linkage disequilibrium between other SNPs and rs6071980 in Thais. Blue line indicates the recombination rates from Chinese and Japanese Hapmap population.
Discussion
Interestingly, GWASs of TB per se in Africans and in our original GWAS from two Asian populations could not identify any obvious genetic risks to TB. The sample sizes of the present TB GWAS were similar to those of other infectious diseases and it appears reasonable that one should be able to identify the common moderate- to high-risk polymorphisms that are present in other common infectious diseases, such as the association of human leukocyte antigen Class II with leprosy,4 HbS is associated with malaria5 and complement factor H variants are associated with meningococcal disease.6 Failure in replication of top signals from the meta-analysis of GWASs of TB per se in Asian populations in the present study reinforce the results of GWASs in African TB;7 no common polymorphisms with moderate to high risks to TB per se could be identified despite the ubiquitous nature of TB in global populations. GWASs in Asian populations do not have the problem of sparse linkage disequilibrium like GWASs in African populations, and they have been successfully utilized to identify common variants with large effects in Thais.16 Thus, the failure of the initial GWAS in Asian TB is not due to the problem of SNP coverage in the current microarray genotyping platform.
Taking into consideration the caveats that might affect the power of a GWAS in identifying genetic risks to common diseases17 and our experience of the increasing power of linkage analysis of TB by the subset-ordered analysis based on the age at onset of TB sibpairs,10 we hypothesized that a gene–environmental effect (G X E) interaction, such as bacille calmette guerin (BCG) vaccination, exposure to different strains of M. tuberculosis,18 or misclassification, might cause heterogeneity in TB. All of these factors contribute to different age effects on TB development. Thus, GWAS analysis of TB stratified by the age at onset might lead to identification of moderate- to high-risk polymorphisms by reducing the genetic heterogeneity.
Having access to two GWAS data sets, the TB cases were empirically split into two groups based on the age at onset of TB: the young TB group (age at onset <45 years) and the old TB group (age at onset ⩾45 years). This age cut-off is based on the bimodal distribution of age at onset of TB of patients within this study and the age distribution of TB patients in Japan19 and Thailand.20 A meta-analysis of young and old TB provided non-overlapping lists of SNPs and suggested that the top signals that contributed to young TB and old TB are distinct (see Supplementary Tables 1 and 2). Major limitation of this approach is smaller number of samples in the age-stratified analysis resulting in lower statistical power to reject null hypothesis.
A replication study for the top 50 SNPs identified from meta-analyses was performed in young TB and of old TB. A total of 100 SNPs was genotyped in both Thai and Japanese replication samples; replication in two case-control data sets identified the HSPEP1-MAFB locus as a novel susceptibility locus on chromosome 20q12 for young TB on GWASs. This locus had moderate effect sizes with an odds ratio (confidence interval) of 1.73 (1.42–2.11). rs6071980 is located 450 kb proximal to MAFB, a transcription factor determining the fate of monocyte/macrophage differentiation, and 300 kb distal to HSPEP1 (Chaperonin 10), a heat shock 10-kDa protein suggested to be an auto-antigen in autoimmune hepatitis and type I diabetes. Interestingly, rs6028945 and rs6071980, two closely located SNPs with high correlation in Caucasians, have also been suggested as genetic markers for anti-TNF responsiveness by a GWAS.21 MAFB was reported to be highly expressed in active TB compared with lower expression in latent TB in an extensive study of whole-blood gene expression signatures that differentiated active TB cases from health individuals.22 These variants might influence the development of TB through an effect on the reactive response to M. tuberculosis infection by expression of MAFB from non-lymphocytic population of white cells. Other loci with lower association evidence (P4M-H<10−5) need additional replication evidence in other populations.
This study demonstrated that consistent replication in TB gene identification is achievable by stratified analysis based on the age at onset of TB despite the smaller number of cases available for analyses. The heterogeneity by age might reflect the complexity of the epidemiology of TB in Asia: introduction of Bacille Calmette Guerin (BCG) Japan vaccination in the late 40s in Thailand23 and Japan;24 viability of individuals who carried genetic risks of TB; host adaptation to various strains of M. tuberculosis;25, 26 misclassification in the control population due to non-exposure to M. tuberculosis; temporal immunosuppression by immunosuppressive drugs;27 immune senescence; and other conventional risk factors of TB. In different age cohorts of TB, the intertwining of these factors contributed to disease development and caused genetic heterogeneity. Stratification based on the age at onset of TB as a classifier to homogenize other uncontrollable factors might be a simple and efficient method for identifying TB susceptibility genes. We suggest that consortium-level meta-analysis of age-stratified TB might provide further insight into the genetic landscapes of TB.
