Abstract
Clinical tissue specimens are useful for pathological diagnosis, which is, in some cases, supported by visualization of biomolecule localization. In general, diagnostic specificity in molecular pathology is increased by the acquisition of a probe to distinguish the modification of isomers. Although glycosylation is one of the candidate modifications in a protein, comparative glycan analysis of disease-associated proteins derived from a single tissue section is still challenging because of the lack of analytical sensitivity. Here we demonstrate a possible method for differential glycoform analysis of an endogenous tumor-associated glycoprotein MUC1 by an antibody-overlay lectin microarray. Tissue sections (5 μm thick) of patients with cholangiocarcinoma (CCA; n=21) and pancreatic ductal adenocarcinoma (PDAC; n=50) were stained with an anti-MUC1 antibody MY.1E12 that was established as a monoclonal antibody recognizing an MUC1 glycosylation isoform with a sialyl-core 1 structure (NeuAcα2-3galactosyl β1-3-N-acetylgalactosamine). MY.1E12-positive tissue areas (2.5 mm2) were selectively dissected with a laser capture microdissection procedure. The membrane MUC1 was enriched by immunoprecipitation with MY.1E12 and subjected to lectin microarray analysis. Even though the reactivities of MY.1E12 between CCA and PDAC were similar, the lectin-binding patterns varied. We found Maackia amurensis leukoagglutinin and pokeweed lectin distinguished MY.1E12-reactive MUC1 of CCA from that of PDAC. Moreover, MUC1 with M. amurensis hemagglutinin (MAH) reactivity potentially reflected the degree of malignancy. These results were confirmed with MAH-MY.1E12 double fluorescent immunostaining. These glycan changes on MUC1 were detected with high sensitivity owing to the cluster effect of immobilized lectins on a tandem repeat peptide antigen covered with highly dense glycosylation such as mucin. Our approach provides the information to investigate novel glycodynamics in biology, for example, glycoalteration, as well as diseases related to not only MUC1 but also other membrane proteins.
Similar content being viewed by others
Main
Recent high-throughput technologies for (macro-)biomolecules have developed considerably in terms of high resolution and high sensitivity, thereby facilitating comprehensive and comparative analysis (-omics) using clinical specimens.1, 2 Based on integrative approaches to understanding functional molecules, we have realized the strategic development of therapeutic targets and biomarkers.
Glycan is the so-called third bio-chain that in nature conjugates with the other bio-chains, proteins, and lipids. For the discovery of glycan-based biomarkers (glyco-biomarkers), glycoproteomics and glycomics that target a specific glycoprotein can be used for comprehensive and comparative analyses. Indeed, such approaches were reported as important studies for unsolved biological events,3 and thereafter several comprehensive analysis tools have been developed.4, 5 However, the comparative detection of characteristic glycosylation in an analyte glycoprotein located in a histopathologically specified area of clinical specimens is still challenging.
Membrane-associated mucin-type 1 (MUC1) is a well-known O-glycosylated protein that localizes to the apical membrane of normal secretory epithelial cells and contributes to the protective mucous gel through ectodomains of heavily O-glycosylated tandem repeats.6, 7, 8 In various cancers, aberrant expression and glycosylation of MUC1 have been demonstrated to have important roles in many events, such as proliferation, invasion, and metastasis.9, 10, 11 For example, aberrant MUC1 glycosylation facilitates integrin clustering by funneling active integrins into adhesions and altering integrin state by applying tension to matrix-bound integrins.9 This phenomenon leads to metastasis of tumor cells by mechanically enhancing cell-surface receptor function. Another study indicated that the highly varied sialylated structures of MUC1 contribute to adhesion in pancreatic cancer perineural invasion.10 These studies clearly show that MUC1 glycosylation has been regarded as an essential target for cancer therapeutic and diagnostic agents.9, 10, 11 Therefore, to satisfy such clinical needs, comprehensive analysis of O-glycoform on MUC1 from clinical specimens is an important key. Nevertheless, there are few reports of comprehensive analysis of O-glycoform even in MUC1, which is a typical O-linked glycoprotein, because quantification of the small amount of MUC1 glycoform derived from clinical specimens (eg, serum, bile, and tissue sections) is still difficult even with current glycomic technologies employing mass spectrometry (MS).12 Development of MS technology has facilitated glycomics research and identification of glyco-biomarkers.13, 14 However, MUC1 glycoform analysis remains difficult because mucins are modified heavily with O-glycosylation and are generally protease-resistant. Moreover, complete MUC1 purification from clinical samples is difficult because MUC1 components are complex with various proteins and other mucins.15 Recently, imaging MS technology using desorption/ionization MS (MALDI MS) has been developed to visualize biomolecules in clinical tissue sections16 and there are some reports on the imaging MS analysis of N-linked glycan expression in tissue sections.17, 18, 19, 20 However, there are no reports for O-linked glycan analysis because this technology depends on glycan digestion with N-glycosidase and there are no O-glycosidases for the specific digestion of O-linked glycans. Therefore, a robust quantitative and qualitative methodology with high sensitivity and high throughput is required for comprehensive analysis of MUC1 O-glycoforms from various clinical samples. Compared with technologies based on MS, the lectin microarray-based glycan profiling method has the sensitivity to obtain the glycan profile of the tiny amounts of analytes derived from tissue sections. Moreover, this technology can obtain the profiles of both N- and O-glycans in diverse complex biological samples without releasing the glycan moieties from glycoproteins.21, 22, 23, 24
In this study, we examined the reliability of differential O-glycoform analysis targeting membrane-tethered MUC1 isolated from 21 cholangiocarcinoma (CCA) and 50 pancreatic ductal adenocarcinoma (PDAC) formalin-fixed paraffin-embedded (FFPE) tissue specimens. To explore the relationship of MUC1 glycoform with characteristics of tumors in clinical tissue sections, we adopted a key basic methodology, namely lectin-assisted glycan profiling using a microarray-based, non-MS system.21, 22, 25, 26 Although both CCA and PDAC sections were stained strongly with an anti-MUC1 antibody MY.1E12 that specifically binds to the sialylated O-glycoform, the results from glycan profiles drawn by the binding signals of 45 lectins enable the MUC1 glycoforms between CCA and PDAC to be distinguished. The results suggest that the difference in the glycoform of MUC1 may be related to the difference in tumors and tumor behaviors. To understand the relationship between aberrant O-glycosylation and various diseases, we propose an innovative methodology to obtain MUC1 O-glycoform information based on the histopathological characteristics of FFPE clinical tissue sections. Our approach will facilitate the development of novel therapeutic targets and biomarkers associated with aberrant O-glycosylation.
