Abstract
Microarray-based gene expression profiling is well suited for parallel quantitative analysis of large numbers of RNAs, but its application to cancer biopsies, particularly formalin-fixed, paraffin-embedded (FFPE) archived tissues, is limited by the poor quality of the RNA recovered. This represents a serious drawback, as FFPE tumor tissue banks are available with clinical and prognostic annotations, which could be exploited for molecular profiling studies, provided that reliable analytical technologies are found. We applied and evaluated here a microarray-based cDNA-mediated annealing, selection, extension and ligation (DASL) assay for analysis of 502 mRNAs in highly degraded total RNA extracted from cultured cells or FFPE breast cancer (MT) biopsies. The study included quantitative and qualitative comparison of data obtained by analysis of the same RNAs with genome-wide oligonucleotide microarrays vs DASL arrays and, by DASL, before and after extensive in vitro RNA fragmentation. The DASL-based expression profiling assay applied to RNA extracted from MCF-7 cells, before or after 24 h stimulation with a mitogenic dose of 17β-estradiol, consistently allowed to detect hormone-induced gene expression changes following extensive RNA degradation in vitro. Comparable results where obtained with tumor RNA extracted from FFPE MT biopsies (6 to 19 years old). The method proved itself sensitive, reproducible and accurate, when compared to results obtained by microarray analysis of RNA extracted from snap-frozen tissue of the same tumor.
Similar content being viewed by others
Main
Breast cancer (MT) is a heterogeneous and complex disease, characterized by molecular and genetic diversity, which causes qualitatively and quantitatively aberrant gene expression, and, as a result, significantly divergent biological and clinical behaviors. Subtle differences in the expression of a limited number of genes among otherwise undistinguishable MTs may indeed underscore substantial differences in the prognostic outcome of the disease, in particular concerning its recurrence and responsiveness to therapy. Treatment decisions, as well as prognostic evaluation, are currently guided by efforts to determine ‘a priori,’ at the time of diagnosis or surgical removal of the primary lesion, the metastatic potential of the disease. Clinical variables that reflect this potential, including tumor size, histological grade and lymph node status, or that help predict responsiveness to chemotherapy or targeted therapy (hormone receptor status, HER-NEU oncogene amplification/overexpression, and so on) are routinely used to classify tumors into subtypes predictive of outcome. These parameters, however, are unable to predict with sufficient accuracy patients that will primarily benefit from a given therapy, and it is currently accepted that the ability to precisely determine the molecular profile of a tumor at diagnosis would provide the clinician with information relevant for an individualized medicine, including selection of the most appropriate therapy regimen.
Several studies using gene expression profiling of tumor samples with microarrays have shown that clinical heterogeneity of MT may be resolved at the molecular level and, more important, that gene expression signatures underlying specific biological properties of cancer cells provide better stratification of patients than established prognostic variables.1, 2, 3, 4, 5 However, the different molecular predictor gene profiles discovered so far were not found always concordant for classification of the risk of recurrence and death, when evaluated in a single MT cohort.6, 7 Furthermore, the two main prognostic gene signatures derived so far2, 5 do not validate in the other's data set, even when cohort differences are taken into account,8, 9 raising questions about the true clinical usefulness of the molecular signatures being currently proposed.10
The basis for such uncertainties can be only partly explained on technical and conceptual grounds. The wide differences in inclusion criteria of patients, evaluation of disease outcome and protocols of treatment among the different MT cohorts from different studies make it difficult to correctly compare the effective prognostic power of each molecular signature, while comparison of gene expression data generated with different microarray platforms and evaluated with different statistical tools is, at present, quite unreliable. The latter problem is being addressed by the MicroArray Quality Control (MAQC) project,11 which aims at improving inter- and intraplatform reproducibility of gene expression measurements, but the former one needs a substantial improvement in study design. Ein-Dor et al8 suggested that very large numbers of samples, analyzed in parallel under identical assay conditions, are required to generate a robust gene list for predicting outcome in cancer. In addition, as functional gene expression and functional data relative to MTs are already available, such as in the case of estrogen- and anti-estrogen-responsive gene sets for example,12, 13 selection of gene lists based upon known biological functions of MTs (molecular classifiers)14 may, in some specific circumstances, yield more accurate and reliable gene expression results, directly linked to functional data. Indeed, this approach allows excluding from analysis a large number of unrelated and uninformative genes, thereby decreasing background noise during statistical evaluation of the microarray data.15
The possibility to use RNA from archived formalin-fixed, paraffin-embedded (FFPE) samples would greatly help solve many of these problems. Indeed, given the wide availability of paraffin-embedded tissue blocks dating back several years and including valuable clinical annotation, such as those derived from clinical trials, would provide enough material for large and uniform retrospective studies. Unfortunately, FFPE samples provide in most cases RNA unfit for standard analysis by microarray-based methods,16 due to extensive RNA degradation by formalin treatment and during storage.17, 18
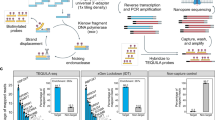
We tested here the recently developed cDNA-mediated annealing, selection, extension and ligation (DASL) methodology for gene expression profiling of highly degraded human RNA samples,19, 20 applying it to in vitro degraded RNA from human MTs and FFPE MT biopsies. In the DASL assay, total RNA is converted into cDNA in a reverse transcription reaction using biotinylated primers, followed by hybridization to query oligonucleotides (in general up to three distinct for each mRNA), primer extension and ligation, fluorescence labeling by polymerase chain reaction (PCR) and annealing (capture) into the array substrate (beads). The limited size of the RNA target sequence and the use of random priming during cDNA synthesis allow analysis of very small RNA fragments, whereas fluorescence labeling by PCR yields high specific activity probes and greatly increases sensitivity of the assay, allowing detection of low-abundance transcripts.