For genetic association studies of TB, the genetic risks for early disease after latent infection and late disease after latent infection might be totally different, and analysis that combines these similar but distinct clinical phenotypes will need large samples to overcome this heterogeneity. We chose to analyze these two clinical phenotypes separately, using age at onset as a proxy to distinguish the clinical diseases. In this study, despite the relatively small numbers of the GWAS and replication samples, we identified genetic risks that reliably replicated in two additional sample groups. Confirmatory evidences are needed in additional study from other populations. Decreasing the heterogeneity by stratification is not uncommonly used in genetic mapping of complex traits, and because it may have been overlooked, it may have contributed to the large samples required in GWASs in some diseases. For diseases with hidden heterogeneity, a case-classification system that clusters cases into more homogeneous groups might be an alternative approach to overcome genetic heterogeneity.
References
Mira, M. T., Alcais, A., Van Thuc, N., Thai, V. H., Huong, N. T., Ba, N. N. et al. Chromosome 6q25 is linked to susceptibility to leprosy in a Vietnamese population. Nat. Genet. 33, 412–415 (2003).
Mira, M. T., Alcais, A., Nguyen, V. T., Moraes, M. O., Di Flumeri, C., Vu, H. T. et al. Susceptibility to leprosy is associated with PARK2 and PACRG. Nature 427, 636–640 (2004).
Alcais, A., Alter, A., Antoni, G., Orlova, M., Nguyen, V. T., Singh, M. et al. Stepwise replication identifies a low-producing lymphotoxin-alpha allele as a major risk factor for early-onset leprosy. Nat. Genet. 39, 517–522 (2007).
Zhang, F. R., Huang, W., Chen, S. M., Sun, L. D., Liu, H., Li, Y. et al. Genome-wide association study of leprosy. N Engl J Med 361, 2609–2618 (2009).
Jallow, M., Teo, Y. Y., Small, K. S., Rockett, K. A., Deloukas, P., Clark, T. G. et al. Genome-wide and fine-resolution association analysis of malaria in West Africa. Nat. Genet. 41, 657–665 (2009).
Davila, S., Wright, V. J., Khor, C. C., Sim, K. S., Binder, A., Breunis, W. B. et al. Genome-wide association study identifies variants in the CFH region associated with host susceptibility to meningococcal disease. Nat. Genet. 42, 772–776 (2010).
Thye, T., Vannberg, F. O., Wong, S. H., Owusu-Dabo, E., Osei, I., Gyapong, J. et al. Genome-wide association analyses identifies a susceptibility locus for tuberculosis on chromosome 18q11.2. Nat. Genet. 42, 739–741 (2010).
Sutherland, I., Svandova, E. & Radhakrishna, S. The development of clinical tuberculosis following infection with tubercle bacilli. 1. A theoretical model for the development of clinical tuberculosis following infection, linking from data on the risk of tuberculous infection and the incidence of clinical tuberculosis in the Netherlands. Tubercle 63, 255–268 (1982).
Alcais, A., Fieschi, C., Abel, L. & Casanova, J. L. Tuberculosis in children and adults: two distinct genetic diseases. J Exp Med 202, 1617–1621 (2005).
Mahasirimongkol, S., Yanai, H., Nishida, N., Ridruechai, C., Matsushita, I., Ohashi, J. et al. Genome-wide SNP-based linkage analysis of tuberculosis in Thais. Genes Immun 10, 77–83 (2009).
Bellamy, R., Beyers, N., McAdam, K. P., Ruwende, C., Gie, R., Samaai, P. et al. Genetic susceptibility to tuberculosis in Africans: a genome-wide scan. Proc. Natl Acad. Sci. USA 97, 8005–8009 (2000).
Cooke, G. S., Campbell, S. J., Bennett, S., Lienhardt, C., McAdam, K. P., Sirugo, G. et al. Mapping of a novel susceptibility locus suggests a role for MC3R and CTSZ in human tuberculosis. Am. J. Respir. Crit. Care Med. 178, 203–207 (2008).
Mahasirimongkol, S., Chantratita, W., Promso, S., Pasomsab, E., Jinawath, N., Jongjaroenprasert, W. et al. Similarity of the allele frequency and linkage disequilibrium pattern of single nucleotide polymorphisms in drug-related gene loci between Thai and northern East Asian populations: implications for tagging SNP selection in Thais. J. Hum. Genet. 51, 896–904 (2006).
Hirakawa, M., Tanaka, T., Hashimoto, Y., Kuroda, M., Takagi, T., & Nakamura, Y. JSNP: a database of common gene variations in the Japanese population. Nucl. Acids Res. 30, 158–162 (2002).
Ohnishi, Y., Tanaka, T., Ozaki, K., Yamada, R., Suzuki, H. & Nakamura, Y. A high-throughput SNP typing system for genome-wide association studies. J. Hum. Genet. 46, 471–477 (2001).