Materials and methods
Clinical Samples
Surgically resected tissues of mass-forming-type CCA from 62 patients and those of mass-forming-type PDAC from 126 patients were prepared. All tissue specimens were retrieved from the archive of the Department of Pathology, Faculty of Medicine, Kagoshima University, Japan, and informed consent for the use of clinical specimens was obtained from all participants. All patients were also involved in a previous study.27, 28 All specimens were FFPE, and cut into 5-μm-thin sections for hematoxylin and eosin (HE) staining, followed by immunohistochemistry and laser capture microdissection (LCM). The tissue specimens were selected for an antibody-overlay lectin microarray based on the criteria described in Supplementary Figure S1. Consequently, differential glycan profiling was performed on 21 CCA and 50 PDAC sections. Clinicopathological data of the subjects are described in Table 1. The pathological stages of the patients were assessed according to the Union for International Cancer Control Tumor, Node, Metastasis Classification.29
Immunohistochemistry with MY.1E12 Staining
To visualize MUC1-positive cells in FFPE tissue sections, immunohistochemistry was performed with an antisialylated MUC1 monoclonal antibody MY.1E12, which recognizes MUC1 with a sialyl-core 1 structure (NeuAcα2-3Galβ1-3GalNAc).30, 31 Immunohistochemistry was performed by a standard procedure with an immunoperoxidase method using an avidin–biotin complex as described previously.32 In brief, each section was deparaffinized with xylene and rehydrated using an alcohol gradient followed by submersion in tap water. The antigen was retrieved using 10 mM citrate buffer and autoclaved for 10 min at 115 °C. Endogenous peroxidase activity was quenched with 0.3% hydrogen peroxide in absolute methanol and then blocked with 2% horse serum in phosphate-buffered saline (PBS) to prevent nonspecific staining. Following incubation with MY.1E12 (1 μg/ml in PBS with 1% bovine serum albumin (BSA)) and biotinylated goat anti-mouse IgG, all sections received avidin–biotinylated peroxidase complex (Vector Laboratories, Burlingame, CA) for 30 min. 3,3′-Diaminobenzidine substrate was added and counterstained with hematoxylin.
Preparation of Tissue Fragments by LCM
LCM of surgical FFPE sections was performed to recover specific cell populations using Laser Micro Dissection System 6000DM (Leica Microsystems, Wetzlar, Germany). A portion of the FFPE serial sections (5 μm thick) from CCA and PDAC patients was thaw-mounted on a film-coated glass slide (PEN-Membrane, Leica Microsystems). The MY.1E12-positive cells in the tissue were visualized with standard HE stain and specified by microscopic observation, followed by microdissection. Dissected tissue fragments were collected from an area of 2.5 mm2 within the MY.1E12-positive cells using LCM.
Protein Extraction
Protein extraction from dissected tissue fragments was performed following previous reports.23, 24 In brief, tissue fragments were collected into microtubes and 10 mM citrate buffered saline (pH 6.0) was added, and the fragments were then heat denatured at 95 °C for 30 min for antigen retrieval. After centrifugation at 20 000 g for 5 min at 4 °C, tissue fragments were washed with PBS. After the PBS was discarded, PBS containing 0.5% NP-40 was added to the tissue fragments and then sonicated gently. The resultant solutions were incubated for 1 h on ice and the supernatants were collected as protein extraction samples.
Immunoprecipitation of Sialylated MUC1 with MY.1E12
For the antibody-overlay lectin microarray, sialylated MUC1 was purified from the protein extraction samples described above as previously reported.26 Sialylated MUC1 was immunoprecipitated using biotinylated antibody-conjugated streptavidin magnetic beads (Dynabeads, Life Technologies, Waltham, MA, USA). MY.1E12 was biotinylated with Biotin Labeling Kit-NH2 (Dojindo Laboratories, Kumamoto, Japan). Biotinylated MY.1E12 (500 ng) was incubated with streptavidin beads in 20 μl of PBS containing 1% Triton X-110 (PBSTx) for 1 h at 4 °C. The beads conjugated with MY.1E12 were washed with PBSTx, and 40 μl of the sample (diluted with PBSTx) was added to the beads. The antigen and MY.1E12 were incubated for 1 h at 4 °C. After the beads were washed with PBSTx, 10 μl of PBS containing 0.2% SDS was added to the beads and the bound material was eluted by heat denature for 10 min at 95 °C. For the lectin microarray analysis, contaminant biotinylated MY.1E12 was completely depleted by incubation with 40 μl of the streptavidin beads for 1 h at 4 °C. The supernatants were collected as immunoprecipitated samples.