Comparison of results obtained for the same RNA samples on standard vs DASL arrays, and of intact vs highly degraded samples, showed that this assay provides parallel quantification of large numbers of RNAs derived from FFPE samples with excellent sensitivity, high reproducibility and accuracy.
MATERIALS AND METHODS
Cell Cultures
Human MT MCF-7 cells (ATCC, Cat No. HTB-22) were cultured in Dulbecco's modified Eagle's medium, supplemented with 10% fetal bovine serum (FBS) (both from Invitrogen SpA, San Giuliano, Milanese, Italy), 100 U/ml penicillin, 100 μg/ml streptomycin and 250 ng/ml Amfotericin-B at 37°C in a humidified atmosphere composed of 95% air and 5% CO2. Cells were provided with fresh medium every 2 days and after reaching a confluence of 40–60%, they were used for the experiment. To evaluate the effect of estrogens, cells were grown in phenol red-free Dulbecco's modified Eagle's medium containing 5% dextran-charcoal-stripped FBS for 5 days and stimulated with 10−8 M 17β-estradiol (E2) as described previously.12, 13
Tumor Tissue Samples
All bioptic tissues for this study were obtained from different patients following their informed consent and belong to the Tumor Bank collections of the Department of Obstetrics and Gynecology, University of Turin. All samples were FFPE invasive ductal carcinomas, according to standard tissue acquisition protocols. We obtained 13 FFPE MT samples, of which 3 were also available as cryopreserved samples in liquid nitrogen, and 2 FFPE bladder cancer (BLT) samples. The paraffin blocks were prepared 6 to 19 years before analysis.
RNA Extraction, Purification and Quality Assessment
RNA was extracted from MCF-7 cells before or after stimulation with E2 for 24 h, using the standard RNA Extraction with TRIzol (Invitrogen SpA) method, as described previously.12, 13
FFPE tissue samples were cut into 5 μm-thick sections on a microtome with a disposable blade, and the sections were stored in xylene at room temperature until use, before washing in fresh xylene and RNA extraction.
Up to four RNA extractions were performed either from three or eight sequential sections from the same paraffin block using the High Pure RNA Paraffin Kit (Roche Diagnostics GmbH, Mannheim, Germany). An initial deparaffinization was first performed with xylene, sections were then extracted in ethanol and homogenized by overnight incubation in Proteinase K. Solubilized nucleic acids were bound to a glass fiber filter in the presence of guanidine salts, and filter-bound nucleic acids were washed and RNA was selectively eluted. DNase I was then used to digest residual DNA, followed by an additional Proteinase K digestion step. RNA was bound to another glass fiber filter, washed and eluted for higher purification.
Total RNA from three FFPE MT tissues was isolated from cryopreserved tissue samples of the same tumors, according to standard protocols, as described earlier,21 total RNA was then precipitated at −20°C with isopropyl alcohol and RNA pellet was washed with 75% ethanol and dissolved in water.
RNA concentration was determined with a ND-1000 spectrophotometer (NanoDrop, Wilmington, DE, USA), and its quality was assessed with an Agilent 2100 Bioanalyzer using Agilent RNA 6000 Nano kit (Agilent Technologies, Santa Clara, CA, USA). To estimate its level of degradation, the RNA integrity number (RIN22) was calculated. This value takes into consideration the whole electrophoretic profile of the RNA sample, including the presence or absence of degradation products.
RNAs from MCF-7 cells and from one of the FFPE MT samples were degraded by incubation at 95°C, as follows: MCF-7 cell RNA was exposed to heat degradation for 0, 15, 30, 60 or 90 min and FFPE MT RNA was treated for 0, 5, 10, 20, 30 or 40 min.
A series of mixings of MT and BLT RNAs were also prepared, according to the following MT RNA:BLT RNA proportions: 100:0, 80:20, 60:40, 40:60, 20:80 and 0:100. RNA was stored at −80°C until use.
Quantitative Real-Time Reverse Transcription Polymerase Chain Reaction
cDNA synthesis was performed with the Qiagen QuantiTect Reverse Transcription kit (Qiagen SpA, Milano, Italy). Fluorescent quantitative real-time reverse transcription-PCR (RT-qPCR) analyses were performed with an ABI Prism 7900HT sequence detection system (Applied Biosystems, Foster City, CA, USA). PCR primers for the ribosomal protein L13a (RPL13A) transcript (RefSeq Accession Number: NM_012423.2) were designed to amplify 82 and 112 bp fragments, with two probes sets (28 and 43) from the Exiqon Human Probe Library (Exiqon S/A, Vedbaek, Denmark). The following primers were used for PCRs:
RPL-28: GAGGCCCCTACCACTTCC (forward) and CTCGCTTGGTTTTGTGG (reverse), RPL-43: GAGGCCCCTACCACTTCC (forward) and AACACCTTGAGACGGTCCAG (reverse).
Each reaction contained 4 μl of RealMasterMix (Eppendorf, Hamburg, Germany), 12 ng of cDNA template and 0.2 nM of each forward and reverse primers, in a total reaction volume of 10 μl. The PCR consisted of 45 cycles of 94°C for 90″, 94°C for 20″ and 60°C for 60″.