Nuinoon, M., Makarasara, W., Mushiroda, T., Setianingsih, I., Wahidiyat, P. A., Sripichai, O. et al. A genome-wide association identified the common genetic variants influence disease severity in beta(0)-thalassemia/hemoglobin E. Hum. Genet. 127, 303–314 (2010).
Manolio, T. A., Collins, F. S., Cox, N. J., Goldstein, D. B., Hindorff, L. A., Hunter, D. J. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Rienthong, D., Ajawatanawong, P., Rienthong, S., Smithtikarn, S., Akarasewi, P., Chaiprasert, A. et al. Restriction fragment length polymorphism study of nationwide samples of Mycobacterium tuberculosis in Thailand, 1997–1998. Int. J. Tuberc. Lung Dis. 9, 576–581 (2005).
Research Institute of Tuberculosis J. Tuberculosis Annual Report 2008–Series 1. Summary of TB notification statistics in 2008. Kekkaku 84, 693–696 (2009).
Siriarayapon, P., Yanai, H., Glynn, J. R., Yanpaisarn, S. & Uthaivoravit, W. The evolving epidemiology of HIV infection and tuberculosis in northern Thailand. J. Acquir. Immune Defic. Syndr. 31, 80–89 (2002).
Liu, C., Batliwalla, F., Li, W., Lee, A., Roubenoff, R., Beckman, E. et al. Genome-wide association scan identifies candidate polymorphisms associated with differential response to anti-TNF treatment in rheumatoid arthritis. Mol. Med. 14, 575–581 (2008).
Berry, M. P., Graham, C. M., McNab, F. W., Xu, Z., Bloch, S. A., Oni, T. et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature 466, 973–977 (2010).
Ritz, N. & Curtis, N. Mapping the global use of different BCG vaccine strains. Tuberculosis (Edinb) 89, 248–251 (2009).
Yamamoto, S. & Yamamoto, T. Historical review of BCG vaccine in Japan. Jpn. J. Infect. Dis. 60, 331–336 (2007).
Gagneux, S., DeRiemer, K., Van, T., Kato-Maeda, M., de Jong, B. C., Narayanan, S. et al. Variable host-pathogen compatibility in Mycobacterium tuberculosis. Proc. Natl Acad. Sci. USA 103, 2869–2873 (2006).
Caws, M., Thwaites, G., Dunstan, S., Hawn, T. R., Lan, N. T., Thuong, N. T. et al. The influence of host and bacterial genotype on the development of disseminated disease with Mycobacterium tuberculosis. PLoS Pathog. 4, e1000034 (2008).
Ledingham, J., Wilkinson, C. & Deighton, C. British Thoracic Society (BTS) recommendations for assessing risk and managing tuberculosis in patients due to start anti-TNF-{alpha} treatments. Rheumatology (Oxford) 44, 1205–1206 (2005).
Acknowledgements
This work was partly supported by a Grant in-Aid for Scientific Research (B), (KAKENHI 21406006) from Japan Society for the Promotion of Science (JSPS). The clinical information and samples in Thailand, in this part was supported by JSPS KAKENHI 17256005 and later by research grant from Ministry of Health, Labor and Welfare (MHLW) H21-aids-12. We would like to thank all the subjects and the members of the Rotary Club of Osaka-Midosuji District 2660 Rotary International in Japan who donated their DNA for this work. We thank all members of BioBank Japan, Institute of Medical Science, The University of Tokyo and of RIKEN Center for Genomic Medicine for their contribution to the completion of our study. This work was conducted as a part of the BioBank Japan Project that was supported by the Ministry of Education, Culture, Sports, Science and Technology of the Japanese government. As for Thai samples, we thank all of the staffs and collaborators of the TB/HIV Research Project, Thailand, a research project between the Research Institute of Tuberculosis, the Japan Anti-tuberculosis Association and the Thai Ministry of Public Health for collecting clinical data and DNA samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Mahasirimongkol, S., Yanai, H., Mushiroda, T. et al. Genome-wide association studies of tuberculosis in Asians identify distinct at-risk locus for young tuberculosis. J Hum Genet 57, 363–367 (2012). https://doi.org/10.1038/jhg.2012.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.35
Keywords
This article is cited by
-
The immunogenetics of tuberculosis (TB) susceptibility
Immunogenetics (2023)
-
Macrophage migration inhibitory factor − 794 CATT5−8 microsatellite polymorphism and susceptibility of tuberculosis
Infection (2021)
-
Fine-mapping analysis of a chromosome 2 region linked to resistance to Mycobacterium tuberculosis infection in Uganda reveals potential regulatory variants
Genes & Immunity (2019)
-
Association between CD53 genetic polymorphisms and tuberculosis cases
Genes & Genomics (2019)
-
Human genetics of mycobacterial disease
Mammalian Genome (2018)