Antibody-Overlay Lectin Microarray
The antibody-overlay lectin microarray was performed as described previously.22, 26 In brief, immunoprecipitated sialylated MUC1 from the protein extraction samples described above was diluted with PBSTx and then applied to the lectin array slide, LecChip (GlycoTechnica, Yokohama, Japan) containing triplicate spots of 45 lectins (Supplementary Figure S2). After overnight incubation at 20 °C, human serum polyclonal IgG (20 μg) was added and incubated for 30 min at 20 °C. After the slide was washed three times with PBSTx, 60 μl of the biotinylated MY.1E12 solution (100 ng) in PBSTx was applied to the array and then incubated for 1 h at 20 °C. After the slide was washed three times with PBSTx, 100 ng of a Cy3-labeled streptavidin (GE Healthcare UK, Little Chalfont, UK) solution in PBSTx was added to the slide and then incubated for 25 min at 20 °C. The slide was washed with PBSTx and scanned with an evanescent-field fluorescence scanner (GlycoStation Reader 1200, GlycoTechnica). All data were analyzed with Array-Pro Analyzer, version 4.5 (Media Cybernetics, Bethesda, MD, USA). The net intensity of each spot was calculated by subtracting the background value from the total signal intensity of three spots. The mean lectin signals of triplicate spots were normalized to the mean value of 45 lectins immobilized on the array as described previously.33
Lectin-MY.1E12 Double Fluorescent Immunostaining
To investigate the localization of lectin epitopes and sialylated MUC1, we performed lectin-MY.1E12 double immunostaining with fluorescence on the CCA and PDAC FFPE tissue sections as in a previous study.25 After deparaffinization, the sections were washed with PBS and then soaked in 10 mM glycine-PBS to quench free aldehyde residues. Blocking was performed by incubation with Carbo-Free Blocking Solution (Vector Laboratories) for 30 min at room temperature. After washing with 10 mM glycine-PBS, tissue sections were incubated with biotin-conjugated Maackia amurensis lectin (biotin-MAL, 10 μg/ml: diluted with 10 mM HEPES buffer, Vector Laboratories), biotin-conjugated Phytolacca americana lectin (biotin-PWM, 20 μg/ml: diluted with 10 mM HEPES buffer, Vector Laboratories), biotin-conjugated Wisteria floribunda agglutinin (biotin-WFA, 5 μg/ml: diluted with 10 mM HEPES buffer, Vector Laboratories), and biotin-conjugated M. amurensis agglutinin (biotin-MAH, 10 μg/ml: diluted with 10 mM HEPES buffer, Vector Laboratories) for 1 h at room temperature for MAL, WFA, and MAH or overnight at 4 °C for PWM. After washing with 0.1% BSA in PBS (BSA-PBS), the sections were incubated with Alexa Fluor 594-conjugated streptavidin (1 μg/ml diluted with PBS, Life Technologies) for 1 h and then washed with 0.1% BSA-PBS and incubated with MY.1E12 (1 μg/ml: diluted PBS) for 1 h at room temperature. After washing with 0.1% BSA–PBS, the sections were incubated with Alexa Fluor 488-conjugated anti-mouse IgG antibody (1 μg/ml: diluted with PBS, Life Technologies) for 45 min at room temperature. The sections were then incubated with Hoechst 33258 (Dojindo Molecular Technologies, Kumamoto, Japan) to stain the nuclei for 20 min at a dilution of 1:500. They were then washed with PBS and sealed by mounting with ProLong Diamond Antifade Mountant (Life Technologies) and were observed using fluorescence microscopy (BZ-X700, Keyence, Osaka, Japan).
Statistical Analysis
Mann–Whitney U-test was used to compare the clinicopathological data between CCA and PDAC. Postoperative survival was analyzed with the Kaplan–Meier method and the differences in the survival curves were compared with log-rank test. These calculations and graph constructions were performed with GraphPad Prism version 5.00 for Windows (GraphPad Software, San Diego, CA, USA). We performed hierarchical clustering analysis (HCA) and principal component analysis (PCA) for multivariable data using JMP Pro 11 (SAS Institute, Cary, NC, USA).
Results
LCM-Assisted Glycan Profiling of Membrane-Tethered MUC1 with Antibody-Overlay Lectin Microarray from FFPE Tissue Sections
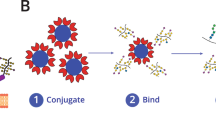
The primary objective of this study was to obtain the glycan profile of MUC1 from FFPE tissue sections by a lectin microarray. Figure 1a illustrates an overview of the sample processing from LCM to MUC1 glycan profiling with an antibody-overlay lectin microarray. In brief, we prepared three FFPE tissue sections of a typical MUC1-producing tumor from a patient with CCA. One of the three sections was stained with HE for clinicopathological observation. Another serial section was stained with antisialylated MUC1 monoclonal antibody, MY.1E12, for confirmation of MUC1-expressing cells. The remaining serial section was used for tissue dissection by LCM. We collected tissue fragments of tumor cells from the MUC1-positive area with LCM under microscopy (Figure 1b). Glycoproteins were extracted from the MUC1-positive (containing tumor cells) and MUC1-negative (containing non-tumor cells) tissue fragments as described in Materials and Methods section. After the immunoprecipitation of MUC1 from the extracted glycoprotein mixtures, MUC1 was applied to the lectin microarray slides, and significant glycan signals were detected only in the MUC1-positive cancer cells (Figure 1c). Finally, we succeeded in obtaining a distinctive glycan profile of MUC1 from CCA patients using FFPE tissue sections of MUC1-positive CCA cells.