Microarray Analyses
For DASL arrays, the Illumina DASL Human Cancer Panel gene set (Illumina Inc., San Diego, CA, USA) were used, represented by a pool of selected probe groups that target 502 gene mRNAs collected from publicly available cancer gene lists (SNP500 Cancer Database and Cancer Genome Anathomy Project; see Table 1 in Supplementary Information), each mRNA being targeted in three locations by three separate probes. A 200 ng portion of total RNA was converted into cDNA using biotinylated random nonamers, oligo-deoxythymidine 18 primers and Illumina-supplied reagents, according to manufacturer's instructions. The resulting biotinylated cDNA was annealed to assay oligonucleotides and bound to streptadivin-conjugated paramagnetic particles to select the cDNA/oligo complexes. After oligo hybridization, mis-hybridized and non-hybridized oligos were washed away, while bound oligos were extended and ligated to generate templates to be subsequently amplified with shared PCR primers. The fluorescent-labeled complementary strand was hybridized at 45°C for 18 h to Illumina Sentrix Universal-96 Array Matrix (SAM) platform and Universal-16 BeadChips. The SAM platform is a fiber-optic assembly composed of 96 individual arrays, while the Universal-16 BeadChip platform is composed of 16 individual arrays manufactured on a microscope slide-shaped substrate. For each sample, at least three technical replicates were performed. After hybridization, the arrays were scanned by laser confocal microscopy using the Illumina BeadArrary Reader 500 system.
Illumina HumanRef-8_V1 Expression BeadChip microarrays (whole genome (WG)), including 24 613 gene-specific oligonucleotide probes, were used. RNA samples were prepared for array analysis using the Illumina TotalPrep RNA Amplification Kit (Ambion, Austin, TX, USA). To synthesize the first cDNA strand, 500 ng total RNA were reverse transcribed using T7 Oligo (dT) Primers, dNTP Mix, RNase inhibitor and an engineered Reverse Transcription ArrayScript. Samples were incubated at 42°C for 2 h. The second cDNA strand synthesis reaction employed DNA polymerase and RNase H. After incubation at 16°C for 2 h, double-stranded cDNA was purified and biotinylated cRNA was synthesized by in vitro transcription with T7 RNA polymerase and biotin-NTP, purified and its concentration and quality were assessed as described above. For each sample, 700 ng of cRNA were hybridized at 55°C for 18 h to the array, followed by washing and staining with streptavidin-conjugated cyanine 3 (GE Healthcare Italy SpA, Milano, Italy). BeadChips were dried and subsequently scanned with the Illumina BeadArray Reader 500.
Data Analysis
Data analyses were performed with BeadStudio software and R/BioConductor programming environment for statistical computing.23 A quality control (QC) report was generated for all arrays and outliers were discarded on the basis of internal BeadStudio controls. Data were normalized with the average normalization algorithm and genes were considered as detected if the detection P-value was lower than 0.01. Arrays with less than 50% detected genes were not included in further analyses.
Differential expression analysis was performed with Illumina Diffscore, a proprietary algorithm24 that uses bead standard deviation to build an error model.
Cluster analysis of samples was carried out in BeadStudio,24 using the Average Linkage method, and distance was expressed as correlation coefficient.
Heat maps were generated with the Multiexperiment Viewer 4.0 software25 after median centering signal intensity values and performing hierarchical clustering of genes with the average linkage method and Euclidian distance.
RESULTS
Test of DASL Assay Performance with In Vitro-Fragmented RNA from Human Breast Cancer Cells and FFPE Breast Cancer Samples
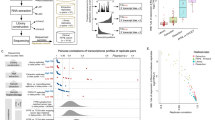
With the aim of verifying DASL protocol performance upon degraded mRNA, two samples of RNA extracted from estrogen-deprived and -stimulated (24 h) MCF-7 cells were fragmented in vitro to various extent by exposure to 95°C for different times and profiled by microarray, according to the DASL method (Figure 1). Samples containing RNA at various level of degradation were obtained in this way, as shown by the loss of ribosomal RNAs and the accumulation of RNA molecules of decreasing average size, that by 90 min reached a plateau (Figure 1a; data not shown). As an indicator of mRNA degradation, the transcript encoded by the housekeeping gene encoding the ribosomal protein L13a (RPL13A) was monitored in all samples by RT-qPCR (Figure 1a). RT-qPCR protocol was set first by determining the appropriate amount of RNA to start with (12 ng) and by using two different probe sets (data not shown). As shown by side-by-side comparison of the gel-like image of micro-capillary electropherograms (Figure 1a, left panel) with the quantity of RPL13A mRNA detectable in the same sample, represented by the (threshold cycle) Ct of the RT-qPCR (Figure 1a, right panel), this method provides a reliable way to measure precisely the degree of RNA degradation in a sample. Data displayed in Figure 1a refer to RNA extracted from E2-treated MCF-7 cells. The same analysis was carried out also on the RNAs from hormone-starved cells, yielding essentially the same results (data not shown).
Evaluation of DASL array technology with intact and in vitro-degraded RNA from MCF-7 cells. (a) QC of MCF-7 RNA samples at different stages of heat-induced degradation was carried out by micro-capillary electrophoresis (gel-like image of the electropherogram in the left panel) and RT-qPCR evaluation of RPL-13A mRNA (right panel). (b) Representative fluorescence intensity scatter plots of non-degraded vs fragmented RNA samples. (c) Scatter plot data analyses showing the results of pair-wise comparison of the estrogen-induced gene expression changes measured with non-degraded vs degraded RNAs. Fold-change values of detected genes in samples showing different levels of RNA were plotted against those obtained with intact RNA samples. To improve visualization of the differences between samples, genes with an absolute fold change value lower than 1.2 were excluded from the plot. Dashed lines in (b and c) point to identity (x=y) between samples and solid lines the ‘best fit’ linear regression experimentally calculated. In each panel are also reported the r2 value resulting from comparison of the two (non-degraded vs degraded) samples and the number (n) of genes in common between all datasets used for statistical evaluations.