Overview of laser capture microdissection (LCM) targeting MY.1E12-positive cells in cholangiocarcinoma (CCA) tissue sections and glycan profiling with MY.1E12-overlay lectin microarray. (a) Scheme of MUC1 glycan profiling from the targeted tissue sections. (b) Typical image of CCA tissue stained with MY.1E12 (left panel). MY.1E12 stain and tissue slides after LCM of the dissected tissue areas (right panels). (c) Scan images and bar graphs of MUC1 glycan profiling of tissue fragments from tumor (i) and non-tumor (ii) regions. Details of the 45 lectins on LecChip and the platform are available in Supplementary Figure S2.
Differential Glycan Profiling of MUC1 in FFPE Tissue Sections from 21 CCA and 50 PDAC Patients
We performed differential glycan profiling to investigate the difference of MUC1 glycoforms between CCA and PDAC using MUC1-positive FFPE tissue sections from 21 CCA and 50 PDAC patients. In general, CCA and PDAC are categorized mainly into three histological types by their growth pattern: intraductal growth type, periductal infiltrating type, and mass-forming type.34, 35 In this study, we used the mass-forming type only to suppress histological variation between CCA and PDAC because the mass-forming type is the most common among the three. The clinical information of CCA and PDAC patients is shown in Table 1. We measured the 45 lectin signals for MUC1 from all CCA and PDAC FFPE tissues and normalized them relative to the mean signal intensity (Figure 2a). HCA was used to discriminate CCA from PDAC by their glycan profiles (Figure 2b). Although the HCA of 71 patients partly overlapped, the MUC1 glycan profiling with 45 lectins classified CCA and PDAC patients into several clusters. This suggested that MUC1 glycoforms from CCA and PDAC origins are different. Subsequent PCA on 45 lectin signal intensities showed that the principal components (PCs) 1 and 2 appeared to correlate to CCA toward PDAC (Figure 2c), although it also showed that some of the CCA and PDAC patient profiles overlapped, possibly because the MUC1 glycoforms of CCA origin seem to vary more than those of PDAC. We then prepared a biplot and bar graphs of lectin replications based on the PCA to investigate specific glycan structures on MUC1 in CCA and PDAC to find lectins that contribute to these PCs (Figure 2d). The data showed that CCA was influenced by various glycans, while PDAC was mainly influenced by modifications in NeuAcα2-3Gal (MAL, M. amurensis leukoagglutinin) and terminal fucose (AOL, Aspergillus oryzae lectin; Aleuria aurantia lectin). To our knowledge, this is the first experimental proof that glycoforms of MUC1 derived from CCA and PDAC have characteristic differences.
Differential glycan profiling of sialylated MUC1 and multivariable analysis between 21 cholangiocarcinoma (CCA) and 50 pancreatic ductal adenocarcinoma (PDAC) patients. (a) Glycan profiles of 21 CCA and 50 PDAC patients listed in Table 1 with 45 lectins and a typical image of immunostaining with MY.1E12. The horizontal line inside each box represents the median, and the lower and upper borders of the box encompass the interquartile range. Vertical lines from the ends of each box encompass the 5th and 95th percentiles. (b) Hierarchical clustering analysis using the glycan profiling data of all CCA and PDAC formalin-fixed paraffin-embedded tissue samples. The heat map shows a two-way cluster analysis carried out on the data of the tissue specimens listed in Table 1. The data were obtained as averages of multiple tissue samples. Levels of lectin signals are indicated by color changes from blue (high expression level) to red (low expression level). CCA and PDAC patients are indicated by blue and red lines, respectively. (c) Principal component analysis of glycan profiling of MUC1 in 21 CCA and 50 PDAC patients. CCA and PDAC are indicated in red and blue, respectively. (d) Relative intensity of 45 lectins shown as a biplot and bar graphs.
Selection of Lectin Probes That Bind to Specific MUC1 Derived from CCA and PDAC
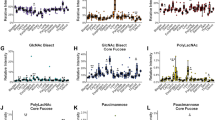
As the MUC1 glycoforms of CCA and PDAC origins appeared to be different, we performed further univariate analysis to obtain an overview of the lectins that show the greatest difference of signal patterns between CCA and PDAC samples. We have preliminarily analyzed all lectins for those that are significant in discriminating CCA from PDAC (Supplementary Table S1). In addition, from the result of the PCA (Figures 2c and d), we selected nine lectins showing significant differences between CCA and PDAC. In the comprehensive lectin profiles, seven (Figure 3a) and two (Figure 3b) lectins were identified as showing increased intensities in CCA and PDAC samples, respectively (P<0.05). We investigated the PWM lectin from P. americana, a polylactosamine binder, in CCA (area under the receiver operating characteristic curve (AUC)=0.8388, P<0.0001) and MAL in PDAC (AUC=0.8146, P<0.0001) as the best lectins showing the most significant statistical scores (P<0.0001, Table 2).
Dot graphs of nine lectins showing a significant difference in glycan profiling between cholangiocarcinoma (CCA) and pancreatic ductal adenocarcinoma (PDAC) patients. (a) Dot plots of seven lectins (PWM, PHA-E, BPL, DSA, ECA, STL, and UDA) with a significant increase in CCA patients. (b) Dot plots of two lectins (MAL and AOL) with a significant increase in PDAC patients.