DASL analysis of these RNA samples was then performed in replicate with either 16 × BeadChip and 96 × SAM array formats, for a total of 36 arrays of which only one did not pass the array quality verification. All technical replicates showed a remarkable reproducibility, with a correlation index (r2) ranging between 0.95 and 0.99 (data not shown). Figure 1b shows the correlations between representative arrays of intact (0 min degradation time, see Figure 1a) vs degraded RNA samples. Upon increasing RNA fragmentation, the differences in signal intensity between all transcripts detected within a single sample decreased, with a progressive worsening in correlation with respect to the corresponding values measured in the same, non-degraded RNA. This is particularly true for 60 and 90 min samples, where the r2 value becomes 0.52 and 0.38, respectively (lower panels in Figure 1b). Interestingly, we observed a linear inverse correlation between RPL13A Ct measured by RT-qPCR and r2 values detected by array analysis. By comparing hybridization data, we found that a Ct value of 26.5, relative to the cellular RNA sample that underwent heat-induced fragmentation for 30 min, represents a threshold value for RNA samples apt for array analysis. In preliminary experiments, we observed that estrogen-induced expression changes of 45 responsive genes (including known hormonal targets such as AREG, BAK1, BRCA2, EGR1, ERBB2, TIMP3, MYB, PGR and TFF1) measured by DASL arrays showed a good correlation with the same measured by WG oligonucleotide microarrays, as both methods detected quantitatively and qualitatively comparable expression changes (data not shown). More important, when fragmented RNA samples showing RT-qPCR Ct values for RPL13A mRNA within or below this threshold were analyzed with DASL arrays, the gene expression changes measured were comparable to those observed with the same RNAs prior to fragmentation (Figure 1c, upper panels). This is in good agreement with what observed earlier for microarray analysis of RNA extracted from FFPE prostate cancer biopsies.19
DASL assay performance was further evaluated with an RNA sample extracted for three sequential 5 μm sections of a FFPE breast tumor sample (MT1). Even in this case, RNA quality was evaluated by micro-capillary electrophoresis and quantitative real-time RT-PCR of RPL13A mRNA. As expected,26 the FFPE tissue-derived RNA showed extensive degradation, revealed by loss of 16S and 28S ribosomal RNA bands and accumulation of low-molecular-wheight species (Figure 2a). A Ct value of 25.2 for RPL13A mRNA was calculated by RT-qPCR (time=0 in the left panel of Figure 2b). Aliquots of this RNA were subject to further fragmentation by in vitro exposure to high temperatures for up to 40 min. Heat-induced loss of RNA integrity in these samples was confirmed by electrophoretic analysis of the samples (data not shown) and RT-qPCR analysis of RPL13A mRNA, showing a linear increase in Ct values upon increasing time of exposure to heat (Figure 2b, left panel; data not shown). Replicate DASL assays were then performed on all these RNA samples. As documented in Figure 2b (right panel), the number of mRNAs detectable by DASL was constant up to the 20 min sample, after which it decreased linearly with the time of RNA exposure to high temperature. When the DASL gene expression profiles relative to these samples were confronted by cluster analysis, RNA samples exposed to heat ≤20 min (RPL13A Ct≤27.5) showed very small differences with respect to the starting RNA, indicating that under those condition the DASL arrays still allowed correct assessment of the RNA expression signature of the sample. Interestingly, 20 min exposure to 95°C already induced extensive RNA fragmentation (Figure 2b, left panel data not shown), suggesting that DASL can be applied to perform gene expression profiling with extensively degraded FFPE tissue-derived RNAs. Furthermore, these results indicate that it is possible to precisely determine RNA suitability for array analysis by a preliminary RT-qPCR test that provides quantitative assessment of a housekeeping gene mRNA detection levels in the sample, proving a handy and cost-effective pre-analytical screening method for exclusion of RNA samples that, because of their poor quality, would give rise to erroneous and misleading data (see below).
Evaluation of DASL array technology with RNA extracted from a FFPE MT biopsy and tested at different levels of fragmentation. (a and b) QC of RNA extracted from the FFPE MT biopsy MT1 was carried out by micro-capillary electrophoresis (gel-like image in the left panel and electropherogram in the right panel of a). Ctrl, control, non-degraded RNA. (b) The results of real-time RT-qPCR quantization of RPL-13A mRNA in MT1 RNA before and after heat-induced fragmentation (left panel) and of RNA analysis by DASL microarrays, expressed as number (n) of detected genes (left panel). The arrow points to the Ct limit value under which samples are still adequate for DASL array analysis (see text). (c) Hierarchical cluster analysis of the results obtained by gene expression profiling with DASL microarray of MT1-derived RNA before (0 min) and following heat-induced fragmentation, carried out for the indicated times. Data refer to average results of three, independent DASL analyses.