Lectin and MY.1E12 Staining to Confirm Localization in FFPE Tissue Sections from CCA and PDAC Patients
To investigate the localizations of lectin-reactive glycans and MUC1 in tissues, we performed double fluorescent immunostaining (Figure 4) with MY.1E12 and the lectins that showed significant signals (PWM and MAL) in CCA or PDAC FFPE tissue sections in Figure 3. WFA-MY.1E12 was used as a positive control because we have already reported its localization by double fluorescent immunostaining in CCA samples.25 In WFA-MY.1E12 staining, we could observe colocalization of WFA epitopes and MUC1 in both CCA and PDAC cancer cells as predicted (lower panels in Figure 4). In MAL-MY.1E12 staining of CCA, although MY.1E12 stained specifically on the apical surface of cancer cells, MAL stained in the stroma around the cancer cells and did not stain on the apical surface of the cancer cells. Therefore, stains of MAL-MY.1E12 were separated completely in CCA samples. However, MAL epitopes expressed well in PDAC cancer cells, and colocalization of MY.1E12 and MAL was observed in the cancer cells of PDAC samples. In contrast, colocalization of PWM and MY.1E12 was observed specifically in CCA rather than in PDAC. These results agree with those in Figure 3 and show that, although the expression levels of MUC1 from CCA and PDAC samples were comparable, their glycosylation profiles are quite different between CCA and PDAC. These results may provide useful information to develop a glyco-biomarker for differential diagnosis between CCA and PDAC.
Lectin-MY.1E12 double fluorescent immunostaining of cholangiocarcinoma (CCA) and pancreatic ductal adenocarcinoma (PDAC) formalin-fixed paraffin-embedded sections. Double fluorescent immunostaining with two lectins (MAL and PWM) shows significant differences between CCA and PDAC and MY.1E12 and between WFA and MY.1E12 as a control. Each lectin was labeled with Alexa 592 streptavidin (red), and MY.1E12 was labeled with Alexa 488 anti-mouse IgG (green). Marginal areas are shown in yellow. Nuclei are visualized with Hoechst 33258 (blue). Each bar indicates 20 or 200 μm. HE staining of tissue specimens using double fluorescent immunostaining is shown in Supplementary Figure S4.
Relationship of MUC1 Glycosylation Change with Survival in CCA Patients
We investigated aberrant glycosylation in MUC1 related to cancer properties, such as tumor stages and survival. To assess lectins related to prognosis, we investigated the correlation between the 45 lectin signals and survival periods after surgery in 21 CCA patients. MAH, a NeuAcα2-3 or desialyl-T (NeuAcα2-3Galβ1-3[NeuAcα2-6]GalNAcα) antigen binder,36, 37 was the lectin that showed a significant correlation with the survival period (P=0.0463) and tumor stages (Figures 5a and b). In analysis of the cumulative survival rate using the Kaplan–Meier method, high reactivity of MAH (relative intensity >0.5) on MUC1 was also significantly related to a poor prognosis (P=0.0002; Figure 5c). To verify these results, we performed double fluorescent immunostaining with MAH and MY.1E12. The MAH and MY.1E12 stains merged well on the surgical tissue sections of patients with a poor prognosis (patient 19, high MAH signal) but did not merge in patients with a good prognosis (patient 12, low MAH signal) (Figure 5d). These results suggest that the increase of MAH signal on MUC1 relates to prognosis in CCA patients.
Relationship between prognosis of cholangiocarcinoma (CCA) patients and aberrant MUC1 glycosylation. (a) 2D plot of the survival period after surgical dissection and M. amurensis hemagglutinin (MAH) signal indicating the most significant correlation with the survival month. (b) Relationship of MAH signal and tumor stage. (c) Kaplan–Meier analysis of MAH signal intensity. Analysis was performed with a MAH signal cutoff value of 0.5. (d) MAH-MY.1E12 double fluorescent immunostaining of CCA FFPE tissue sections from patients 12 and 19 shown in panel (a) and Supplementary Table S1. The clinical stages and survival periods were stage III and 26 months in patient 12 and stage IVa and 6 months in patient 19, respectively.