Application of DASL Arrays to Expression Profiling of FFPE Breast Cancer Specimens
Accuracy and reproducibility of the DASL assay applied to FFPE cancer specimens were then evaluated. To this aim, 17 paraffin blocks, corresponding to 15 distinct biopsies (with a storage age of 6–19 years), were used as source of RNA. The tests were carried out performing multiple RNA extractions from either three or eight sequential 5 μm sections of the same paraffin block and, in two cases, from two blocks derived from different portions of the same tumor, for a total of 48 independent extractions. RNA yield and quality were assessed in all cases by optical density (OD) and micro-capillary electrophoresis. Thirty-one such samples met the minimal quality pre-requisites for further use, that is, RNA yield of at least 25 ng/μl with OD260/OD280 ratio >1.8, according to criteria slightly more stringent than what previously proposed for similar studies.16 Such values were set based both on the electrophoretic profile of the starting RNA samples and the results of multiple test DASL assays, where the number of detected genes was taken as parameter for quantitative evaluation of array performance. It is worth noting that, in case of failure, inspection of Hematoxylin-eosin-stained microphotographs of a section from the same paraffin block helped explain most of the cases with low RNA yield was obtained, as we observed in such cases one or more of the following conditions: (i) small size of the tissue fragment included (<0.5–0.8 cm2); (ii) prevalence in the sample of fat tissue or (iii) presence of large patches of necrotic or fibrotic tissue (data not shown). Twenty-three of the 31 RNA samples mentioned above, corresponding to 8 distinct specimens (7 breast and 1 BLT) were included in this study. RNA integrity was evaluated by RT-qPCR of RPL13A mRNA as described above, which yielded Ct values ranging from 25.38 to 28.77. Furthermore, RIN—a parameter based on micro-capillary electrophoresis results that provides quantitative evaluation of RNA quality,22 was calculated for each sample. RIN values were found to range from 1.0 to 2.6. Each sample was then assayed on 3–6 array replicates, for a total of 101 arrays performed both on 16 × BeadChips and a 96 × SAM array formats.
Data shown in Figure 3a are representative of the results obtained with ‘good-quality’ samples, that is, RNAs that met all the QC criteria in our pre-test screening (see below). The upper-left panel of the figure shows a comparison between replicate arrays of the same RNA, while the upper-right panel shows between two RNAs extracted form either 3 (MT3_3 × _d) or 8 (MT3_8 × _d) sequential sections of the same paraffin block; the lower panel reports a comparison between two RNAs extracted from sections derived from two different paraffin blocks of the same tumor. In all cases, the r2 revealed a close proximity, to near identity, among the two expression profiles being compared. Furthermore, the very high number (n) of genes detected by both arrays, out of the 502 analyzed, demonstrates the good reproducibility and reliability of the assay. Our experience led us to set the following parameters for evaluation of measures obtained with replicate arrays: (i) results are unreliable when one or more (depending on the initial number of replicates) replica arrays show outlier values in its internal controls, indicating technical failure of the assay; (ii) the same applies when n is <200 and r2 between replicas <0.69; (iii) for arrays showing n between 200 and 250, data are acceptable only when r2 between replicates is ≥0.70; (iv) when r2 falls between 0.69 and 0.75, array data quality is good only if n is >250. In all other cases, gene expression data are to be considered reliable.
Evaluation of DASL array technology with RNA extracted from multiple FFPE MT biopsies and comparison with results obtained by analysis of RNA extracted from snap-frozen samples of the same tumors with ‘standard’ oligonucleotide microarrays. (a) Scatter plot data analyses showing the results of pair-wise comparison of replicate RNA samples extracted from two sets of three sequential sections from the same paraffin block (MT3_3 × _b vs _d, upper-left panel), from a set of three vs a set of eight sequential sections of the same (MT3_3 × _d vs 8_d, upper-right panel) or from two sets of three sequential sections from two different paraffin block prepared from different portion a tumor (MT2_DASL_A vs _B, lower panel). (b) Venn diagrams showing the results of pair-wise comparison relative to the number of genes detected with DASL vs WG microarrays in RNA extracted by FFPE and snap-frozen samples of three different MTs. Numbers between parentheses refer to the total number of transcripts, from the common list, detected with each microarray platform. (c) Scatter plot data analyses showing genes differentially expressed between tumor samples MT1 and MT2, as detected with WG (upper-left panel) or DASL (upper-right panel) microarrays. Data refer to 290 genes detected with both platforms in either tumor. The relative position of most genes remains the same in the two cases, whereas TOP2A and CXCL9 (encircled) showed opposite behavior with the two platforms. For these, the relative position of the three different probes present in the DASL arrays are shown in the lower panel. Dashed lines in (a and c) point to identity (x=y) between samples and solid lines the ‘best fit’ linear regression experimentally calculated. In each panel are also reported the r2 resulting from sample comparison and the number n of genes used for statistical evaluation.
Overall, 83% (84/101) successful hybridizations were achieved here, but array data quality and results were found to be highly dependent on the initial RNA sample status. In fact, when considering the percentage of success in replicate arrays from the same RNA sample, we observed that this varied between 40 and 100%, the main factor influencing the result being RNA quality. Indeed, apart from few tests that failed because of technical problems encountered during array set up and hybridization, in all cases we were able to link array performance to RNA degradation status, with arrays used to analyze ‘poor-quality’ RNAs showing a lower rate of success that those applied to ‘good-quality’ RNA. As RNA quality is a parameter that can be easily defined in a preliminary test, we suggest the following RNA QC criteria to define in advance suitability of a given sample for DASL assay: (i) RNA with RPL13A Ct<26.5 are most suited for DASL array analysis, whereas those with Ct>28 should be discarded; (ii) when RPL13A Ct is between these two values, the RNA sample should be further evaluated by micro-capillary electrophoresis to determine the RIN index; (iii) samples showing a RIN score >2.0 can be considered apt for analysis, whereas those with RIN≤2.0 are still suitable when the prevalent RNA size is >200 nt. By applying these simple QC criteria, we were able to obtain 100% DASL array success rate (data not shown).