Discussion
Aberrant glycosylation of MUC1 during cancer development is well known to contribute to various cancer events. Although many studies have attempted the clinical application of MUC1 glycosylation,38, 39, 40 such attempts have been unsuccessful. This is largely because of the difficulty in qualitative analysis of glycosylation targeting small amounts of MUC1 derived from clinical specimens such as body fluids and tissue sections even with the recent development of glycan analysis technologies. Thus a robust methodology is required for qualitative analysis of the MUC1 glycoform. In this study, we described the establishment of a qualitative method of the MUC1 glycoform, namely, glycan profiling, with high sensitivity and throughput targeting in FFPE tissue sections and its application in differential glycan profiling between MUC1-producing cancers such as CCA and PDAC. Some studies have reported on MUC1 glycan analysis derived from serum and culture cells.12, 38, 39, 40 However, although it is important to analyze clinical tissue sections for the development of tumor markers and to understand the tumor biology more deeply, no reports have been published on the direct analysis of MUC1 glycan in FFPE tissue sections. Glycan analysis in FFPE tissue sections using MS imaging technology has been reported. However, as described in the Introduction section, the detection sensitivity could not determine the localization of glycan epitopes, including O-glycosylation, on specific glycoproteins in FFPE tissue sections. Therefore, a reliable methodology with high sensitivity was required to qualify O-glycosylation of mucins in FFPE tissue sections. LCM is ideal for isolating a specific population of cells, and several different types of well-designed devices are commercially available.41 Although LCM has contributed to the proteomics approach for tissue analysis targeting FFPE tissue sections and has facilitated molecular cancer biomarker discovery,42 the use of LCM in glycome analysis containing O-glycosylation has not been reported. We previously established that the lectin microarray system is a highly sensitive and high-throughput glycan profiler.21 The antibody-overlay lectin microarray also facilitates glycan-associated biomarker discovery focusing on distinctive glycoproteins.22 In this study, we attempted to obtain MUC1 glycan profiles in MUC1-positive cancer cells in FFPE tissue sections using lectin microarrays combined with LCM technology. Significant advantages of our approach include: (1) highly sensitive and high-throughput performance with simplicity compared with other technologies such as MS; (2) multiplex analysis using 45 lectins with defined binding specificities concurrently covering both N- and O-glycans; and (3) glycan profile of MUC1 can be obtained from a small amount of FFPE tissue because the target lesion for analyzing the MUC1 glycoform was selected specifically by visualization of MUC1-positive cancer cells with immunohistochemistry assisted by microdissection. Consequently, we achieved the MUC1 glycoform derived from an area of 2.5 mm2 in 5 μm thickness in FFPE tissue sections. Our approach is thus novel and unique for the direct qualification of MUC1 O-glycoforms in clinical tissue sections and indicates that MUC1 O-glycoforms of CCA and PDAC are quite distinctive.
CCA and PDAC are malignant tumors associated with poor prognosis. Histologically, both carcinomas are represented by similar infiltrating ductal architecture and mild-to-moderate nuclear atypia. This essentially causes important clinical problems because differential diagnosis between CCA and PDAC has significant implications for surgical management, chemotherapy, and patient prognosis.43, 44, 45 It is difficult to distinguish CCA from PDAC based on the protein compositions because the expression patterns of mucins, including MUC1, MUC2, MUC4, MUC5AC, and MUC16, are similar in these carcinomas.27, 28, 32, 45 Our data also indicate that the expression of MUC1 is not different immunohistochemically between CCA and PDAC (Supplementary Figure S3). These results well represent our concept that the glycan moieties on the proteins are different even if the protein levels are the same regardless of the type of cancer or clinical conditions.46 Moreover, specific glycosylation profiles of MUC1 are considered to be biomarkers for various cancers,47 and several groups have reported that glycosylation is useful as a cancer marker.12, 38, 48 Our technology may contribute to detecting specific glycosylation profiles of MUC1 related to various cancer events. In addition, although the sample size was limited, we could identify MUC1 glycosylation related to the prognosis of CCA in Figure 5. Our data showed clearly that the lectin MAH indicated a change of signal intensity due to tumor prognosis. This result experimentally proved an accepted theory using clinical tissue that the heavy sialylation of MUC1 is associated closely with cancer prognosis through invasion and metastasis.10, 48 Previously, we developed WFA-positive sialylated MUC1 as a highly sensitive CCA diagnostic marker in serum and bile specimens.25, 26, 49, 50 Unfortunately, detection of PWM-MUC1, MAL-MUC1, and MAH-MUC1 in serum is difficult compared with detection of WFA-MUC1 because the results indicate their levels are comparatively low (Figure 4). Thus, to determine their feasibility as a serum marker, a highly sensitive detection system is required.
In this study, we showed a dramatic change of glycosylation on MUC1 related to the cancer type or cancer progression. MAL and MAH bind to sialylated glycans such as NeuAcα2-3Gal with little different binding specificities. Because of structural differences in the binding pockets among these lectins, MAH recognizes O-glycans while MAL prefers N-glycans. Maenuma et al.37 reported that MAH preferentially binds to disialyl-core 1 antigen, 3′-sulfated core 1 (HSO3-3Galβ1-3GalNAcα), and 3′-sulfated Lewis c (HSO3-3Galβ1-3GlcNAcβ). Other reports showed that MAL strongly binds to Neu5Acα2-3Galβ1-4GlcNAc.51, 52 Our lectin microarray analysis indicated that the signal intensity of MAL is extremely low compared with that of MAH, which is correlated well with the fact that MUC1 has high-dense O-glycan clusters with few N-glycosylation site. CA19-9 also has a terminal NeuAcα2-3Gal structure. In this study, we did not compare the glycan profiling and CA19-9 value. Further analysis would be efficacious to clarify detailed profiling of the recognition specificity in these lectins.
In conclusion, we have established a robust method for direct analysis of MUC1 O-glycoforms derived from FFPE tissue sections. Our approach opens up new possibilities in research using tissue sections with glycoproteomics. In the future, this method is expected to be utilized for the O-glycome analysis of MUC1 and other mucins related to various diseases for biomarker discovery and a deeper understanding of their biological functions.
References
Hanash S . Disease proteomics. Nature 2008;422:226–232.
Hart GW, Copeland RJ . Glycomics hits the big time. Cell 2010;143:672–676.
Doerr A . Glycoproteomics. Nat Methods 2012;9:36.
Noro E, Togayachi A, Sato T et al. Large-scale identification of N-glycan glycoproteins carrying Lewis x and site-specific N-glycan alterations in Fut9 knockout mice. J Proteome Res 2015;14:3823–3834.