To evaluate the functional significance of the gene expression profiles generated by DASL with RNAs from FFPE tissues, we set forth to compare the gene expression data obtained with this method with the same derived from WG oligonucleotide microarray analysis (see ‘Materials and Methods’ section for details). This comparison was carried out by performing expression profiling of RNA extracted from snap-frozen tissue samples from the MTs matching three FFPE samples (MT1, MT2 and MT5), analyzed by DASL. Results are displayed in Figure 3b and can be summarized as follows. First, a remarkable overlap in gene detection between the two methods was evident with all three samples, as 88–98% of the genes detected with WG arrays were also detected by DASL. Considering that the portion of tumor mass analyzed by WG and DASL were not the same, the slight differences in gene detection rate between the two assays can be easily explained with subtle variations in tissue composition and/or local gene expression patterns between the two samples. Second, DASL showed a considerably higher sensitivity than WG arrays, as the former could detect 21–34% more transcripts than the latter. This higher sensitivity may be explained by the exponential, specific cDNA amplification step included in the DASL protocol, which results in amplification only of the transcripts present in the gene panel for which the specific primers are designed. Thus, any targeted transcript of the gene pool will be amplified and detected by DASL arrays, even if present in very low amounts. By contrast, with standard WG array platforms all transcripts undergo linear amplification,27 so that rare transcripts may remain scarce in the final hybridization mixture, and for this reason undetectable.
As expression profiling is most useful in clinical oncology to pinpoint molecular differences and similarities among tumor samples, we asked whether DASL was reliable also for differential analysis between different MTs. As a comparison, we used once again the data derived from WG array analysis of frozen samples of the same tumors. In the upper panels of Figure 3c, a pair-wise comparison of the expression profiles detected by either method in breast tumors MT1 vs MT2 is shown. In this case, considering a common list of 290 genes detected in either tumor with both platforms, distance between samples, quantitatively expressed by the r2, was essentially the same with the two assays. On the other hand, for each transcript considered, the signal detected with the DASL array was significantly higher than that measured with WG arrays and compressed into a lower dynamic range, for the same reasons mentioned above. When considering the number of differentially expressed genes detected by the two platforms, we observed a slightly higher number of differentially expressed genes in the WG vs the DASL data set (59 vs 34 for MT1 vs MT2 comparison) and a good overlap between the two platforms (14 differentially expressed genes in common, in this same case). This last result should not surprise, as different methods for statistical evaluation had to be applied for data analysis in the two cases, due to the fact that dynamic ranges in the two platforms were different. Nevertheless, despite the remarkable difference in signal scale between the two data sets, most genes were positioned in similar regions of the scatter plots, indicating that comparable qualitative results were obtained with the two assays. This conclusion was confirmed by the results obtained by pair-wise comparisons of the expression profiles of breast tumors MT1 vs MT3 and MT2 vs MT3 detected with DASL and WG arrays (data not shown). Of the 14 differentially expressed genes detected in MT1 vs MT2 biopsies with both platforms, two (TOP2A and CXCL9, circled in Figure 3c) showed opposite behavior with the two microarrays. As DASL arrays included three probes for each of these transcripts, we controlled whether the discrepancy could be explained by one or more DASL probe failing to correctly detect the signal. As shown in the lower panel of Figure 3c, however, this was not the case, as signals relative to all three probes position, in both cases, near to each other in the scatter plot. It is worth noting that expression of these two genes was not very different between the two tumor samples being compared, as they showed a fold-difference <2.0, a value considered not significant in most gene expression studies. It is thus possible to conclude that the apparent discrepancy observed here between the two methods may depend upon differences in specific RNA levels intrinsic to the starting biological materials (snap-frozen and FFPE tissues, respectively), as already commented above.
Sensitivity of the DASL Assay for Detection of Differences in Gene Expression Patterns Between Tissue Samples
To evaluate sensitivity of the DASL array method implemented here for detection of differential gene expression between FFPE tissue samples, we exploited the differences in RNA profiles between BLT and MT (Figure 4a). To this aim, a sample dilution experiment was carried out whereby the RNA extracted from the FFPE breast tumor sample MT3 were mixed in various proportions (100:0, 80:20, 60:40, 40:60, 20:80 and 0:100) with the same extracted from a BLT sample. We then carried out duplicate DASL array analyses of each RNA pool generated in this way and looked for effectiveness of the method to correctly identify progressively divergent gene expression patterns. As shown in Figure 4b, cluster analysis of the data showed a remarkably high correlation between array and sample replicates, as already seen in previous experiments, but, more importantly, it revealed that the DASL assay could indeed discriminate and correctly identify slight differences in gene expression patterns between closely related samples, allowing to classify them accordingly.
Test of DASL assay sensitivity and ability to discriminate large and subtle differences in the gene expression patterns between FFPE-derived RNA samples. (a) Scatter plot data analysis showing significant differences in gene expression patterns between a breast (MT3) and a bladder (BLT) carcinoma. Dashed lines in a point to identity (x=y) between samples and solid lines the ‘best fit’ linear regression experimentally calculated. In the panel are also reported the r2 resulting from MT vs BLT sample comparison and the number n of genes detected in either tumor and used for statistical evaluation. (b) Sample dilution test. Data clustering of DASL microarray results obtained by analysis of RNA samples obtained by mixing with each other MT and BLT RNAs at various relative ratios.