Hinneburg H, Stavenhagen K, Schweiger-Hufnagel U et al. The art of destruction: optimizing collision energies in quadrupole-time of flight (Q-TOF) instruments for glycolide-based glycoproteomics. J Am Soc Mass Spectrom 2016;3:507–519.
Hollingsworth MA, Swanson BJ . Mucins in cancer: protection and control of the cell surface. Nat Rev Cancer 2004;4:45–60.
Kufe DW . Mucins in cancer: function, prognosis and therapy. Nat Rev Cancer 2009;9:874–885.
Agraw B, Krantz MJ, Reddish MA et al. Cancer-associated MUC1 mucin inhibits human T-cell proliferation, which is reversible by IL-2. Nat Med 1998;4:43–49.
Paszek MJ, DuFort CC, Rossier O et al. The cancer glycocalyx mechanically primes integrin-mediated growth and survival. Nature 2014;511:319–325.
Swanson BJ, McDermott KM, Singh PK et al. MUC1 is a counter-receptor for myelin-associated glycoprotein (Siglec-4a) and their interaction contributes to adhesion in pancreatic cancer perineural invasion. Cancer Res 2007;67:10222–10229.
Fuster MM, Esko JD . The sweet and sour of cancer: glycans as novel therapeutic targets. Nat Rev Cancer 2005;5:526–542.
Storr SJ, Royle L, Chapman CJ et al. The O-linked glycosylation of secretory/shed MUC1 from an advanced breast cancer patient’s serum. Glycobiology 2008;18:456–462.
Dell A, Morris HR . Glycoprotein structure determination by mass spectrometry. Science 2001;291:2351–2356.
Rudd PM, Elliott T, Cresswell P et al. Glycosylation and the immune system. Science 2001;291:2370–2376.
Andersch-Björkman Y, Thomsson KA, Holmén Larsson JM et al. Large scale identification of proteins, mucins, and their O-glycosylation in the endocervical mucus during the menstrual cycle. Mol Cell Proteomics 2007;6:708–716.
Stoeckli M, Chaurand P, Hallahan DE et al. Imaging mass spectrometry: a new technology for the analysis of protein expression in mammalian tissues. Nat Med 2001;7:493–496.
Powers TW, Jones EE, Betesh LR et al. Matrix assisted laser desorption ionization imaging mass spectrometry workflow for spatial profiling analysis of N-linked glycan expression in tissues. Anal Chem 2013;85:9799–9806.
Powers TW, Neely BA, Shao Y et al. MALDI imaging mass spectrometry profiling of N-glycans in formalin-fixed paraffin embedded clinical tissue blocks and tissue microarrays. PLoS ONE 2014;9:e106255.
Everest-Dass AV, Briggs MT, Kaur G et al. N-glycan MALDI imaging mass spectrometry on formalin-fixed paraffin-embedded tissue enables the delineation of ovarian cancer tissues. Mol Cell Proteomics 2016;15:3003–3016.
Holst S, Hejis B, de Haan N et al. Linkage-specific in situ sialic acid derivatization for N-glycan mass spectrometry imaging of formalin-fixed paraffin-embedded tissues. Anal Chem 2016;88:5904–5913.
Kuno A, Uchiyama N, Koseki-Kuno S et al. Evanescent-field fluorescence-assisted lectin microarray: a new strategy for glycan profiling. Nat Methods 2005;2:851–856.
Kuno A, Kato Y, Matsuda A et al. Focused differential glycan analysis with the platform antibody-assisted lectin profiling for glycan-related biomarker verification. Mol Cell Proteomics 2009;8:99–108.
Matsuda A, Kuno A, Ishida H et al. Development of an all-in-one technology for glycan profiling targeting formalin-embedded tissue sections. Biochem Biophys Res Commun 2008;370:259–263.
Tan B, Matsuda A, Zhang Y et al. Multilectin-assisted fractionation for improved single-dot tissue glycome profiling in clinical glycoproteomics. Mol Biosyst 2014;10:201–205.
Matsuda A, Kuno A, Kawamoto T et al. Wisteria floribunda agglutinin-positive mucin 1 is a sensitive biliary marker for human cholangiocarcinoma. Hepatology 2010;52:174–182.
Matsuda A, Kuno A, Nakagawa T et al. Lectin microarray-based sero-biomarker verification targeting aberrant O-linked glycosylation on mucin 1. Anal Chem 2015;87:7274–7281.
Saitou M, Goto M, Horinouchi M et al. MUC4 expression is a novel prognostic factor in patients with invasive ductal carcinoma of the pancreas. J Clin Pathol 2005;58:845–852.
Higashi M, Yamada N, Yokoyama S et al. Pathobiological implications of MUC16/CA125 expression in intrahepatic cholangiocarcinoma-mass forming type. Pathobiology 2012;79:101–106.
Sobin L, Gospodarowicz M, Wittekind C et al. Classification of Malignant Tumors7th ednWiley Blackwell: New Jersey, USA, 2009.
Takeuchi H, Kato Y, Denda-Nagai K et al. The epitope recognized by the unique anti-MUC1 monoclonal antibody MY.1E12 involves sialylα2-3galactosylβ1–3N-acetylgalactosaminide linked to a distinct threonine residue in the MUC1 tandem repeat. J Immunol Methods 2002;270:199–209.
Tarp MA, Sørensen AL, Mandel U et al. Identification of a novel cancer-specific immunedominant glycopeptide epitope in the MUC1 tandem repeat. Glycobiology 2007;17:197–309.