Comparative gene expression analysis between breast and BLT RNAs (Figure 4a) highlighted two lists of differentially expressed genes (listed in Supplementary Table 2), the first represented by mRNAs highly expressed in breast tumors, including also estrogen receptor alpha mRNA (ESR1, Figure 5a), and the second including transcripts more abundant in BLTs (Figure 5b). When these gene lists were used to evaluate the array data generated with the sample dilution test described above, they both clearly showed how the pattern of gene expression correlated linearly, in each sample, with the RNA dilution factor (Figure 5). Indeed, we observed that genes completely absent in one tumor were detected also in as low as 20% of the original RNA, thus confirming the sensitivity and reliability of the DASL assay. This is a significant result in view of the fact that tumor biopsies may contain various amounts of cancer cells intermixed with cell of other origin (stromal, vascular, inflammatory, and so on) and that a gene-profiling study must rely on the possibility to detect cancer-specific mRNA expression profiles in extracts showing a wide range of relative abundance of tumor-derived RNA.
Heat-maps of results from the sample dilution test. The pattern of expression of a set of genes predominantly expressed in breast (a) or bladder (b) cancer (listed in Supplementary Table 2) are shown for each of the different samples described in Figure 4. Green and red represent low and high intensity of expression, respectively, whereas gray indicates absence of signal. Above each map are reported the relative color scales, referring to median-centered fluorescence intensity values.
DISCUSSION
Gene expression profiling of tumor biopsies promises to help improve clinical management of breast and other cancer patients by providing new ways to classify tumors and to predict disease outcome and responsiveness to therapy,28, 29 but its application to large-scale studies or in clinical settings has been, so far, seriously hampered by the fact that in most cases RNA extracted from these biopsies is degraded, and thus unsuitable for molecular analysis. This is particularly true for FFPE samples, which are of great potential usefulness for translational cancer research but provide only highly fragmented RNA.
We implemented here a technology for parallel quantitative analysis by oligonucleotide microarrays of a large number of transcripts in highly degraded RNAs extracted from FFPE breast and BLT biopsies that overcomes previously encountered limitations, and describe a pre-analytical RNA sample screening protocol providing QC parameters that, when performed, allow to obtain reproducible and biologically relevant gene expression results even with extensively fragmented RNAs. The results of this study provide a practical method for carrying out gene expression profiling analyses in FFPE MT samples by DASL oligonucleotide microarrays, which is applicable also to normal and pathological samples of other origin. Sensitivity, reproducibility and accuracy of the assay indicate that it is well suited for retrospective clinical studies aiming at identifying prognostic and predictive gene profiles in archival FFPE tissue banks and, predictably, also on laser-capture microdissected tissue samples, following an appropriate set up. Furthermore, the method described is cost-effective and not labor intensive, making it feasible analysis of even large numbers of samples at once. Furthermore, as the limiting factor for successful molecular profiling of the sample is excessive RNA fragmentation, protocols for RNA restoration in vitro, such as that described recently by Loudig et al,30 could be included in the assay when analysis of particularly informative, or unique, specimens is required.
The possibility of using FFPE tissue samples may allow also to combine expression profiling of defined sets of genes by DASL with array Comparative Genomic Hybridization and DNA methylation analysis31, 32 of the same biopsy, an approach that has been recently shown to be extremely powerful to identify functionally and clinically distinct MT subtypes.33
References
Sorlie T, Perou CM, Tibshirani R, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci USA 2001;98:10869–10874.
van de Vijver MJ, He YD, van't Veer LJ, et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 2002;347:1999–2009.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 2004;351:2817–2826.
Chang HY, Nuyten DS, Sneddon JB, et al. Robustness, scalability, and integration of a wound-response gene expression signature in predicting breast cancer survival. Proc Natl Acad Sci USA 2005;102:3738–3743.
Wang Y, Klijn JG, Zhang Y, et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 2005;365:671–679.
Fan C, Oh DS, Wessels L, et al. Concordance among gene- expression-based predictors for breast cancer. N Engl J Med 2006;355: 560–569.
O'Shaughnessy JA . Molecular signatures predict outcomes of breast cancer. N Engl J Med 2006;355:615–617.
Ein-Dor L, Zuk O, Domany E . Thousands of samples are needed to generate a robust gene list for predicting outcome in cancer. Proc Natl Acad Sci USA 2006;103:5923–5928.
Naderi A, Teschendorff AE, Barbosa-Morais NL, et al. A gene-expression signature to predict survival in breast cancer across independent data sets. Oncogene 2007;26:1507–1516.
Brenton JD, Carey LA, Ahmed AA, et al. Molecular classification and molecular forecasting of breast cancer: ready for clinical application? J Clin Oncol 2005;23:7350–7360.
Shi L, Reid LH, Jones WD, et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat Biotechnol 2006;24:1151–1161.
Cicatiello L, Scafoglio C, Altucci L, et al. A genomic view of estrogen actions in human breast cancer cells by expression profiling of the hormone-responsive transcriptome. J Mol Endocrinol 2004;32: 719–775.
Scafoglio C, Ambrosino C, Cicatiello L, et al. Comparative gene expression profiling reveals partially overlapping but distinct genomic actions of different antiestrogens in human breast cancer cells. J Cell Biochem 2006;98:1163–1184.
Teschendorff AE, Naderi A, Barbosa-Morais NL, et al. A consensus prognostic gene expression classifier for ER positive breast cancer. Genome Biol 2006;7:R101.
Weisz A, Basile W, Scafoglio C, et al. Molecular identification of ERalpha-positive breast cancer cells by the expression profile of an intrinsic set of estrogen regulated genes. J Cell Physiol 2004;200: 440–450.
Penland SK, Keku TO, Torrice C, et al. RNA expression analysis of formalin-fixed paraffin-embedded tumors. Lab Invest 2007;87: 383–391.
Karsten SL, Van Deerlin VM, Sabatti C, et al. An evaluation of tyramide signal amplification and archived fixed and frozen tissue in microarray gene expression analysis. Nucleic Acids Res 2002;30:E4.
Benchekroun M, DeGraw J, Gao J, et al. Impact of fixative on recovery of mRNA from paraffin-embedded tissue. Diagn Mol Pathol 2004;13:116–125.
Bibikova M, Talantov D, Chudin E, et al. Quantitative gene expression profiling in formalin-fixed, paraffin-embedded tissues using universal bead arrays. Am J Pathol 2004;165:1799–1807.
Fan JB, Gunderson KL, Bibikova M, et al. Illumina universal bead arrays. Methods Enzymol 2006;410:57–73.
Sorbello V, Fuso L, Sfiligoi C, et al. Quantitative real-time RT-PCR analysis of eight novel estrogen-regulated genes in breast cancer. Int J Biol Markers 2003;18:123–129.
Schroeder A, Mueller O, Stocker S, et al. The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol 2006;7:3.
Gentleman R, Carey V, Huber W . Bioinformatics and Computational Biology Solutions Using R and Bioconductor. Heidelberg: Springer, 2005.
Illumina Inc. BeadStudio User Guide, Doc 1117962 Rev. B., 2004, 2005.
Saeed AI, Sharov V, White J, et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 2003;34:374–378.
Cronin M, Pho M, Dutta D, et al. Measurement of gene expression in archival paraffin-embedded tissues: development and performance of a 92-gene reverse transcriptase-polymerase chain reaction assay. Am J Pathol 2004;164:35–42.
Van Gelder RN, von Zastrow ME, Yool A, et al. Amplified RNA synthesized from limited quantities of heterogeneous cDNA. Proc Natl Acad Sci USA 1990;87:1663–1667.
Gruvberger-Saal SK, Cunliffe HE, Carr KM, et al. Microarrays in breast cancer research and clinical practice—the future lies ahead. Endocr Relat Cancer 2006;13:1017–1031.
Morris SR, Carey LA . Gene expression profiling in breast cancer. Curr Opin Oncol 2007;19:547–551.
Loudig O, Milova E, Brandwein-Gensler M, et al. Molecular restoration of archived transcriptional profiles by complementary-template reverse-transcription (CT-RT). Nucleic Acids Res 2007;35:e94.
Joosse SA, van Beers EH, Nederlof PM . Automated array-CGH optimized for archival formalin-fixed, paraffin-embedded tumor material. BMC Cancer 2007;7:43.
Mehrotra J, Vali M, McVeigh M, et al. Very high frequency of hypermethylated genes in breast cancer metastasis to the bone, brain, and lung. Clin Cancer Res 2004;10:3104–3109.
Chin K, DeVries S, Fridlyand J, et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 2006;10:529–541.
Acknowledgements
Research supported by Italian Ministry of University and Research (PRIN 2005063915_003 and 2006069030_003), Italian Ministry of Health (Research Grants 2003, 2005); Regione Piemonte (Ric. Sanitaria e Scientifica Applicata, Grants 2006 and 2007), UE (CRESCENDO IP, contract no. LSHM-CT2005-018652), AIRC (Italian Association for Cancer Research) and PhD Programs: ‘Pathology of Cellular Signal Transduction’ of the Second University of Naples (O Paris, R Tarallo and OMV Grober) and ‘Toxicology, Oncology and Molecular Pathology’ of the University of Cagliari (M Ravo). We thank Paola Bontempo, Nicoletta Raverino, Raffaele Rossiello, Ivana Sarotto and the pathologists of the IRCC Hospital of Candiolo (TO), Mauriziano ‘Umberto I’ Hospital of Turin and Second University of Naples for the supply of tumor tissues and their assistance in collection of FFPE samples for analysis. M Mutarelli is recipient of a Post-doctoral Fellowship of the Second University of Naples and O Paris of a Doctoral Fellowship from the AIRC Naples Oncogenomics Center.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website (http://www.laboratoryinvestigation.org)
Disclosure
The authors have no conflict of interest.
Supplementary information
Rights and permissions
About this article
Cite this article
Ravo, M., Mutarelli, M., Ferraro, L. et al. Quantitative expression profiling of highly degraded RNA from formalin-fixed, paraffin-embedded breast tumor biopsies by oligonucleotide microarrays. Lab Invest 88, 430–440 (2008). https://doi.org/10.1038/labinvest.2008.11
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2008.11
Keywords
This article is cited by
-
DNA methylation markers predict recurrence-free interval in triple-negative breast cancer
npj Breast Cancer (2020)
-
Clinical characteristics and surgical outcome in USP8-mutated human adrenocorticotropic hormone-secreting pituitary adenomas
Endocrine (2019)
-
Early immune modulation by single-agent trastuzumab as a marker of trastuzumab benefit
British Journal of Cancer (2018)
-
Gene expression modules in primary breast cancers as risk factors for organotropic patterns of first metastatic spread: a case control study
Breast Cancer Research (2017)
-
Reliable gene expression profiling of formalin-fixed paraffin-embedded breast cancer tissue (FFPE) using cDNA-mediated annealing, extension, selection, and ligation whole-genome (DASL WG) assay
BMC Medical Genomics (2016)