Higashi M, Yonezawa S, Ho JJ et al. Expression of MUC1 and MUC2 mucin antigens in intrahepatic bile duct tumors: its relationship with a new morphological classification of cholangiocarcinoma. Hepatology 1999;30:1347–1355.
Tateno H, Toyota M, Saito S et al. Glycome diagnosis of human induced pluripotent stem cells using lectin microarray. J Biol Chem 2011;28:20345–20353.
Blechacz B, Komuta M, Roskams T et al. Clinical diagnosis and staging of cholangiocarcinoma. Nat Rev Gastroenterol Hepatol 2011;8:512–522.
Vincent A, Herman J, Schulick R et al. Pancreatic cancer. Lancet 2011;378:607–620.
Konami Y, Yamamoto K, Osawa T et al. Strong affinity of Maackia amurensis hemagglutinin (MAH) for sialic acid-containing Ser/Thr-linked carbohydrate chains of N-terminal octapeptides from human glycophorin A. FEBS Lett 1994;342:334–338.
Maenuma K, Yim M, Komatsu K et al. Use of library of mutated Maackia amurensis hemagglutinin for profiling the cell lineage and differentiation. Proteomics 2008;8:3274–3283.
Yue T, Goldstein IJ, Hollingsworth MA et al. The prevalence and nature of glycan alterations on specific proteins in pancreatic cancer patients revealed using antibody-lectin sandwich arrays. Mol Cell Proteomics 2009;8:1697–1707.
Wu YM, Nowack DD, Omenn GS et al. Mucin glycosylation is altered by pro-inflammatory signaling in pancreatic-cancer cells. J Proteome Res 2009;8:1876–1886.
Remmers N, Anderson JM, Linde EM et al. Aberrant expression of mucin core proteins and O-linked glycans associated with progression of pancreatic cancer. Clin Cancer Res 2013;19:1981–1993.
Espina V, Wulfkuhle JD, Calvert VS et al. Laser-capture microdissection. Nat Protoc 2006;1:586–603.
Kondo T, Hirohashi S . Application of highly sensitive fluorescent dyes (CyDye DIGE Fluor saturation dyes) to laser microdissection and two-dimensional difference gel electrophoresis (2D-DIGE) for cancer proteomics. Nat Protoc 2006;1:2940–2956.
Lok T, Chen L, Lin F et al. Immunohistochemical distinction between intrahepatic cholangiocarcinoma and pancreatic ductal adenocarcinoma. Hum Pathol 2014;45:394–400.
Ney JT, Zhou H, Sipos B et al. Podocalyxin-like protein 1 expression is useful to differentiate pancreatic ductal adenocarcinomas from adenocarcinomas of the biliary and gastrointestinal tracts. Hum Pathol 2007;38:359–364.
Hooper JE, Morgan TK, Grompe M et al. The novel monoclonal antibody HPC2 and N-cadherin distinguish pancreatic ductal adenocarcinoma from cholangiocarcinoma. Hum Pathol 2012;43:1583–1589.
Narimatsu H, Sawaki H, Kuno A et al. A strategy for discovery of cancer glycol-biomarkers in serum using newly development technologies for glycoproteomics. FEBS J 2010;277:95–105.
Irimura T, Denda K, Iida Si et al. Diverse glycosylation of MUC1 and MUC2: potential significance in tumor immunity. J Biochem 1999;126:975–985.
Fujita K, Denda K, Yamamoto M et al. Expression of MUC1 mucins inversely correlated with post-surgical survival of renal cell carcinoma patients. Br J Cancer 1999;80:301–308.
Yamaguchi T, Yokoyama Y, Ebata T et al. Verification of WFA-sialylated MUC1 as a sensitive biliary biomarker for human biliary tract cancer. Ann Surg Oncol 2016;23:671–677.
Shoda J, Matsuda A, Shida T et al. Wisteria floribunda agglutinin-sialylated mucin core polypeptide 1 is a sensitive biomarker for biliary tract carcinoma and intrahepatic cholangiocarcinoma: a multicenterstudy. J Gastroenterol 2017;52:218–228.
Kawaguchi T, Matsumoto I, Osawa T . Studies on hemagglutinins from Maackia amurensis seeds. J Biol Chem 1974;249:2786–2792.
Knibbs RN, Goldstein IJ, Ratcliffe RM et al. Characterization of the carbohydrate binding specificity of the leukoagglutinating lectin from Maackia amurensis. Comparison with other sialic acid-specific lectins. J Biol Chem 1991;266:83–88.
Acknowledgements
We thank Ms Azumi Takahashi for assistance in preparing this manuscript. This work was supported by a Grant-in-Aid for Young Scientists (B) (30722590).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website
A new and robust methodology for differential O-glycome profiling on an endogenous tumor-associated glycoprotein, MUC1, is proposed in this paper. This method enables glycome analysis of small quantities of samples collected from single formalin-fixed, paraffin-embedded tissue sections and is applicable to other O-glycosylated proteins. This technique is promising for discovery of biomarkers and therapeutic targets for a large number of diseases.
Supplementary information
Rights and permissions
About this article
Cite this article
Matsuda, A., Higashi, M., Nakagawa, T. et al. Assessment of tumor characteristics based on glycoform analysis of membrane-tethered MUC1. Lab Invest 97, 1103–1113 (2017). https://doi.org/10.1038/labinvest.2017.53
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2017.53
This article is cited by
-
Role of Wisteria floribunda agglutinin binding glycans in carcinogenesis and metastasis of cholangiocarcinoma
Histochemistry and Cell Biology (2024)